On the Iberian peninsula, there seems to be a tradition to give well-connected scientists suspected (or even convinced) of data fudging an award. In Spain, Carlos López-Otín, Professor of Biochemistry and Molecular Biology at the University of Oviedo, was given a Mentoring Award from the elite journal Nature, on recommendation from Spanish academia and despite evidence of data irregularities in his papers. This prompted my readers, in particular the famous pseudonymous data integrity sleuth Clare Francis, to comment on on PubPeer and on my site (as “Zebedee”) with additional evidence, which made Lopez-Otin’s scientific credibility look progressively worse and worse, with each new post.
Eventually, an image of a Northern blot (showing expression of mRNAs which code for proteins) was found to appear recurrently across several papers from that Oviedo lab, where the authors pretended it was a newly produced analysis. In reality, it was a “library” loading control reused so the authors could re-run same RNA gel of human tissue lysates over the years and never check ever again what they have actually loaded on their gels. Eventually Lopez-Otin et al even stopped caring what order of samples that original loading control had.
Clare Francis was soon joined on his quest for the Perennial Northern Blot of Oviedo by Elisabeth Bik, famous microbiology blogger and image duplication detective, and my regular contributor (also pseudonymous) Smut Clyde, who now presents you the findings of no less than 23 appearances of that same northern blot in 23 publications from Lopez-Otin’s lab in the guest post below. It is just as convincing as if the Spanish actor Antonio Banderas appeared in 23 different films still dressed in same costume from his 1995 hit Desperado, carrying same guitar case. Incidentally, also Lopez-Otin’s Perennial Northern Blot made its first appearance at around that year.

The cancer researcher Lopez-Otin is an actual real-life celebrity in Spain. On 29 June 2017, he helped the King of Spain open the Princess of Girona Foundation Awards ceremony, together with another Spanish celebrity of the creative art of the pretend, Antonio Banderas. The Princess Foundation wrote about their 2017 gala host:
“Carlos Lopez-Otín is an academician of the European Academy and the Royal Academy of Sciences of Spain, and Doctor Honoris Causa by several Spanish and foreign universities. Throughout his scientific career he has received several awards such as the European FEBS Prize for Biochemistry, the DuPont Award for Life Sciences, the “Carmen and Severo Ochoa” Award, the Mexico Prize for Science and Technology, King Jaime I Prize Research and the National Research Award “Santiago Ramón y Cajal”
That multiple award winning research star is also EMBO member, just like some other of his Spain-based colleagues of questionable research integrity are: Maria Pia Cosma and Pura Munoz-Canoves. These two also regularly receive juicy research grants and recognition: Munoz-Canoves’ most recent was the “Vanguardia de la Ciencia” award, while Cosma was given in 2016 “Ciutat de Barcelona” award. Another Spanish researcher with shady data in his papers is Manel Esteller, he also gets awarded regularly, in 2016 it was the Catalonia International Prize from (now fugitive) President Puigdemont. In Portugal, Esteller’s former PhD student and now zombie scientist Sonia Melo was given a Prémio FAZ Ciência award from Fundação AstraZeneca, only two months ago. There are surely more of such questionable Iberic awardees, readers are welcome to salute these stars of Photoshop in the comment section.
I personally have a theory that in this way the system of Iberian academia announces who is untouchable, in order to intimidate critics of research misconduct in their own ranks. Probably a pathetic left-over from the fascist past of Spain and Portugal. The subliminal message is: yes, we all know what these award-winners did to create those big papers and we don’t mind at all. The award-giving farce shows who the role models are and what Spanish and Portuguese scientists are expected to do with their silly notions of research integrity: shove it, start making big papers whatever it takes, or lose your job. In fact, not even the arch-zombie scientist Susana Gonzalez had to suffer unemployment, unlike masses of honest young Spanish scientists.
A Perennial Northern Blot, by Smut Clyde
The title of this post refers to the famously picaresque Western blot belonging to a Brazilian diabetes researcher.  In its protean versatility, Mario Saad‘s pentadecaplicating blot could transform itself into any protein — tubulin, actin, GLUT4, IRS1 — from any combination of source conditions. It thereby appeared in at least 15 versions, spread across 10 papers in “an intricate publishing web“, serving as the loading control in that many different experiments (that is, as a measure of the total level of extracted protein, for normalising the measurements of the protein of interest). In my imagination it spoke with the voice of Robin Williams. This site forwarded a report on the Wandering Western… the ensuing saga included editorial Expressions of Concern, lawsuits, an investigation by Saad’s university that saw no evidence of misconduct, and 13 retractions so far (RetractionWatch are keeping score).
In its protean versatility, Mario Saad‘s pentadecaplicating blot could transform itself into any protein — tubulin, actin, GLUT4, IRS1 — from any combination of source conditions. It thereby appeared in at least 15 versions, spread across 10 papers in “an intricate publishing web“, serving as the loading control in that many different experiments (that is, as a measure of the total level of extracted protein, for normalising the measurements of the protein of interest). In my imagination it spoke with the voice of Robin Williams. This site forwarded a report on the Wandering Western… the ensuing saga included editorial Expressions of Concern, lawsuits, an investigation by Saad’s university that saw no evidence of misconduct, and 13 retractions so far (RetractionWatch are keeping score).
The present case also concerns re-use of a loading control, but this time featuring a Northern blot. The compass-point tradition for naming gel-electrophoresis techniques began with Sir Edwin Southern, pioneer of Southern blotting, for this is how humor works in molecular biology. It has been explained to me that Northern blots do not directly measure the popularity of a protein in the cellular economy; instead, mRNA (encoding for a protein) is the chemical species, extracted from various sources (lanes), and spread out into bands according to molecular weight. Then transferred (blotted) from the electrophoresis gel to a filter for stability, and detected by inducing the mRNA to bind to a matching and radiotracing DNA probe.
So in this case, a team of researchers have a bank of 28 “cell smoothies”: two sets of eight tissue types, one set of eight cancer-cell lines, and four fetal-tissue samples. In a series of papers published over a decade, the team have characterised numerous proteins from within the self-organising complexity of the human cell — sequencing the DNA for each protein and specifying its chromosomal location, describing its role within that complexity, and checking which tissues express it (which depends on which genes remained active in each lineage of cells that differentiated and specialised and became a tissue). That is to say, the Northern Blots were just one aspect of the papers, and they are all outside my comfort grade and above my pay zone.

Each study took a few drops from the stored samples, blotted it (“Filters containing about 2 µg of polyadenylated RNAs from the indicated human tissues”), and probed for the mRNA of choice. But there are limits to the precision that a pipette can provide — even in the hands of a trained gene-modified laboratory monkey — so the final stage is to wash the probe DNA out of the filter and probe it again for Actin (a background “housekeeping” protein, required by cells to maintain their architecture, unless they are dead) to correct for the actual aliquots that were used. Thus papers in this sequence typically include a phrase along these lines:
“Filters were subsequently hybridized with a human actin probe to ascertain differences in RNA loading”.
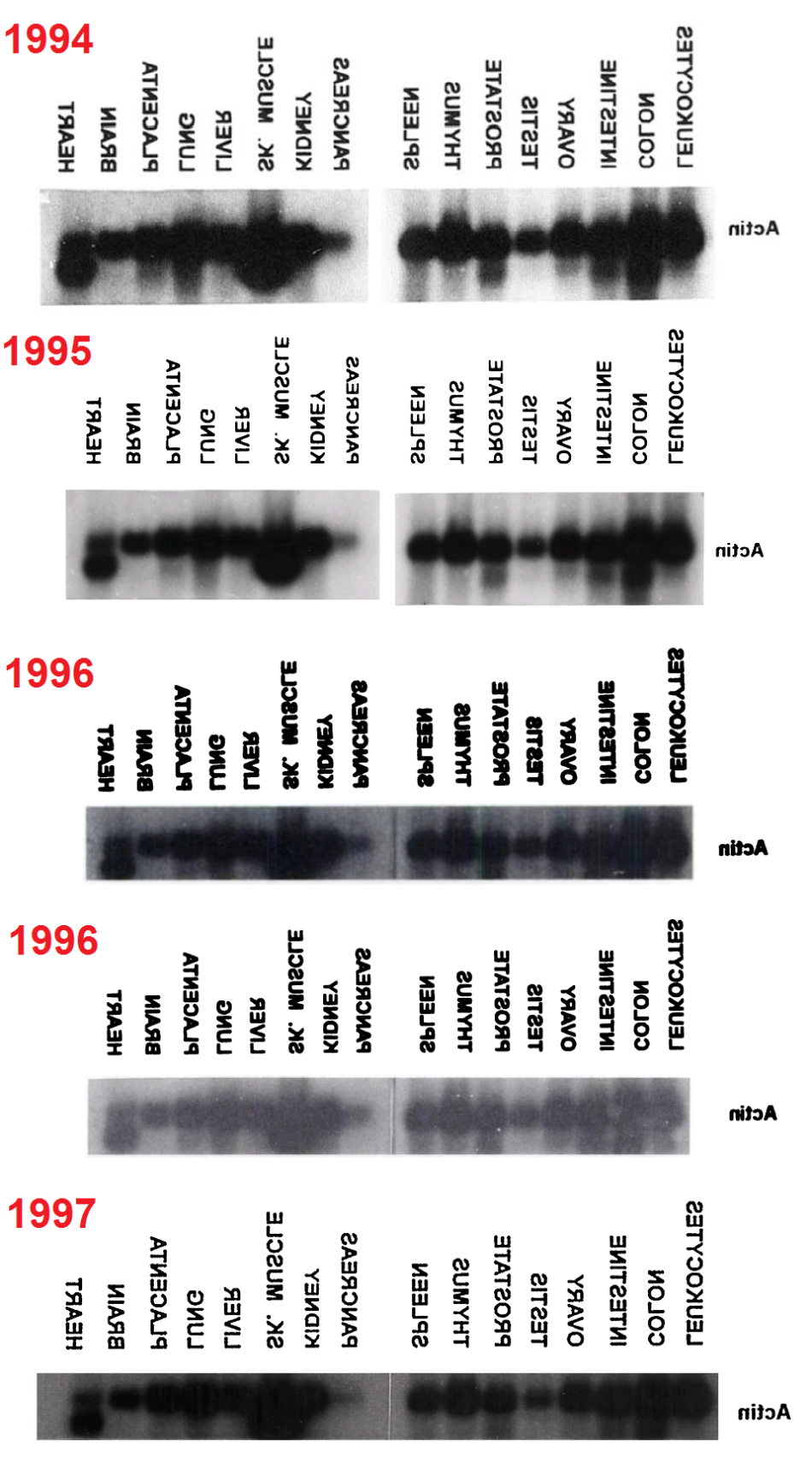
It is conceivable, however, that this phrase was repeated from the first paper, along with the loading blot itself. Comparing 23 papers, there appears to be one original blot for each bank: four blots, which are variously compressed and clipped according to the exigencies of publication, and varying also in exposure, rather than a separate measurement after each separate exercise in tissue localisation. The sources are ‘Zebedee’, commenting on threads at this site; anonymous contributors to PubPeer threads; Elisabeth Bik; and myself.
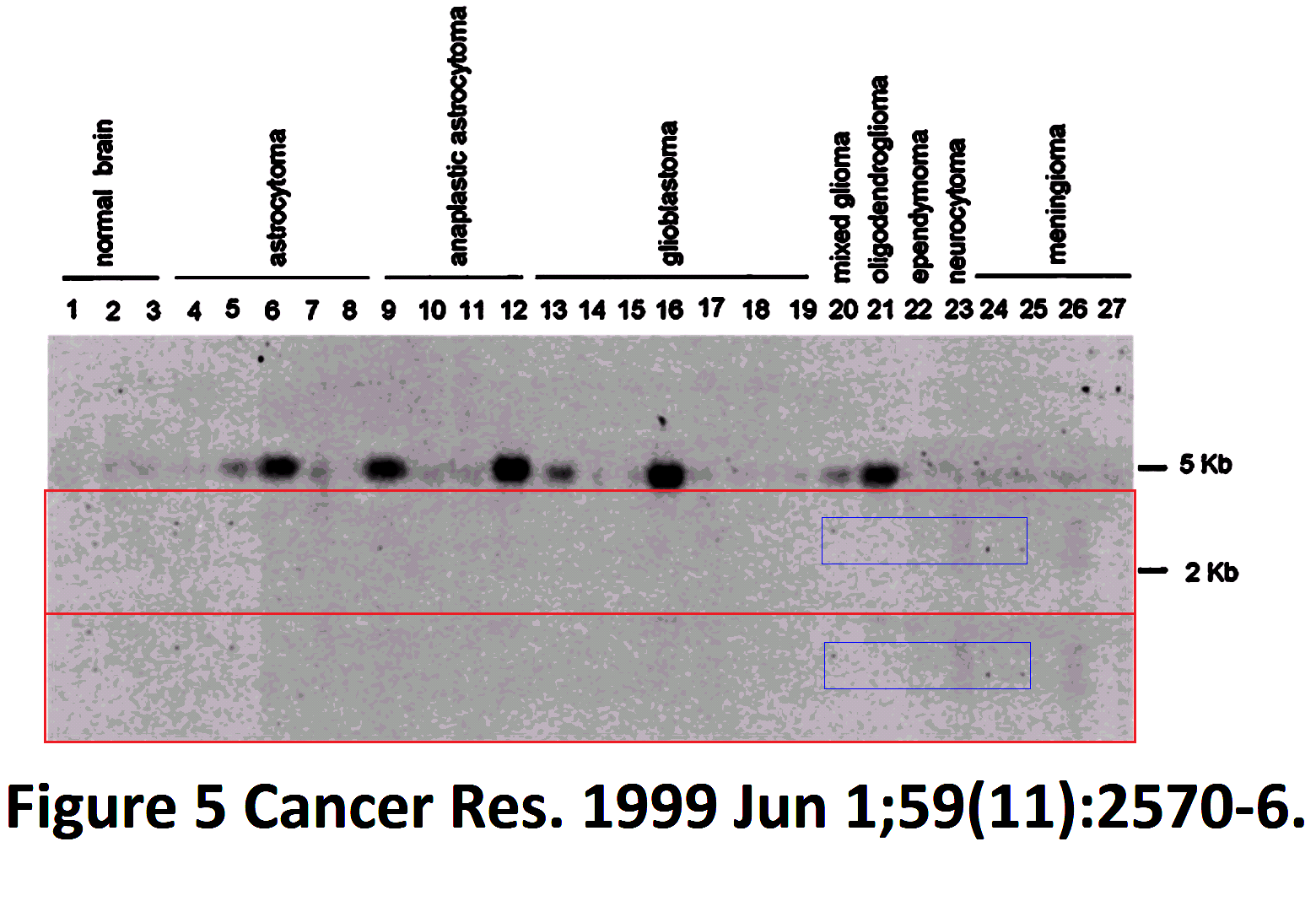
This comes to our notice because a 23-fold replication beats the 15-fold record of the Brazilian wanderer. Crucially, though the possible copies are consistently identified as Actin, and the authors have tried to label the sources of the lanes consistently. The reuse of a ‘loading library’ is deprecated, but this does not begin to approach the problematic level of the Brazilian Western: there was no attempt to mislead (other than the claim that the control in each study was specific to it, made subsequently to the data to be controlled). It is a perennial blot, always in the same place, rather than a wanderer or vagrant.
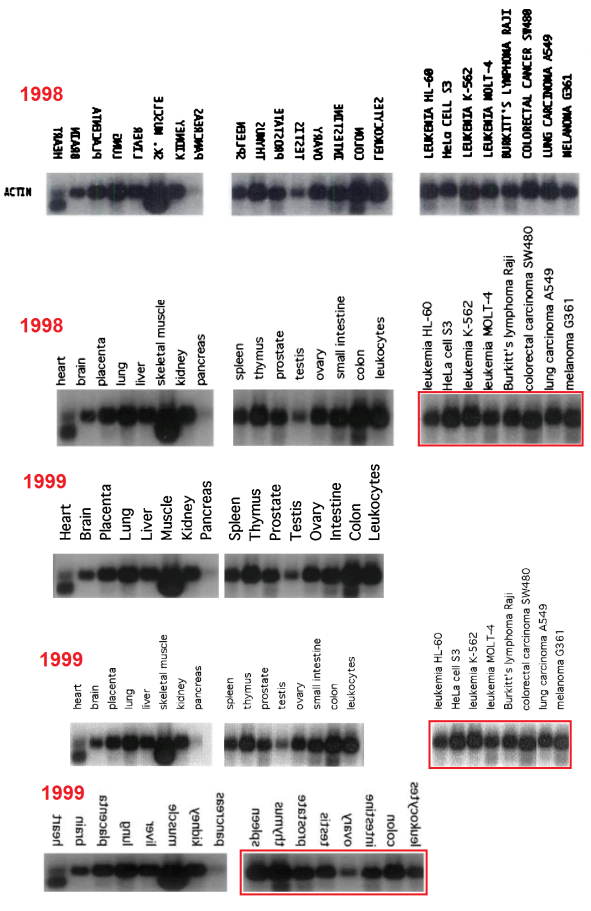

Regrettably, the labelling of lanes was not as consistent as was intended. In a 2003 appearance of the fetal-tissue blot, it was flipped horizontally relative to the lane labels, as marked with a red box in the Figures. Note that in some publications the lanes are listed in reverse order — from Leucocytes to Heart rather than vice versa — and in the Figures I have flipped each band and labels in such cases, to keep a single sequence of tissue types (hence the mirror-image text in places).
Red boxes were also necessary in some cases where the cancer-cell blot was flipped relative to its lane labels, and for the #2 array of tissue cells in the 1999 paper. In addition, that blot was rotated through 180° from 2001 onwards (so that the Actin background for Thymus cells becomes that of Colon cells, and vice versa, while Testes and Ovary change places, and Spleen with Leucocytes). This is marked with orange boxes. One can only hope that these pictorial labelling issues did not extend into the measurements of Actin from the blots, as used in the quantitative results.
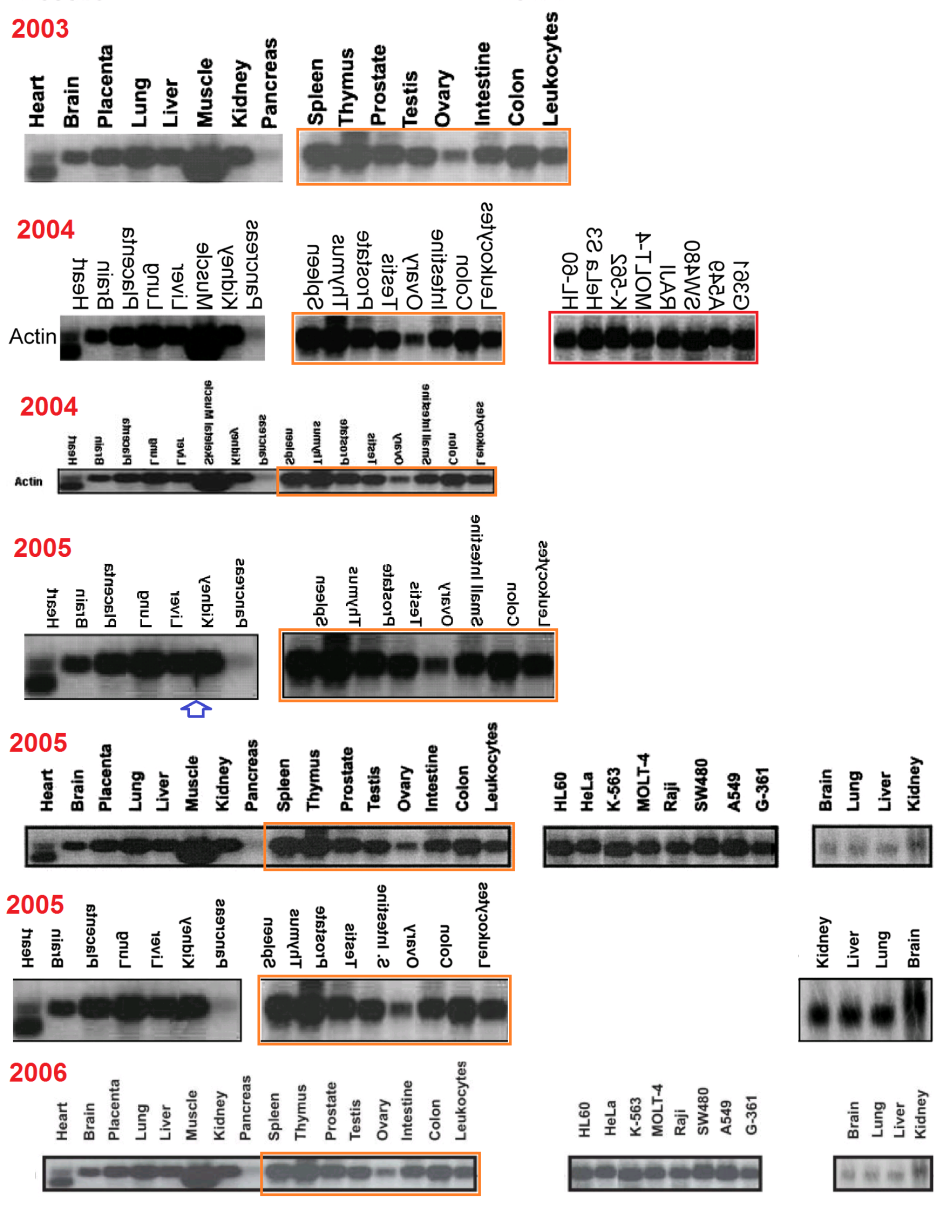
Finally, two blue arrows mark the omission of ‘Pancreas’ and ‘Skeletal muscle’ from one study each, with the loading band spliced to remove that lane.
I am going to play ‘good cop’ here, and propose that the corner-cutting absence of study-specific controls probably made little difference to the results. Corrigenda to the paper acknowledging the use of archival controls would be appropriate (along with correction of any flipped and rotated bands). Other issues have been raised about other figures in some of the papers, but I do not address those here.
Details of the 23 publications follow. We are still hopeful of finding a few more examples of the Perennial Northern Blot.
- 1994. “Human cathepsin O. Molecular cloning from a breast carcinoma, production of the active enzyme in Escherichia coli, and expression analysis in human tissues“, Velasco et al; J Biol Chem., 269(43):27136-42.
- 1995. “Cloning and expression analysis of a novel human serine hydrolase with sequence similarity to prokaryotic enzymes involved in the degradation of aromatic compounds“, Puente & López-Otín; Journal of Biological Chemistry 270, 12926-12932. DOI 10.1074/jbc.270.21.12926 Figure 5.
- 1996. “Cloning and Expression Analysis of Human Bleomycin Hydrolase, a Cysteine Proteinase Involved in Chemotherapy Resistance“, Ferrando et al.; Cancer Research 56: 1746-1750. PMID: 8620487
- 1996. “Molecular Cloning of a Novel Membrane-type Matrix Metalloproteinase from a Human Breast Carcinoma“, Puente et al; Cancer Research 56:944-949.
- 1997. “Identification and characterization of a novel human matrix metalloproteinase with unique structural characteristics, chromosomal location, and tissue distribution“, Pendás et al; J Biol Chem. 272(7):4281-6. doi: 10.1074/jbc.272.7.4281 Figure 7.
- 1998. “Cathepsin L2, a Novel Human Cysteine Proteinase Produced by Breast and Colorectal Carcinomas“, Santamaría et al; Cancer Res. 58(8):1624-30.
- 1998. “Cathepsin Z, a novel human cysteine proteinase with a short propeptide domain and a unique chromosomal location“, Santamaría et al; J Biol Chem. 273(27):16816-23. doi: 10.1074/jbc.273.27.16816 Figure 5.
- 1999. “Cloning and characterization of human MMP-23, a new matrix metalloproteinase predominantly expressed in reproductive tissues and lacking conserved domains in other family members“, Velasco et al; J Biol Chem. 274(8):4570-6. doi: 10.1074/jbc.274.8.4570 Figure 6.
- 1999. “Molecular cloning and structural and functional characterization of human cathepsin F, a new cysteine proteinase of the papain family with a long propeptide domain“, Santamaría et al; J Biol Chem. 274(20):13800-9. doi: 10.1074/jbc.274.20.13800 Figure 6.
- 1999. “Identification and Chromosomal Location of Two Human Genes Encoding Enzymes Potentially Involved in Proteolytic Maturation of Farnesylated Proteins“, Freije et al; Genomics 58, 270–280. DOI: 10.1006/geno.1999.5834
- 2000. “Human MT6-matrix metalloproteinase: identification, progelatinase A activation, and expression in brain tumors“, Velasco et al; Cancer Research 60, 877–882. pubmed: 10706098
- 2001. “Identification, Characterization, and Intracellular Processing of ADAM-TS12, a Novel Human Disintegrin with a Complex Structural Organization Involving Multiple Thrombospondin-1 Repeats“, Cal et al; Journal of Biological Chemistry 276, 17932-17940. doi: 10.1074/jbc.M100534200 Figure 5.
- 2002. “Matriptase-2, a Membrane-bound Mosaic Serine Proteinase Predominantly Expressed in Human Liver and Showing Degrading Activity against Extracellular Matrix Proteins“, Velasco et al; J Biol Chem. 277(40):37637-46. doi: 10.1074/jbc.M203007200 Figure 8.
- 2002 “Cloning, expression analysis, and structural characterization of seven novel human ADAMTSs, a family of metalloproteinases with disintegrin and thrombospondin-1 domains“, Cal et al; Gene 283 49-62. doi: 10.1016/S0378-1119(01)00861-7
- 2003. “Polyserase-I, a human polyprotease with the ability to generate independent serine protease domains from a single translation product“, Cal et al; PNAS 100(16): 9185–9190. doi: 10.1073/pnas.1633392100 Figure 4.
- 2003. “Human Autophagins, a Family of Cysteine Proteinases Potentially Implicated in Cell Degradation by Autophagy“, Mariño et al; Journal of Biological Chemistry 278, 3671-3678. doi: 10.1074/jbc.M208247200 Figure 3.
- 2003. “Identification and Characterization of ADAMTS-20 Defines a Novel Subfamily of Metalloproteinases-Disintegrins with Multiple Thrombospondin-1 Repeats and a Unique GON Domain“, Llamazares et al; Journal of Biological Chemistry 278(15):13382-13389. doi: 10.1074/jbc.M211900200 Figure 4.
- 2004. “Identification and Characterization of Human and Mouse Ovastacin“, Quesada et al; JBC 279 (25) 26627-26634. doi: 10.1074/jbc.M401588200 Figure 3.
- 2004. “Cloning and enzymatic analysis of 22 novel human ubiquitin-specific proteases“, Queseda et al; Biochemical and Biophysical Research Communications 314, 54-62. doi: 10.1016/j.bbrc.2003.12.050
- 2005. “Identification of Human Aminopeptidase O, a Novel Metalloprotease with Structural Similarity to Aminopeptidase B and Leukotriene A4 Hydrolase“, Díaz-Perales et al; Journal of Biological Chemistry 280, 14310-14317. doi: 10.1074/jbc.M413222200 Figure 4.
- 2005. “Human Polyserase-2, a Novel Enzyme with Three Tandem Serine Protease Domains in a Single Polypeptide Chain“, Cal et al; JBC 280, 1953-1961. doi: 10.1074/jbc.M409139200 Figure 3.
- 2005. “Identification and Characterization of Human Archaemetzincin-1 and -2, Two Novel Members of a Family of Metalloproteases Widely Distributed in Archaea“, Diaz-Perales et al; JBC 280(34):30367-30375. doi: 10.1074/jbc.M504533200 Figure 4.
- 2006. “Identification and characterization of human polyserase-3, a novel protein with tandem serine-protease domains in the same polypeptide chain“, Cal et al; BMC Biochemistry. doi: 10.1186/1471-2091-7-9 Figure 7.
Update 7.05.2018. Based on reader comment:


Donate!
If you are interested to support my work, you can leave here a small tip of $5. Or several of small tips, just increase the amount as you like (2x=€10; 5x=€25). Your generous patronage of my journalism will be most appreciated!
€5.00


I am glad that you discuss this case, which has disturbed me since I noticed all the problematic cases at Pubpeer. I am part of a group of scientist who collected all the evidence, did our own analysis and sent the home institution (University of Oviedo) a Powerpoint report describing the numerous problematic cases in detail.
The answer from the Research ethics committee of the University of Ovidedo was surprising:
“After listening to the experts in the field, this Committee has concluded that there are no relevant irregularities in the papers you mentioned and that the research has been performed following the standards established by the different journals.
The University of Oviedo fully support professor Carlos López Otín and his research group and we are very concerned about the potential consequences of publishing this kind of doubts on the internet.”
The letter was signed by José Ramón Obeso, Vice-rector of Research.
When we responded asking for details in the report, especially regarding the Research ethics committee of the University of Ovideo and the “experts in the field” we understood that there is not much transparency at Spanish universities:
“…..The deliberations of the Ethics commitee are confidential.
Anyway, we suggest you to review the Material and Methods sections of the journals your examined, because they explain in detail the procedures used in the experiments and it will clarify most of your concerns. I remind you that this material is also subjected to revision of the journals editors and referees.
Best regards,
Ethics Committee.”
Since the University of Oviedo is a public university I would assume that transparency in this kind of cases is an obligation. I do not know the law here, but at the home page of the university I found some information regarding transparency:
“In accordance with the provisions of Law 19/2013, of December 9, transparency, access to public information and good governance, the Transparency Portal of the University of Oviedo makes available to the university community and the whole of the citizenship, relevant information and general interest about its organization and functioning as a public service of higher education and scientific and technical research, guaranteeing compliance with the obligations of active advertising provided by law.”
Original text:
“De conformidad con lo establecido en la Ley 19/2013, de 9 de diciembre, de transparencia, acceso a la información pública y buen gobierno, el Portal de Transparencia de la Universidad de Oviedo pone a disposición de la comunidad universitaria y del conjunto de la ciudadanía, información relevante y de interés general sobre su organización y funcionamiento como servicio público de educación superior e investigación científica y técnica, garantizando el cumplimiento de las obligaciones de publicidad activa previstas legalmente.”
I will be grateful if someone can provide us with more information regarding transparency laws in Spain.
LikeLike
Here is José Ramón Obeso holding the full Lopez-Otin report: https://twitter.com/uniovi_info/status/915955456182497280

LikeLike
The gentlemen to the left of the image had to resign from his position as Director of a Public Science Foundation last year because of serious suspicions of corruption:
https://elpais.com/elpais/2017/10/04/ciencia/1507137429_162069.html
LikeLike
You can forget about Spanish transparency laws actually guaranteeing transparency. As you have seen any institution systematically withdraws any information it considers “confidential” unless there is a judicial resolution forcing it to do so. Most Spanish people accept a hierarchical relationship with the Administration blindly, Spain is formally a democracy but the dominant mindset is closer to Russia or China – do not challenge power and wait for your turn sheepishly
LikeLike
If uniovi does not take the ppt report, why do not you upload that on youtube? It had worked well in the case of Shigeaki Kato. See the example. https://www.youtube.com/watch?v=on5lmd-pxiU . After this, Kato left science.
LikeLike
Nothern Blots are not only trick for Lopez-Otin. In previous thread, readers pointed out more doubts on WB, immunostaining, and colony countings. https://forbetterscience.com/2017/12/19/nature-rewards-data-manipulation-with-a-mentoring-award/
Each technique reserves independent thread.
LikeLike
But what type of reviewers/ journal editors let this type of things pass? For sure Lopez-Otin wouldn’t publish this alone
LikeLike
They have learned Northern Blot was such an out-dated technique. They jumped to WB, then immunostaining, and subjective colony countings.
Their 2015 paper in NCB put those techniques together. Also, their tricks were put together.
https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18
NF-κB activation impairs somatic cell reprogramming in ageing
Nature Cell Biology (2015) – 14 Comments
doi: 10.1038/ncb3207 issn: 1465-7392 issn: 1476-4679 pubmed: 26214134
Clara Soria-Valles , Fernando G. Osorio , Ana Gutiérrez-Fernández , Alejandro De Los Angeles , Clara Bueno , Pablo Menéndez , José I. Martín-Subero , George Q. Daley , José M. P. Freije , Carlos López-Otín
LikeLike
Pingback: Boletim de Notícias: Comissão da Câmara vota pacote de facilidades para agrotóxicos | Direto da Ciência
Looking forward to seeing how or whether uniovi reacts to the pile of new evidence, 23 papers! and for sure more papers to come. They cannot pretend the ignorance.
LikeLike
Doubts on subjective colony countings on their 2015 paper in NCB paper continues.
NF-κB activation impairs somatic cell reprogramming in ageing
Nature Cell Biology (2015) – 15 Comments
doi: 10.1038/ncb3207 issn: 1465-7392 issn: 1476-4679 pubmed: 26214134
Clara Soria-Valles , Fernando G. Osorio , Ana Gutiérrez-Fernández , Alejandro De Los Angeles , Clara Bueno , Pablo Menéndez , José I. Martín-Subero , George Q. Daley , José M. P. Freije , Carlos López-Otín
LikeLike
Their 2015 NCB paper is still accumulating lots of comments.
NF-κB activation impairs somatic cell reprogramming in ageing
Nature Cell Biology (2015) – 17 Comments
doi: 10.1038/ncb3207 issn: 1465-7392 issn: 1476-4679 pubmed: 26214134
Clara Soria-Valles , Fernando G. Osorio , Ana Gutiérrez-Fernández , Alejandro De Los Angeles , Clara Bueno , Pablo Menéndez , José I. Martín-Subero , George Q. Daley , José M. P. Freije , Carlos López-Otín
https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18
None of their submitted original films of WB were labeled. Only retrospectively annotated for figure preparation, that allows possibility of calling any sample as they wanted. Even though they annotated molecular weight (KDa), I do not see molecular weight markers run on the whole membrane.
LikeLike
More comments on 2015 NCB paper.
NF-κB activation impairs somatic cell reprogramming in ageing
Nature Cell Biology (2015) – 21 Comments
doi: 10.1038/ncb3207 issn: 1465-7392 issn: 1476-4679 pubmed: 26214134
Clara Soria-Valles , Fernando G. Osorio , Ana Gutiérrez-Fernández , Alejandro De Los Angeles , Clara Bueno , Pablo Menéndez , José I. Martín-Subero , George Q. Daley , José M. P. Freije , Carlos López-Otín
https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18#20
“I am following the former comments on the paper, considering arguments around NF-kB that affecting the main message of this study. NF-kB blots (EMSA) in this paper were too much intensely developed and lost its resolution as you can see in raw data. Nevertheless, their main figures showed relatively cleaner blots that can not found in raw image.
Because of NF-kB being main points of this study, I would say that the author should go back to check the original blots.
However, even if it were done, the problem still remains that none of the blots were formally labeled, so they could claim any lane as their ‘actual sample’…”
LikeLike
More comments on 2015 NCB paper.
NF-κB activation impairs somatic cell reprogramming in ageing
Nature Cell Biology (2015) – 23 Comments
doi: 10.1038/ncb3207 issn: 1465-7392 issn: 1476-4679 pubmed: 26214134
Clara Soria-Valles , Fernando G. Osorio , Ana Gutiérrez-Fernández , Alejandro De Los Angeles , Clara Bueno , Pablo Menéndez , José I. Martín-Subero , George Q. Daley , José M. P. Freije , Carlos López-Otín
https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18#23
“The blots of p16 were flipped. The blots of lamin B1 seem to come from different membrane of unrelated samples. It is hard to presume which blots correspond to which sample, because 1) The order of sample differ between each blots even in the same set of experiment; 2) The orginal mambranes were not labeled. I would share the view of Purpuradusta Barbieri that “allows possibility of calling any sample as they wanted”.”
LikeLike
2015 NCB paper (Soria-Valles et al.) still gains more comments on Pubpeer.
Suspicious flipping and jumping bands from different films, and inconsistent colony countings generate more attention.
NF-κB activation impairs somatic cell reprogramming in ageing
Nature Cell Biology (2015) – 30 Comments
doi: 10.1038/ncb3207 issn: 1465-7392 issn: 1476-4679 pubmed: 26214134
Clara Soria-Valles , Fernando G. Osorio , Ana Gutiérrez-Fernández , Alejandro De Los Angeles , Clara Bueno , Pablo Menéndez , José I. Martín-Subero , George Q. Daley , José M. P. Freije , Carlos López-Otín
https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18#30
“Lack of clear and solid standard of colony countings yields inconsistent results and representation.”
“Weird… It is hard to see how they conducted experiments. Did they run all the samples same day to make corresponding raw data to fulfill demand of NCB?”
“It is very concerning that each band were flipped, intensity of signal was artistically modulated, and came from random film. Rubrimonas Cliftonensis pointed the cherry-picking of bands from irrelevant films.”
“Ok, extensive posts on this paper might be reconciled into two major issues.
a. The blots are flipping and jumping aroud different films. And contrasts of bands are too much modified. This issue accounts for most half of the paper (figure 2a, 3c, 4g, 4i, 5f, 6b, 7a, 8b, s1e, s2b, s4c, s6a, s6c, s6e, s8a-e).
b. The criteria of colony counting is unclear and subjective. This issue accounts for another half of the paper (figure 1b, 1d-g, 3c, 3e-g, 5a-b, 5d-f, 6a, 6c-d, 7b-c, s1g, s4a-b, s5b-c, s5e-f, s6b, and s6d).”
LikeLike
New doubts on 2015 NCB paper (Soria-Valles et al.). Where are their beta-actin control bands coming from…?
NF-κB activation impairs somatic cell reprogramming in ageing
Nature Cell Biology (2015) – 32 Comments
doi: 10.1038/ncb3207 issn: 1465-7392 issn: 1476-4679 pubmed: 26214134
Clara Soria-Valles , Fernando G. Osorio , Ana Gutiérrez-Fernández , Alejandro De Los Angeles , Clara Bueno , Pablo Menéndez , José I. Martín-Subero , George Q. Daley , José M. P. Freije , Carlos López-Otín
https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18#32
“The raw data of control beta actin bands are missing.
P-p53, p53 and p16 bands were flipped.
P21 bands seem to come from irrelevant membrane. Their layout and lanes never match with other samples.
There is some weird baked membrane (or black box) on p16 membrane.”
LikeLike
You guys are starting a witch hunt against one of the most honest scientists in Spain (something I would not say of some of the other names you mention in your article, by the way).
There might be some sloppiness (specially in the changing the names from different wells across different articles) but, if you knew López-otin, you would never question his integrity.
As far as I know, that blot was a commercial blot and not a cheap one, so they didn’t blot it over and over against actin to extend the lifespan of the blot itself, and be able to reuse it for longer period of times.
Also, it actually means it was not totally needed to re-blot against the actin probe, since it was the same blot (same thing as in a WB, where you wouldn’t blot against actin every time you do the stripping if you are analyzing several proteins on the same gel), not different blots loaded by “a trained gene-modified laboratory monkey”, but maybe you didn’t care enough to read the methods sections where it states that the blot was purchased from Clontech?
I’m all up for detecting fraud and making science better, but you should be more careful with your claims and your judgements before starting an unfair crusade.
Also, you should keep to yourself your very-much-not-hidden disdain against the Iberian peninsula as a whole. There are definitely rotten apples, as in every other country, but there are no systematic “untouchables” nor anything close to your statement that “we all know what these award-winners did to create those big papers and we don’t mind at all”.
LikeLike
Am I to understand that a single blot was acquired with proteins already gelled and transferred, then repeatedly stained (and stripped) for 23 different proteins?
LikeLike
RNA actually, not proteins. Which is much more unstable than protein (western blot) or DNA (southern blot) or bullshit (eastern blot). I wonder if anyone ever stripped an RNA blot more than once or twice, because you lose lots of bound material with each stripping. After 20 such washes over the years, I woudl expect the blot membrane to disintegrate, not just lose all traces of RNA it ever used to contain. But as we learned, that is a “special”, custom-made magical super-blot.
LikeLike
They use a commercial Northern blot membrane, and if you see the commercial instructions from Clontech you can see that it can be reuse multiple times. However, it is very easy to talk about fraud and image duplication without having an idea of what is a Northern-blot and without having ever perform one.
LikeLike
“There might be some sloppiness (specially in the changing the names from different wells across different articles)”.
That’s O.K. then. What a relief!
LikeLike
No, sloppiness should be corrected in any paper where data is not correctly presented, of course.
All i’m saying is that those are probably honest mistakes, mistakes that can happen to all of us (it did happen to me as well after publication, that we realized a couple of mistakes in one of our figures). Specially when you’re talking about someone with over 400 publications.
As for Leonid’s comment, cause it doesn’t let me reply to you directly, i do have run quite some Northern Blots myself (have you?) and, precisely because stripping is not a good thing for northerns, what you typically do is let the radioactive signal decay by itself, before you probe it against whatever other gene of interest.
I can assure you that, if you do it that way and you keep the blots correctly stored (not totally dried, and in a sealed bag at -20C), the blots can last for a LOONG TIME and can be reused many times.
That’s the exact explanation as to why they wouldn’t strip and reblot against actin, to preserve the lifespan of the blot.
Again, i would be surprised if you yourself had done many (or any) of these northern blots, but still you seem to indicate that you have the higher truth about it…
LikeLike
Thank you for your insights. Now we know how Lopez-Otin lab produced so many papers. Others have to do really boring, pedestrian things, like experiments. But the Lopez-Otin lab had one blot, and used it to make many papers, over many years. The loading control remains same old picture from years ago. And at some point, that blot grew new tissues on it!
Ah well. You are right: I never did Northern Blots, so maybe the practice of data re-use is different in that particular community of Northern blot researchers.
Also I never transplanted a trachea, so maybe I should also shut up there.
I did quite a lot of Western Blots though, but then again never for Nature Cell Biology level, so I better shut up also there.
Thanks again, anonymous expert!
LikeLiked by 1 person
Carolyn is right to point out that radioactive decay is an alternative to stripping.
If this theory is correct, and the same gel was preserved and used repeatedly, what level of agreement should we expect between the shapes and widths of the lanes and blots for the different RNAs it was probed for?
LikeLike
With all due respect, arguments cannot work one way and the oposite, particularly among scientists. The decisions that Lopez-Otin deserved many prizes, many grants and many interviews in mass media were precisely based in the fact that his productivity had been much greater than that of his peers. Many scientists that competed for the same prize, grant or visibility had to accept that it was just fair that everything went to Lopez-Otin. After all, his group had published more than 400 papers! Now you cannot argue that because he has published 400 papers you cannot expect the same standards of quality that one would ask to someone with lower productivity (and more methodical rigour).
Now we know that at least one aspect of that productivity was artificial, because many of us have done many, many, Northern blots, we can testify that publishing 23 times the same loading control is wrong, dead wrong. He could just had referred to the first control in the rest of the papers that followed the first one, clean an easy! What was wrong with admitting that he had not reprobed the original Northern blot for actin? Nothing, but his group would had lost 23 panels and would have received the expected 23 criticisms from the reviewers: you are relying in a membrane that you had stripped 23 times and stored over more than one decade? Give me a break! alent.
The last point is that we only can identify the obvious flaws (panel repetitions or fabrications). How can we know if those are the exceptions or the tip of the iceberg?
LikeLike
And EMBO gave the first author (Soria-Valles) of 2015 NCB paper long-term fellowship… “C.S.V. is an EMBO (ALTF 1240-2015) fellow”.
She is from EMBO member Lopez-Otin lab. Traverse lineage of perennial bands.
@schneiderleonid
“@NatureCellBio asks authors to deposit full uncropped western blots. But @ERC_Research awardee & @EMBO member Carlos Lopez-Otin made a travesty of it. Nothing matches. Does the journal mind? @Magda_Skipper”
LikeLike
Leonid, may worth covering Lopez-Otin’s more updated “sloppiness” in a new post. Pubpeer discussing lots of their new skills; flipping&jumping bands, subjective colony count techniques, and de novo synthesis of actin bands (NEW!). https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18
LikeLike
Salute to Carlos Lopez-Otin’s ground-breaking new technology of de novo synthesis of proteins independent from “perennial” template RNAs – Soria-Valles et al., 2015 Nature Cell Biology.
LikeLike
Uniovi has indeed failed in some transparency index on Spanis universities. Its also typical on spain universities institucional endogamy, wit a lot of dark tins involvin teachers assignations
LikeLiked by 1 person
Readers found that 2015 NCB paper (Soria-Valles et al.) violates data deposition policy of the journal.
NF-κB activation impairs somatic cell reprogramming in ageing
Nature Cell Biology (2015) – 35 Comments
doi: 10.1038/ncb3207 issn: 1465-7392 issn: 1476-4679 pubmed: 26214134
Clara Soria-Valles , Fernando G. Osorio , Ana Gutiérrez-Fernández , Alejandro De Los Angeles , Clara Bueno , Pablo Menéndez , José I. Martín-Subero , George Q. Daley , José M. P. Freije , Carlos López-Otín
https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18#35
“The raw blots were too much modulated to correlate with images in figures. Raw films were so black, and figures were too white! The submitted raw blots (supplementary figure 9) lacks molecular size markers in every single film, and constrast was adjusted too high.
NCB has policy below to raw blots. “Positive and negative controls, as well as molecular size markers, should be included on each gel and blot” “High-contrast gels and blots are discouraged, as overexposure may mask additional bands. ”
And the criteria of the authors do not meet with the policy…
LikeLike
More violation to NCB policy by Soria-Valles et al., 2015 from Lopez-Otin lab.
Not only WB, but also microscopic image processing.
NF-κB activation impairs somatic cell reprogramming in ageing
Nature Cell Biology (2015) – 37 Comments
doi: 10.1038/ncb3207 issn: 1465-7392 issn: 1476-4679 pubmed: 26214134
Clara Soria-Valles , Fernando G. Osorio , Ana Gutiérrez-Fernández , Alejandro De Los Angeles , Clara Bueno , Pablo Menéndez , José I. Martín-Subero , George Q. Daley , José M. P. Freije , Carlos López-Otín
https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18#37
“Another NCB policy regarding quantification of microscopic images was not followed in this paper. It has been unclear and unnamed how the authors processed software of image quantification that occupied half of their data (figure 1b, 1d-g, 3c, 3e-g, 5a-b, 5d-f, 6a, 6c-d, 7b-c, s1g, s4a-b, s5b-c, s5e-f, s6b, and s6d). Tetramerium Zeta pointed that intuitive counting skill ended up similar percentage in two different experiments, but obvioulsy looking very different…
NCB policy; “Processing software should be named and manipulations indicated (such as type of deconvolution, three-dimensional reconstructions, surface and volume rendering, “gamma changes,” filtering, thresholding and projection).”
LikeLike
It is clear now that Lopez-Otin’s magical Northern blotting skill was descended to new generation in a form of elusive bands and phantasmagoric images as we are appreciating in Soria-Valles, 2015 Nature Cell Biology.
Their wonder wand casts a spell of evasion to journal policy.
https://pubpeer.com/publications/836AB3A8AB4FD562A2D4CBFBF8ED18#37
LikeLike