The elite journal Nature Cell Biology (NCB) has a very commendable policy on data integrity: the deposition of raw data, in particular original scans of western blots and other gel analyses. This is to avoid dishonest authors playing tricks with loading controls, antibody specificity and stealthy gel splicing or even band duplications. The rule however starts applying only after the paper passed peer review and was accepted for publication, and because of that, some authors deposit any odd gel picture and count on nobody bothering to check, or to take action.
This was exactly what the Spanish star cancer researcher Carlos López-Otín, professor of Biochemistry and Molecular Biology at the University of Oviedo, did. For his lab, the disregard for loading controls is standard practice: he even managed to reuse same picture of one no less than 23 times (read here). Recently, some of those papers, all at Journal of Biological Chemistry (JBC), had to be corrected.
The embarrassment for the Nature family journal NCB is the bigger because the flagship Nature issued in 2017 a Mentoring Award to Lopez-Otin (read here), where one really wonders what exactly Nature aimed to reward here. The skill of publishing in their own high impact journals using a sod-all, who-cares attitude to research integrity, which Lopez-Otin taught his mentees?
The paper in question is the following, and though it was not yet discussed in my articles, the comment section under my previous two articles (here and here) was largely dedicated to it:
Clara Soria-Valles , Fernando G. Osorio , Ana Gutiérrez-Fernández , Alejandro De Los Angeles , Clara Bueno , Pablo Menéndez , José I. Martín-Subero , George Q. Daley , José M. P. Freije , Carlos López-Otín
NF-κB activation impairs somatic cell reprogramming in ageing
Nature Cell Biology (2015) doi: 10.1038/ncb3207
The first author Clara Soria Valles is presently postdoc with another significant co-author of that paper, George Q Daley. Do you know who Professor Daley is? Merely the Dean of Harvard Medical School and one of the most important and influential people worldwide in the field of stem cell research. According to Author Contributions statement, “G.Q.D. provided critical materials and participated in the preparation of the manuscript“. It is not clear if that participation involved the checking of data integrity and if those “crucial materials” Daley provided were already published (if they were, he should not have claimed co-authorship). In any case, with Daley and his postdoc on board, the correction of that paper will become a rather sensitive and political issue. If it ever happens, that is. With that paper and the help of Daley, the cancer researcher Lopez-Otin was able to conquer another piping hot and highly lucrative research field: that of stem cells and regenerative medicine. It earned him an ERC grant right away, worth of €2.5mn from 2017 to 2022, to work on continuation of his NCB study, on “Deconstructing Ageing: from molecular mechanisms to intervention strategies“. It is very convenient that the elite European public funder ERC does not seem to care a bit about research integrity, and is rather proactive in that regard.
The paper in Nature Cell Biology which made all this possible, postulated the decisive role of the transcription factor protein NF-kB in cellular senescence as barrier to reprogramming of cells to pluripotency, and offered a potential cure to child patients suffering from the deadly premature ageing syndrome, which is caused by genetic and cell nucleus defects, specifically the Néstor–Guillermo progeria and the Hutchinson–Gilford progeria.

Lopez-Otin’s impactful publication of 2015 must have given hopes to the families of sick children. If only they knew how sloppily the data was assembled.
Much of the sleuthing came from the pseudonymous Clare Francis, who comments and shares image evidence on my site as “Zebedee”, several others also contributed on PubPeer. There are many issues with that NCB paper, the PubPeer thread went over 120 comments, despite moderation. Some of the criticism appears misguided, when two or more western blots photographed together on one film were confused for a single gel. But most of the PubPeer evidence has its validity. For brevity, I will present below only select evidence.
The debate started almost exactly one year ago, in September 2017 with this comment, where a mismatch between figure and raw data in 3 figures was reported:
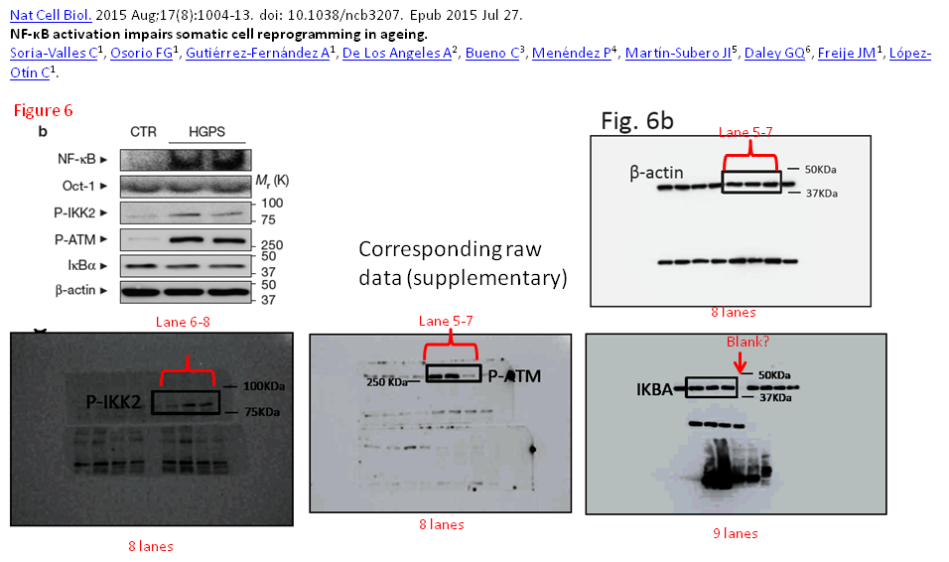
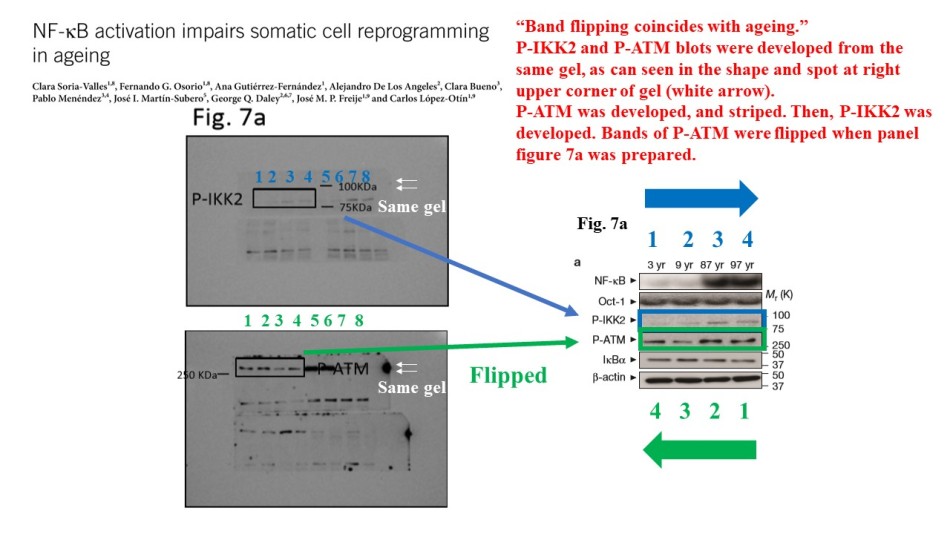
Figure 6b shows an array of 6 Western blots, with b-actin being the loading control, and no two of these panels came from the same gel. They all, p-IKK2, p-ATM, IkBa and the loading control b-actin came from a separate gel each, presented in the final figure as a single blot. Further PubPeer debate proved that also NF-kB and Oct-1 signals came from two more gels. Note that the lanes of original blot scans are not labelled, there is also no legend.
The signal for NF-kB protein in that figure and in another one, Figure 3c, does not really match the original scan. Basically, the authors failed to provide correct gels for their NF-kB data. And the ones they did provide show fat and rather specific looking bands underneath. If the authors cropped this image data, it was against Nature‘s own instructions for authors. But then again, those do not seem the correct blots anyway.
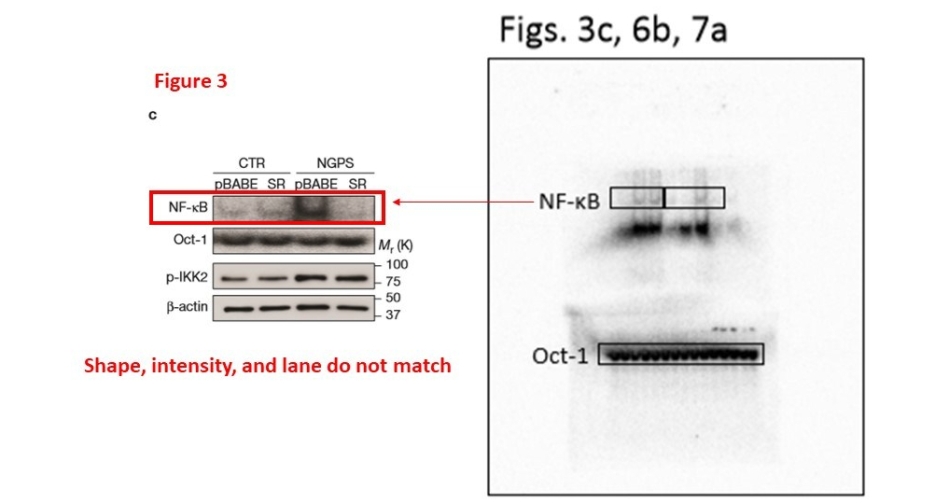
The NF-kB is the central analysis of the study, it is the hub protein of the provided key mechanism and even part of the paper’s title. Surely the NF-kB data should be the most clear and reliable? Yet also in Figures 3c and 6b, the data in the figure doesn’t match what the authors provided as raw data. From Oct-1 analyses it also becomes rather obvious that the Lopez-Otin lab cannot be bothered with proper loading controls. Some 14 samples were loaded on a gel and used for each of the 3 separate experiments in need of some Oct-1 data. The bands don’t seem to match. Is this even the Oct-1 signal the gel shows, or maybe a forgotten loading control from yet another experiment?

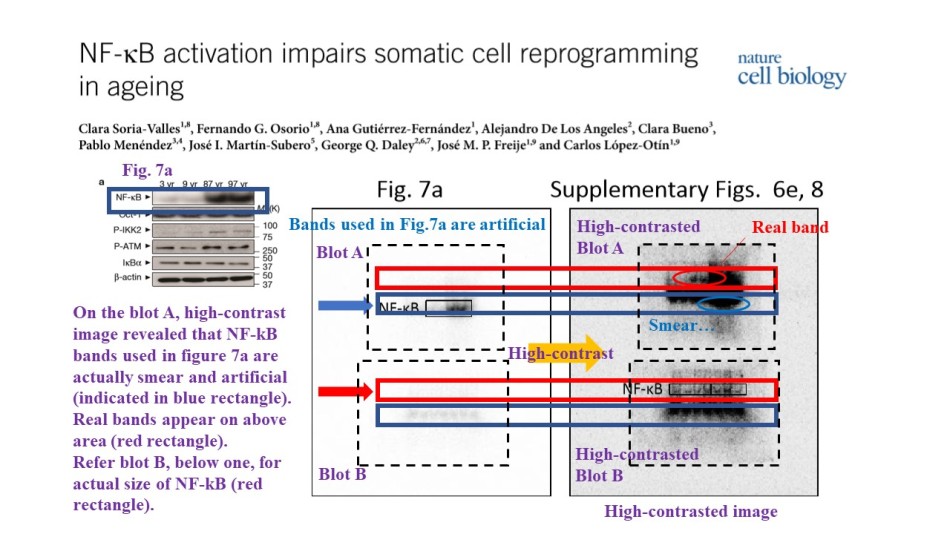
The authors did same again, with yet another “Oct-1” and “NF-kB” gels for Supplementary Figure 8, as commented on PubPeer. In fact, what are those published NF-kB bands anyway? After a PubPeer user boosted the contrast of raw gel scans, more bands appeared. Because the authors did not bother to provide molecular weigh marker scale as requested by the publisher, we cannot know which of the smear (if any) might correspond to the correct NF-kB band.
This is what Nature Publishing Group itself has to say as their expectations towards the authors about the data integrity of western blots (highlights mine):
Electrophoretic gels and blots
Positive and negative controls, as well as molecular size markers, should be included on each gel and blot—either in the main figure or an expanded data supplementary figure. […]
The display of cropped gels and blots in the main paper is encouraged if it improves the clarity and conciseness of the presentation. In such cases, the cropping must be mentioned in the figure legend. (Some journals require full-length gels and blots in supplementary information wherever possible [as does Nature Cell Biology, -LS].)
- Quantitative comparisons between samples on different gels/blots are discouraged; if this is unavoidable, the figure legend must state that the samples derive from the same experiment and that gels/blots were processed in parallel. Vertically sliced images that juxtapose lanes that were non-adjacent in the gel must have a clear separation or a black line delineating the boundary between the gels. Loading controls (e.g. GAPDH, actin) must be run on the same blot. Sample processing controls run on different gels must be identified as such, and distinctly from loading controls
- Cropped gels in the paper must retain important bands.
- Cropped blots in the body of the paper should retain at least six band widths above and below the band.
- High-contrast gels and blots are discouraged, as overexposure may mask additional bands. Authors should strive for exposures with gray backgrounds. Multiple exposures should be presented in supplementary information if high contrast is unavoidable. […]
Of course, those rules might not apply to friends of Daley’s like Lopez-Otin. This is also why three physically different gels were used to make the small Figure 2a. The gel for Lamin B1 has 9 lanes (plus molecular weight marker), the gels for p16 and b-actin 6 lanes. Yet also those latter two must be separate gels, because the p16 signal is in different orientation. Either that, or the authors didn’t like the result of their p16 experiment and flipped it around so the progeria samples (NGPS) would show more of the senescence-associated protein p16.
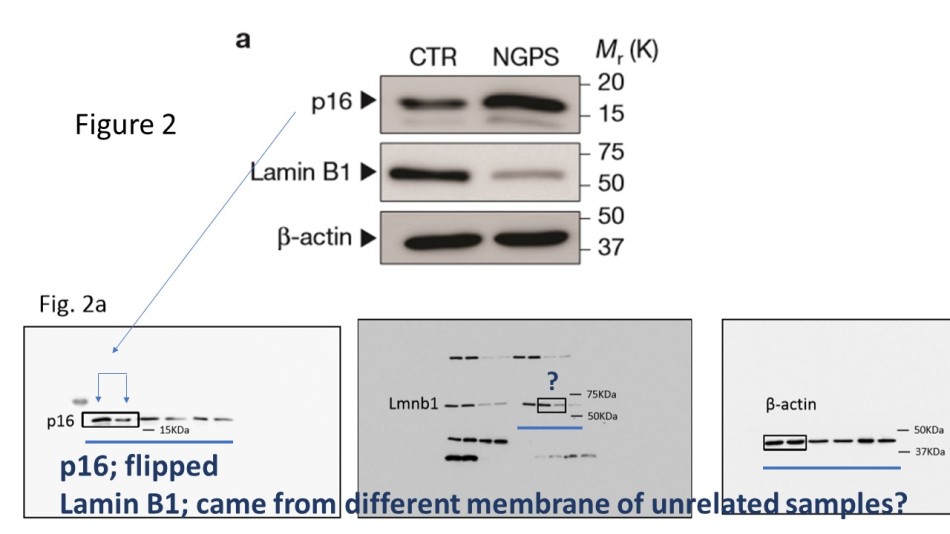
Figure 4i has another flip, the gel bands for the methylated histone protein H3K79me2 are reversed. Why would one do that? The reverse order to load a gel like that would be highly unusual and illogical. What was going on there? The original loading control seems to have been already used in Figure 3C. It definitely was a different gel from the methylated histone analysis from Figure 4i shown below. Never mind all that, the figure was good enough to be quantified into a bar diagram. Normally one needs triplicates for that, but the authors forgot to upload them, or maybe simply couldn’t find something which might match distinctly.

Yet here it is again, this loading control, reused in the best Lopez-Otin manner. Yes, admittedly those are same samples, according to figure legends. But not from the same gels, as raw data indicates. Why was it such a problem for the authors to probe each of these gels for equal loading and to provide correct loading controls? Why do we have to trust them blindly to have loaded all samples in equal amounts on different gels (if interested, here is more on importance of loading controls)? I am not sure NCB‘s impact factor and elite status of peer review is any argument on why controls should be suspended.
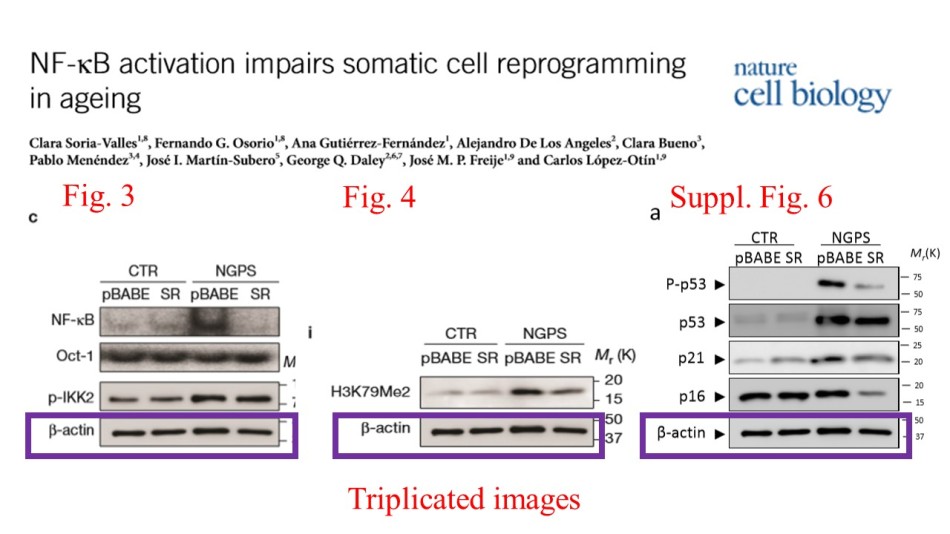
There was even more band flipping, in Figure 7a (a similar event took place also in Figure 6b). It seems to be same gel, stripped of the previous antibody signal and re-probed. So why the flip? A mistake? Was one of authors bored, and wished to troll? Or was something wrong with the results, did they need some fixing? Of course the loading control doesn’t match, and it is anyone’s guess what “NF-kB” and “Oct-1” bands might have been in their other life.
Also in Figure 8B, the authors apparently probed the same gel, but maybe once again found themselves unhappy with the result. The methylated histone lanes used were samples 2, 3, 4, but the b-actin loading controls were 3, 2, 1, in this orientation. From the same gel, mind you. Maybe the loading control was “unusable”, because the differences in H3k79me2 signal would be less prominent if b-actin was matched correctly?

Lopez-Otin and his co-authors made a mockery of good scientific practice and the concept of loading controls by uploading as raw data whatever they had at hand. The publisher must now decide if it wishes to defend its guidelines of “Loading controls (e.g. GAPDH, actin) must be run on the same blot” and “Cropped gels in the paper must retain important bands” or in fact, that “the final image must correctly represent the original data and conform to community standards“.
And of course maybe the scientific community and the progeria patients’ families should be cautioned through a correction notice how the data which gave them hope was assembled.
Update 17.10.2018. Lopez-Otin left Spain and University of Oviedo and now works in France, as a reader noted in a comment. As La Nueva Espana reported already in June:
“I do not know when Carlos Lopez Otín will return to the University of Oviedo”, said yesterday the rector, Santiago García Granda, who considers the stay of Professor of Molecular Biology at the University Pierre and Marie Curie in Paris as a natural matter. “Moving is a normal thing”.
The 60 year old Lopez-Otin now set up a research group at the Centre de Recherche des Cordeliers in Paris, obviously on the invitation of his past collaborator and that institute’s deputy director Guido Kroemer. Several of Kroemer’s papers are flagged for suspected data manipulation on PubPeer, and he was one of signatories of a recent Stalinist letter organised by CNRS which defended research misconduct and denounced whistleblowing and critical journalism (more on that here).


Donate!
If you are interested to support my work, you can leave here a small tip of $5. Or several of small tips, just increase the amount as you like (2x=€10; 5x=€25). Your generous patronage of my journalism will be most appreciated!
€5.00

Thanks Leonid for covering the issues on NCB paper from Lopez-Otin, George G. Daley and others.
PubPeer did not take my last post, but I am glad you conveyed it.
Gracias!
“I have been following PubPeer comments on Carlos Lopez-Otin’s Nature Cell Biology paper (Soria-Valles et al). “this paper is dedicated to the memory of Néstor M.O.” a donor of Néstor-Guillermo progeria syndrome (NGPS) used in figure 2a that was distrustfully flipped to make up over-expression of p16 in Néstor’s cells. And sadly more sample switches of HGPS (another type of progeria) at figure 6b and of >87 yrs aged-donor cells at figure 7a to make up activation of NF-kB pathway.
As a researcher involved in ageing study, I cannot believe their working ethics for samples provided by progeria patients and aged donors (>87 yrs). The patients (and their families) had donated their own cells and tissues hoping to advance science, but Lopez-Otin and colleagues, including Pabro Menéndes (Jossep Carreras Imstitute) and George Q. Daley (Dean of Harvard Medical School) who provided crucial materials, bit the hand that feeds. Lopez-Otin and colleagues should be rewarded on their disrespectful manner on science and their patients.”
LikeLike
Sure these guys will find the right blots in their drawer and NCB will believe in that joke. I will send a suggestion to our Institute to stop subscription of NCB and list it as a predatory journal.
LikeLike
NPG and other high impact journals need to do do a serious introspective analysis and opt for a more transparent publishing process if they want to survive in the not so far future
LikeLike
As a certain world-wide renowned scientist I refuse to submit anything to Nature…I hope many other scientists follow this example
LikeLike
This NCB paper is connecting dots of Carlos Lopez-Otin’s progeria legacy with that of George Daley in the manner of both groups. NF-kB (aka “David Baltimore pathway” who is a mentor of George Daley https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2742082/ ), Dot1L (George Daley’s big achievement in stem cell field https://www.ncbi.nlm.nih.gov/pubmed/22388813 ), and reprogramming (as everyone knows, George Daley’s top discovery). It’s like “pizza inside a burger inside a pizza”. Central paper for both labs.
LikeLike
Almost record number of issues that one paper can have. Not only reusing blots, but Lopez-Otin developed more skills of band flipping, capturing smears and switching samples. So any action to this paper, NCB?
LikeLike
Good post, Leonid. Yes, PubPeer does lots of moderation to George Q. Daley, dean of Harvard Medical School. PubPeer could be as tough as CNS journals when it comes to putting comments on his papers.
Pubpeer is biased, unfortunately.
5 out of 7 comments deleted.
https://pubpeer.com/publications/32494494DF1BA35389D4DD4215FDB6
2 out 2 comments deleted.
https://pubpeer.com/publications/9FFA7E2C80B6F11BDF006C3E6A1C1D
1 out of 5 comments deleted.
https://pubpeer.com/publications/CC2645E6699CB04102B6632962552C
Several comments had been deleted without traces being left, peers claiming those were totally gone and unseen.
https://pubpeer.com/publications/03698513C41957E1BFDE9396741C97
LikeLike
It is so unfair that we struggle to be able to get funded to do some honest research work
and these kind of people cheats, gets funding, appear in TV like movie stars joking with all of us
LikeLike
Researchers should join forces and more actively put pressure on research institutions, publishers and funding sourches. I think one of the reasons why these guys can continue with their cheating is that too few researchers care about more than their tiny piece of study.
Working researchers of all countries, Unite!
LikeLike
PubPeer comments to Soria-Valles et al., NCB paper are still continuing.
NF-κB activation impairs somatic cell reprogramming in ageing
Nature Cell Biology (2015) – 135 Comments
pubmed: 26214134 doi: 10.1038/ncb3207 issn: 1465-7392 issn: 1476-4679
Clara Soria-Valles , Fernando G. Osorio , Ana Gutiérrez-Fernández , Alejandro De Los Angeles , Clara Bueno , Pablo Menéndez , José I. Martín-Subero , George Q. Daley , José M. P. Freije , Carlos López-Otín
https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18#137
“I thought that two or more western blots photographed together on one film. That explains why blots of different primary abs appear on one film. To check this, I changed contrast of this original blot corresponding to figure 2a. As you see, there is no margin of different membranes on the film. So I would say those bands on the film came from single blot with one type of primary ab. As molecular marker and label are absent, I can not be sure what those bands are.”
“In supplementary figure 1e, samples 10-11 were missing from BAF blot. Lanes 8-9 of BAF and b-actin blots did not seem to come from the same gel. Again, lack of molecular marker and label leaves the identity of bands with dismay.”
“Size of NF-kB was not consistent between figures 3c, 6b, and 7a. Which bands of NF-kB are right?”
LikeLike
At least what is truth in this NCB paper is that Carlos Lopez-Otin is not yet as high-class enough as George Q. Daley for whom PubPeer is seriously censoring comments. It is ironic. If Daley were to be corresponding, PubPeer comments would never grow as it is. And the paper would have been on Nature.
LikeLike
It is weird if PubPeer is favor to some high power PIs of great social status. If true, very dangerous and scary to rigor of science…
LikeLike
It has been a while since this NCB paper becomes a hottest paper on PubPeer. It is unusual a paper can thrive for long with so may obvious issues.
Wondering any action to this paper.
LikeLike
I do not think NCB takes any action for this paper. The rigor of the paper is guaranteed by Spanish top star Carlos Lopez-Otin and the dean of world highest med school George Q. Daley. So there is no mistake.
It all matters who had published, not what to publish.
LikeLike
PubPeer comments on credibility of colony counting techniques and statistics in Soria-Valles et al. NCB paper.
NF-κB activation impairs somatic cell reprogramming in ageing
Nature Cell Biology (2015) – 149 Comments
pubmed: 26214134 doi: 10.1038/ncb3207 issn: 1465-7392 issn: 1476-4679
Clara Soria-Valles , Fernando G. Osorio , Ana Gutiérrez-Fernández , Alejandro De Los Angeles , Clara Bueno , Pablo Menéndez , José I. Martín-Subero , George Q. Daley , José M. P. Freije , Carlos López-Otín
https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18#
“Not only WB band issues, but also their colony counting is cryptic and worth more attention. Their represent images and bar graphs do not match. This is something grilled into me when Method section did not describe how the authors quantified images. This paper depends on colony counting a lot. Figures 1b, 1d-g, 3c, 3e-g, 5a-b, 5d-f, 6a, 6c-d, 7b-c, s1g, s4a-b, s5b-c, s5e-f, s6b, and s6d.”
https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18#132
“More prominent example can be found in Figure 6. Two independent samples, one had very less and the other had much more number of colonies in the original images, had alsmot identical bar with tiny error. p=0.006, n=3. Even if the other two experiments had equal number of colonies between samples, p value cannot be such low as 0.006.”
https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18#137
“I agree that colony countings are too subjective and lack consistency unless employed machine-based countings. The earlier posts focused on the issues in colony countings in this paper. Also, I would like to point out that too tiny error bars throughout this paper is something grilling into me.”
https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18#139
“Following the author’s description of statistical analysis, two-tailed Student’s t-test was done with n=3. Assume IKKi colonies = 15, 16, 17; IKK2-KI colonies = 16, 18, 20. The most favorable numbers for authors that I could estimate from their represent images and bar graphs in figure 6b. t=test will be p= 0.07418. Still never be p=0.006 that authors reported in the paper. Curious if source data is available.”
https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18#141
“The issue of colony counting and statistics all came from lack of source data. Actually, NPG has a policy to deposit source data used for any statistical analysis. Example, another NCB paper of the same year (2015) did deposit source data. https://www.nature.com/articles/ncb3073#supplementary-information ”
https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18#147
“The Methods section described in the paper. The colony counting did noy exploit machine counting, rendering subjective results as discussed in earlier posts. “Reprogramming efficiency. For quantification of iPSC reprogramming efficiency, retroviral transduction was carried out in parallel infections containing all of the retroviruses used for reprogramming plus a tomato-expressing retrovirus. The total number of human iPSC colonies was counted after immunostaining with Tra-1-60 specific antibody as previously described 36, or by detection of alkaline phosphatase activity in murine iPSCs (AP detection kit, Sigma) following the manufacturer’s instructions.””
https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18#152
LikeLike
More comments to Soria-Valles et al. NCB paper on PubPeer.
NF-κB activation impairs somatic cell reprogramming in ageing
Nature Cell Biology (2015) – 149 Comments
pubmed: 26214134 doi: 10.1038/ncb3207 issn: 1465-7392 issn: 1476-4679
Clara Soria-Valles , Fernando G. Osorio , Ana Gutiérrez-Fernández , Alejandro De Los Angeles , Clara Bueno , Pablo Menéndez , José I. Martín-Subero , George Q. Daley , José M. P. Freije , Carlos López-Otín
https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18
“Supplelentary figure 6c has several issues.
pBABE p16 band was highly contrasted to become thicker, although its corresponding IkB-SR p16 band on the same lane was not.
the pattern of 3 black spots at the right corner of IkB-SR p16 band differ between figure and original data. Upper left spot nearest to band is thickest in the figure, but lower right one is thickest in the original data.
b-actin bands are different from raw data.”
“Figure 7f tried to show that expression of NANOG was reduced in aged fibroblasts. However, human fibroblasts do not express NANOG regardless of age (Takahashi, 2007 Cell. PMID 18035408). Actually, another figure (supplementary 2) in this paper clearly showed lack of expression of NANOG in human fibroblasts.”
“The same protein BAF showed different number of bands between figures.”
LikeLike
Not only the right blots in their drawer, but also the authors will find source data of Excel files in their laptops to vindicate their tiny error bars and minimum p values. And you will see reincarnated version of this paper soon!
LikeLike
Pluripotency genes are expressed in human adult fibroblasts, and elicited by a mere inhibition of NF-kB? If it is true, Carlos Lopez-Otin and George Q. Daley will be nominated for Nobel prize. And strip the prize from Shinya Yamanaka whose cumbersome ground-state cell magic was overtuned from the ground. If adult fibroblasts already express pluripotent genes, how could Yamanaka defined his four-factors based on elicitation of NANOG.
I bet on Daley, who pioneered reprogramming and called for cleanse STAP drama, has achieved real and important break-through again!
LikeLike
Inducing pluripotent genes in adult fibroblasts? Yes, we believe in the power of David Baltimore pathway (NF-kB) who formed George Q. Daley into one of the first to discover reprogramming. Better mind that low-pH stressor cannot make it…
LikeLike
PubPeer readers picking up another case of pluripotency gene expression in adult fibroblasts in Soria-Valles et al., 2015 NCB paper.
NF-κB activation impairs somatic cell reprogramming in ageing
Nature Cell Biology (2015) – 152 Comments
pubmed: 26214134 doi: 10.1038/ncb3207 issn: 1465-7392 issn: 1476-4679
Clara Soria-Valles , Fernando G. Osorio , Ana Gutiérrez-Fernández , Alejandro De Los Angeles , Clara Bueno , Pablo Menéndez , José I. Martín-Subero , George Q. Daley , José M. P. Freije , Carlos López-Otín
https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18#155
“Same example in figure 6. Expression of NANOG in adult human fibroblasts was detected, which is contradicting with their own supplementary data.”
https://imgur.com/a/IxwbEkl
LikeLike
Figure duplication and horizontal flip from Osorio et al., 2012 Genes and Development.
Nuclear lamina defects cause ATM-dependent NF-κB activation and link accelerated aging to a systemic inflammatory response
Genes & Development (2012) – 19 Comments
pubmed: 23019125 doi: 10.1101/gad.197954.112 issn: 1549-5477 issn: 0890-9369
Fernando G Osorio , Clea Bárcena , Clara Soria-Valles , Andrew J Ramsay , Félix De Carlos , Juan Cobo , Antonio Fueyo , José M P Freije , Carlos López-Otín
https://pubpeer.com/publications/4E89FBEE8E6605B592593CF801E7BD#20
LikeLike