The Portuguese cancer researcher Sonia Melo has been cleared of all suspicions of scientific misconduct by her employer Instituto de Investigação e Inovação em Saúde (I3S) in Porto. She is now re-installed as research group leader, despite of an earlier EMBO investigation which stripped Melo of her start-up funding and the title of EMBO Young Investigator. Previously, PubPeer users raised strong suspicions of data manipulations as well as concerns about irreproducibility and artefactual results based on questionable reagents. The affected publications were authored by Sonia Melo during her stays in the laboratories of Manel Esteller in the Spanish city Barcelona (see my report here) and Raghu Kalluri at MD Anderson in Texas, USA.
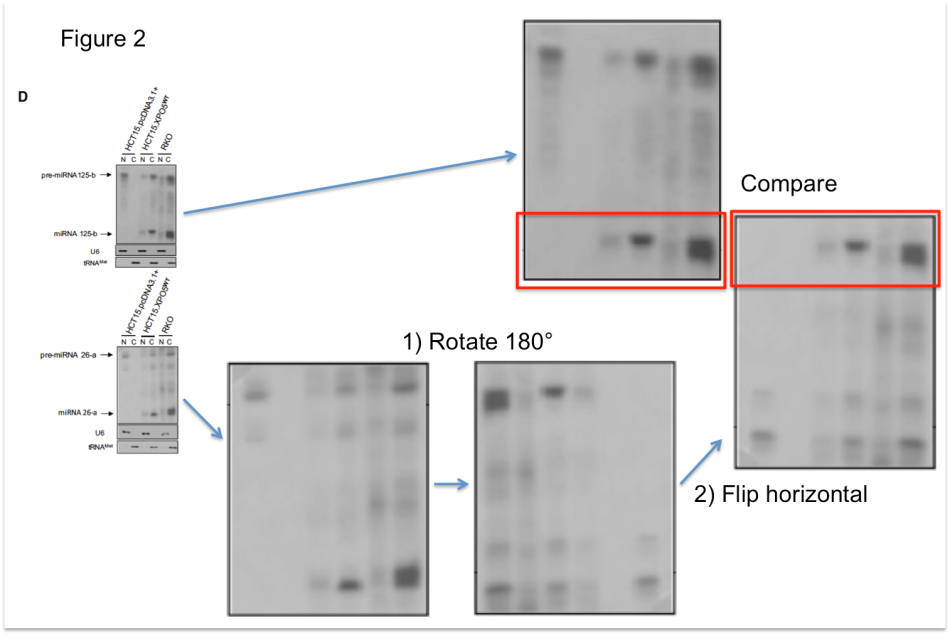
Neither of her former supervisors has been investigated by his respective host institution in connection to PubPeer-posted concerns about their publications with or without Melo. Aside of the EMBO investigation (the findings of which were only made available to Melo’s former and current employers), I3S was the only institution to initiate their own investigation. Unfortunately, its report is not available to the public either. All we now receive is a press release, in which I3S admits to the existence of data manipulations (interpreted as cases of “negligence” which “do not compromise the scientific content”) in 3 of Melo’s papers: the now retracted Melo et al, Nature Genetics, 2009, plus Melo et al, PNAS 2011 and Melo et al, Nature 2015. Both papers will be corrected; the latter was seminal in the fundraising of at least $80 Million for the purpose of developing a commercial cancer diagnostics test. No further Melo publications were investigated, including this one:
The entire letter issued to me today by Júlio Borlido Santos, Coordinator of I3S Communication Unit, is below:
Statement
External Memorandum – File on the Researcher Sónia Melo
i3S would like to announce that researcher Sónia Melo has been encouraged to resume her work as Principal Investigator. This decision follows the fact that Sónia Melo has been acquitted of any suspected fraudulent attitude in the inquiry initiated by i3S regarding her work before integration in this research unit. The analysis of the case was carried out by an independent External Committee, as was publicly announced in March of the current year.
The External Committee has been invited by the i3S Board of Directors (BD) to analyse and issue a report regarding scientific integrity allegations on Sónia Melo’s conduct, following EMBO’s decision to withdraw the Installation Grant that had been awarded to her. As mentioned at the time, Sónia Melo had decided to suspend her work as i3S Principal Investigator with the agreement of i3S’s BD and her group coordinator until the conclusion of the investigation. Meanwhile, Sónia Melo continued her research work under the supervision of her group coordinator.
The External Committee carried out all inquiries independently and included, in its analysis, all the documentation that was available to the i3S BD, as well as the publication record of the researcher Sónia Melo. During the inquiry, Sónia Melo demonstrated an attitude of total cooperation with both the i3S BD and the External Committee. The External Committee’s report formed the basis for the i3S BD decision.
As first author, Sónia Melo assumed her part of the responsibility for the errors underlying the retraction of an article published in Nature Genetics (Nature Genetics 41, 365 – 370, 2009), the publication of a corrigendum in an article published in Nature (Nature 523, 177–182, 2015) and an error in an article published in PNAS (PNAS 108, 4394–4399, 2011), the latter under amendment. These three papers only include work produced before she joined i3S. The External Committee deliberated that the errors identified collide with the need for scientific rigour and that they demonstrate negligence, a fact the researcher was in agreement with. The errors identified are associated with the editing and revision process of A01/00 images included in the referred papers, and do not compromise the scientific content of the papers.
The External Committee, which performed an in-depth analysis of the facts, considered that the errors detected in the three abovementioned articles are not the result of a fraudulent action or attempt to avoid the truth. The i3S BD is pleased that the External Committee has reached this conclusion and hence has encouraged the researcher to resume her functions. Under Sónia Melo’s suggestion, and in accordance with her Group Coordinator, the researcher is resuming her functions this coming Monday, October 31st”.

According to her institutional website, Melo currently receives funding from:
- “Embo Installation Grant” [revoked in February 2016, -LS]
- Exosomes Epigenetic Message to Tumour Microenvironment-understanding how HiC2 regulates breast cancer associated fibroblasts from American Institute for Cancer Research (AIRC)
- “Maratonas da Saúde 2015” from Maratona da Saúde
- “The functional role of exosomes in tumor hetesogeneity and cancer all plasticity.” from Fundação para a Ciência e a Tecnologia (FCT)
- “The role of Exosomes in Tumor Heterogeneity: More than Just Bubbling” from Fundação para a Ciência e a Tecnologia (FCT).
In this regard, it would be surely nothing less but raving madness from I3S to lose such a productive scientist. Question is now: will EMBO re-install Melo’s Young Investigator funding?
Update 30.10.2016. This just in: Maria Leptin, director of EMBO has provided me with this reply regarding EMBO’s cancellation of Melo’s funding:
“Our decision was founded on a critical analysis of extensive files, and we see no reason to reverse it.
The statement we made in March still holds:
This is to confirm that EMBO has withdrawn the installation grant awarded to her. After EMBO had become aware of the allegations against papers authored by her we set up a committee to investigate these allegations. After a thorough analysis of all papers that had formed the basis for her application for the grant, the committee concluded that the body of work upon which the selection for an installation grant was made contained evidence of a level of negligence in handling and presenting data that would have precluded a recommendation for an award. The committee therefore decided that Sonia Melo should not become a member of the EMBO network of Young Investigators and Installation Grantees, and that the installation grant will be revoked. This has been communicated to Sonia Melo and her home institution on February 29.”

If neither Sonia Melo’s ex-supervisors were investigated, it wasn’t fair as well that she carried all the blame alone when perhaps she is the less guilty of the involved persons
LikeLike
I agree that the supervisors should also be investigated, but if the bad practices follow her around, and were not limited to either lab, it’s probably her doing. We must all be accountable, despite the pressures.
LikeLike
Uma desgraça para Portugal, uma tragédia académica, e um insulto ao espirito Lusitano e Camoniano.
LikeLike
Plantarum: the main problem is if we do what our supervisors tell us to do we will get burnt; if we don’t do what they tell us we will get burnt (the second option is my case). Like we say in Portuguese: “Preso por ter cao, e preso por nao ter cao”
LikeLike
If that is indeed the case, then better burnt by doing right, than burnt by doing wrong.
LikeLike
Thank you for the post. There is a scientific bubble full of fake results and unfair awards. We all know that. The question is when it will blow and how far the collateral effects will reach.
I dare to suggest that since we all call ‘biomedicine’ what it should be simply called biochemistry or biology, something has gone wrong (not all fortunately!) in the real science of living organisms (including humans).
I am a scientist from Barcelona, so I feel ashamed of what happened in IDIBELL. This is not how we must proceed. This is really harmfull for many good Catalan scientists and for the whole scientific world.
Thanks again for your post. We will persevere in doing good science, because we like it, allthough nobody knows us and we are fired from labs because of that.
LikeLike
“These three papers only include work produced before she joined i3S.”
OK. But this is the whole point: would she have been given the job at i3S if she hadn’t had those papers?
If not, she should be sacked and the post given to the runner-up applicant (if they no longer want it, it the application process should be reopened.)
LikeLike
As far as I can understand there is nothing wrong about the Nature and the PNAS papers. The Nature paper got corrected, as it happens with dozens of papers every week, and a corrigendum was published. Apparently the PNAS paper too, although I couldn’t find the corresponding corrigendum. Correcting a paper is not fraud or misconduct. Therefore, it looks to me the only issue at stake here is the Nature Genetics paper. Or am I missing something?
LikeLike
“Frank”, I invite you to leave the I3S decree for a second and move to this thread concerning the Nature 2015 paper:
https://pubpeer.com/publications/70714D8ACB8F13164A2752B4335F38#fb43746
as well as to this comment by Ana Pedro, who happens to have been working in exactly same research field:
https://forbetterscience.wordpress.com/2016/03/02/sonia-melo-loses-embo-yip-funding/comment-page-2/#comment-2487
Of course it is perfectly possible that the rest of the scientific community are incompetent idiots with two left hands growing out of their bums, thus unable to reproduce the experiments done by a genius. So far the only lab where the paper was reproducible is that of of Raghu Kalluri at MD Anderson. Or did the I3S commission mention any other successful reproducibility attempts on their premises or did they address the question of the anti-GPC1 antibody at all?
“Frank”, I also invite you to declare potential COI, since your IP address is located to Portugal, specifically Porto.
LikeLike
Dear Mr. Schneider,
I am surprised you don’t care about any COI Ana Pedro, for instance, may have related with this issue. I am not saying she does have one, but there is certainly a difference of treatment here. I looked at Ana Pedro’s post, as you suggested, and would like to know where did Ana Pedro publish her results. Is there any other publication from the rest of the scientific committee failing to reproduce the results? Or are you suggesting that the lack of publications is synonymous to lack of reproducibility?
Yes, I am in Porto and yes, i3S is not indifferent to me. As I suppose it isn’t to any Portuguese citizen. Does that turn me or my opinion less respectable? I am contributing to an important discussion with a perspective which should be considered as valid as that of the other users. I do respect the work you are doing here and hope you don’t selectively accept negative posts only and maintain single-minded a discussion that should be plural.
Thank you for your attention.
LikeLike
Everyone’s comment is welcome, but please note my comment policy regarding anonymous posts. Ana Pedro signed her criticisms by her correct name, unlike you did. In fact, was the choice of “Frank” intended to suggest a non-Iberic origin?
LikeLike
Dear Mr. Schneider,
It took me sometime to look this up. First of all I have to admit I was convinced Ana Pedro was a fake name. Now that you claimed it’s a real name, I looked it up and there is indeed someone called Ana Pedro working with exosomes. As far as I could find out in pubmed, she is the author of a single paper (https://www.ncbi.nlm.nih.gov/pubmed/26910077) about a method to purify exosomes. I hope I did not get the wrong Ana Pedro. I guess she did not publish the GPC1 results, yet. Curious about her I looked up the other co-authors of the paper. David Lyden quickly caught my attention. I found out he is a major author in the exosomes field with many publications in highly ranked journals. Interestingly, he is well represented in pubpeer (https://pubpeer.com/search?q=Lyden&sessionid=C3D73CDAB85985E1C20B&adv=none). Also noteworthy, is that Dr. Lyden is a major competitor of Dr. Kalluri, Sonia Melo’s former boss and senior author of the Nature paper. But maybe I got the wrong Ana Pedro.
My real name is not Frank, although I am not what could be called a classical Iberian. Much to my sadness I don’t think I can reveal my real name. Not yet, at least. I am now getting a glimpse of how this entire process may be so ill-tempered by motivations which are difficult to interpret, that I would put myself in a very dubious situation by revealing my real name. However, with the rare exception of people like Ana Pedro, anonymity seems to be the rule in these posts. Isn’t it?
LikeLike
It is indeed the same Ana Pedro, and I hope she engages into a debate with you. All anonymous comments on my site are checked in regard to their wording for COIs. Trolling did happen before, e.g. by Frontiers, a Macchiarini collaborator and even a research competitor of my ex-boss.
LikeLike
If I understand correctly, the science is correct. The problem stems from the use of wrong images, which Sonia Melo admits she failed to detect. I think we are talking of a different category of problem here. This is not fraud, a deliberate falsification of scientific data, and and I think that should not be misjudged in this case. Justice and righteousness have to work both ways. To punish, but also to correct by educating.
What I don’t understand is why only the most recent employer of Sonia Melo actually did something to clarify the matter. Let’s face it. They certainly are the least prepared part to investigate the problem since none of the work was developed there! I don’t get it…
LikeLike
rank”, I invite you to leave the I3S decree for a second and move to this thread concerning the Nature 2015 paper:
https://pubpeer.com/publications/70714D8ACB8F13164A2752B4335F38#fb43746
as well as to this comment by Ana Pedro, who happens to have been working in exactly same research field:
https://forbetterscience.wordpress.com/2016/03/02/sonia-melo-loses-embo-yip-funding/comment-page-2/#comment-2487
Of course it is perfectly possible that the rest of the scientific community are incompetent idiots with two left hands growing out of their bums, thus unable to reproduce the experiments done by a genius. So far the only lab where the paper was reproducible is that of of Raghu Kalluri at MD Anderson. Or did the I3S commission mention any other successful reproducibility attempts on their premises or did they address the question of the anti-GPC1 antibody at all?
“Frank”, I also invite you to declare potential COI, since your IP address is located to Portugal, specifically Porto.
LikeLike
Dear Mr. Schneider,
I appreciate the publication of my previous post. Meantime, I have been looking into the pubpeer posts as well and I admit the question of the anti-GPC1 antibody is pertinent. Those results certainly need to be reproduced. Regarding your point on the reproduction taking place at the i3S premises, honestly I don’t see how that could happen. That’s why I called the attention in my first post for the fact that the only institution which did an investigation is not the one fully equipped to actually do it! That should be addressed by the institutions where the work took place.
LikeLike
The point remains that (your employer?) I3S declared the Nature 2015 paper as scientifically fully sound, thus fully ignoring the concerns of its suspected irreproducibility and artefactual nature. I wonder why.
LikeLike
Dear Mr. Schneider,
I have no direct relationship whatsoever with the Sonia Melo process. Regarding i3S’s declaration, what I read in the external memorandum you posted is “The External Committee, which performed an in-depth analysis of the facts, considered that the errors detected in the three abovementioned articles are not the result of a fraudulent action or attempt to avoid the truth.” As far as I understand i3S did not make any claims about full soundness of the Nature paper. They made a simple claim about a specific, and meanwhile corrected, error. Yet, your point is one among many others that a single-page memorandum can certainly not elucidate. I am sure you are as eager to get a clearer understanding of what happened here. Why don’t you contact i3S, if you did not do so already, in order to get answers to these questions? Or better still, did you contact Sonia Melo herself to see if she would be available to answer some questions?
LikeLike
Dear Frank,
I do not know if what I have is exactly a “COI” with Dr. Kalluri. I can explain better my situation:
I was a postdoc at the Lyden Lab/ Champalimaud Foundation. Actually, I am the author of the RECI/BIM-ONC/0201/2012 (FCT, Portugal), although I don’t came neither as the RI or a team member. It happens with this project I went to Cornell as a postdoc, and receive some samples of breast cancer patients from CF from Dr. Fatima Cardoso. What happened to me is that I found much time before the publication of Hoshino et al.2015, that my results did not have anything to do to what is described in that paper. Furthermore, mass spec data from Lyden lab itself that I analyzed does not confirm at all anything that was published in Costa-Silva et al, 2015 and in Peinado et al., 2012. Additionally, this same mass spec data does not reproduce at all the findings of Melo/Kalluri GPC1, 2015 paper.
For these reasons I went into a big conflict with Dr. Lyden/CF and they fired me. I would say that I have some sort of a big ‘COI” both with Dr. Kalluri and Dr.Lyden/CF.
If I will be able to publish all of my contradictory findings? Eventually, I will be, I am doing what I can to publish them and to move forward in the scientific field.
I must tell you all of this saddens me deepen, I always thought institutions like IPATIMUP, MD Anderson, Cornell, Champalimaud Foundation were concentrated in developing real good research towards patients and not only interested in publishing faked papers in high impact journals.
If I deserve a second chance? If Sonia Melo deserves a second chance, I think I deserve it as well, at least I am an honest person.
LikeLike
Dear Ana Pedro,
I am very sorry to learn all that you told in your post. And I am very sorry to learn you are facing problems with your scientific career. It’s precisely why I don’t easily board into quick judgements about people. Too often people get disgraced without even the slightest chance of defending themselves. What puzzles me, is that in the situation you described the obvious thing to do would be to get in touch with Sonia Melo and try to work together out why results are not the same. Did you try that?
LikeLike
Oh we all heard many stories about honest Dr. Ana Pedro. Is it true that you accused Lyden and several postdocs of falsifying data and publications and then simultaneously asked to continue to collaborate with them? Is it true that you asked Schneinner to be coauthor on your manuscripts even when he made no contributions to the science? Is it true that for years you still continued to ask Champalimaud to apply to funding with you after all you accuse them of? Did your contract simply run out or can you actually show us proof that you were fired? You know, for the sake of honesty and better science.
LikeLike
Frank, if you bother to actually read the comments on the PubPeer thread as a scientist, rather than someone engaged in a gish gallop, you will see there there are multiple problems with this paper.
I would also note that given the intense effort world-wide on pancreatic cancer, themes remarkable thing is that the wider community has yet to validate the results. Contrast this to the discovery of the association of c-erbB2 with breast cancer prognosis.
First paper 1987, Science
Second paper, from a different group 1987, Oncogene
Third paper, from yet another group 1987, Mol Cell Biol
etc.
It is now 2016 and I have not seen one paper.
The excuse that the experiments are ‘hard’ will not wash, it was hard in 1987, it is hard now. People have tried and failed and for good reason.
So rather than engage in ad hominems on Ana Pedro and gish gallops, why don’t you actually address the scientific questions.
External committee? I am senior enough to know that these are of two types. Type A is appointed to whitewash, type B is appointed to find the truth.
Given that not one query on PubPeer has been addressed properly by the senior author, the committee or yourself, it is fairly simple to determine what type committee was appointed.
LikeLike
Gosh. Going through all the pubpeer comments is no easy task. At the end it’s all about reproducibility. I know not other way of dealing with it execpt for actually doing the experiments and publishing them. And that’s true both for positive or negative results. If someone already did it, an easy task as you suggest, and did not get the same results, I don’t understand why it wasn’t published yet. And I use the same argument you used. Don’t tell me it is hard to publish. Nowadays, with literally hundreds of journals around, publishing is the easiest thing in the world. Hence, until somebody publishes, one has to wait and not assume it’s right or wrong. That’s looking at things like a scientist.
I know nothing about type A or B commissions. Clearly, this one was none of them, as although Sonia Melo was exonerated from fraud, she was clearly made responsible for negligence. Not a label I would like to carry in my CV and certainly not one that will help her carry on with her career. Not clear to me she will be able to survive this label…
LikeLike
Dear Frank,
You don’t understand why negative results are not publish? Publishing is the easiest thing in the world? Really? Are these your best arguments? If it’s on paper it’s true – “That’s looking at things like a scientist”?!? I guess if things were really like you claim the Nature Genetics paper would never been retracted.
LikeLike
Dear Citizen Hugo,
That was not my claim. What I wrote was “until someone publishes one has to wait and not assume it is right or wrong”. I never said the Nature paper is right, but it certainly will not be proven wrong by sterile claims of unpublished, undocumented lack of reproducibility! The Nature Genetics paper had duplicated figures. That’s why it was caught. The Nature paper is waiting to be replicated. A quite different situation. If it’s proven wrong I will be the first to acknowledge it.
And yes, science is about objectivity and drawing conclusions based on hard evidence. It’s not about about pilling sticks and stones to be lit and thrown at the first chance! I am sorry, but reading through many of the comments in pubpeer and here utterly reminds of an inquisitorial process…
LikeLike
Dear Ana Pedro, dear Ferniglab, dear Citizen Hugo,
Coming from a different scientific area is sometimes a nice thing. Maybe that’s why I quickly and easily found evidence that not everything in the Nature paper is perhaps wrong. Take the following paper for instance (https://www.ncbi.nlm.nih.gov/pubmed/?term=Nanomechanical+sandwich+assay+for+multiple+cancer). It’s not a replication of the Nature paper experiments, at all. But, it certainly demonstrates that GPC1 can be detected in cancer-derived exosomes, and that it readily distinguishes cancer-derived from non-cancer-derived exosomes. With great efficiency, according to the authors.
LikeLike
Dear Frank,
I’ m sorry but those were exactly your claims. Although I think that your confidence in the current state of publishing system is heart-warming, I have to tell you: it’s not really easy to publish negative results (there is no real incentive in doing it, quite the opposite really).
In fact, it’s not easy to publish at all! But I guess things must be different in your scientific area. It also explains why you are unaware that systemic perceived lack of reproducibility is a major topic in “biomedicine” nowadays (indeed some authors claim there is a crisis of confidence about the reproducibility of published results), as is the need to reform the publishing system and career evaluation.
And no, you don´t have to publish to dispute published findings (and this is a good thing in my opinion). A good example of this was the controversy surrounding the STAP method for generating pluripotent stem cells. Probably it’s not your scientific area, but this made headlines in generalist media. You can look it up. Curiously, the original paper also included duplicated images and such.
Concerning the paper you mention, “It’s not a replication of the Nature paper experiments, at all.”. Well, I guess it’s self-explanatory. And the problem is not that the Nature paper is or isn’t fully wrong, the problem is that there are disputed evidence in it that raise (legitimate) suspicion on the work as a whole. One thing is that your findings are later proved to be inaccurate or even wrong, the other thing is to use falsified data to prove your hypothesis and publish a paper.
LikeLike
Dear Citizen Hugo,
My claims are in the text I posted. I leave it to the reader to draw its own conclusions about what I did or did not claim.
Do you have any evidence falsified data was used in the Nature paper?
Regarding the publication I provided, could you please comment on the fact that GPC1 was detected in exosomes and distinguishes cancer-derived from non-cancer-derived exosomes? This clearly contradicts the data Ana Pedro claims to have on her own.
LikeLike
If I may interject: with such heavy suspicions of data manipulations raised against Dr. Melo, the first thing any institutional investigation should have done would be asking for her original research data, of every single investigated paper. They are not that old, being from 2011 to 2015. Maybe you as Ipatimup/I3S insider can find out if your institution bothered with finding out if that data actually existed?
LikeLike
Dear Frank,
Yes, this is the point: does Melo’s original data actually exist? and if the answer is yes, does it fits with the publication? My mass spec original data does not show any of that and if you consider that GPC1 could be used to distinguish cancer exosomes from non-cancer exosomes this is a various serious issue because it is not possible to reproduce Melo’s findings. So, for example like mammography screening is done to detect breast cancer, would you submit yourself to an exosomal GPC1 screening to detect pancreatic cancer?
LikeLike
Dear Frank,
Also in relation to your suggestion of contacting Sonia Melo: I challenge Sonia Melo to contact me (anacadavezpedro@hotmail.com) and show me her original mass spec data
LikeLike
Dear Ana,
Believe it or not, but I am not Sonia Melo, and I have no ideia whether she is following this thread…
LikeLike
Dear Ana,
Could you please comment on the paper I just posted? https://www.ncbi.nlm.nih.gov/pubmed/?term=Nanomechanical+sandwich+assay+for+multiple+cancer. As far as I can understand it supports Sónia Melo’s results.
LikeLike
Regarding the publication in Nanoscale:
There are no results showing specificity for the anti-GPC1 antibody they are using. Without any documentation showing antibody specificity in their cell model, the data is in my eyes worthless.
Furthermore the anti-CD63 antibody (Abcam, clone SPM110) they are using is non-existing. This is
quite essential as the anti-CD63 marker is the only one they use to prove that they actually are looking
at exosomes! They include anti-CD24 and -EGFR as markers of exosomes, but these are not the best markers to identify exosomes in general. Detection of CD63, CD81 and CD9 is the best way to analyze exosome samples and see if the isolated vesicles are true exosomes. Abcam has a long list of anti-CD63, some of them showing a single band without smearing. They also fail to mention that these proteins are detected under non-reducing conditions. When you see western blots for anti-CD63 with a nice distinct band without smearing, or when they do not run it under non-reducing conditions, you can assume that the antibody staining is unspecific, as most of the antibodies are.
As long as it is uncertain which anti-CD63 antibody they are using I do not trust this Nanoscale paper.
LikeLike
Dear Morty,
It must be just another fraudulent paper, then. The plot thickens and the evil brotherhood continues to grow!!!! These Canadian authors were probably commissioned to publish a supportive paper to Kalluri’s group…
Forgive me the caustic tone in this post and, believe it or not, I do respect your expert opinion on the paper. Unfortunately, I am not able to discuss the paper with such detail. However, isn’t all this too much of a conspiracy theory?
I have to admit when I challenged Ana Pedro to comment on the paper I was not expecting yet another accusation of manipulated/fake/worthless results…
I better rest my case…
LikeLike
I am confused. the issue is the antibody’s specificity, did I get it right? So what exactly does it prove that yet another lab used the same antibody proven by others to be unspecific? I will now invite this paper’s authors to the discussion.
LikeLike
No reply yet from Dr. Thundat, I now sent a reminder and contacted first author as well. Please bear in mind that the Thundat research group as not a biological one, therefore I am not sure how they can properly test for antibody specificity and exosomal marker expression. nime.eche.ualberta.ca
It seems to me instead they applied their engineering skills to develop a technical assay based on commercial antibodies which they deemed as validated by, well, Melo et al Nature 2015. So what again is this supposed to prove, Frank?
LikeLike
Dear Frank,
Yes, they found GPC1 in breast cancer exosomes, but this does not mean that this is significant. How many times they found GPC1 in cancer exosomes? How many times GPC1 is not present in normal exosomes? If GPC1 is present in both normal and cancer exosomes is there a significant difference in the amount? Moreover, the focus of Melo’s paper in pancreatic cancer and the development of a screening method for pancreatic cancer (PC). With my samples I don’t find GPC1 most of the time in pancreatic cancer exosomes. Nevertheless, contradictory results had always been published and to be sure that GPC1 or other factors may be worth used in PC diagnostic, prognostic or early PC detection many studies with patients samples must be done.
I will do a PPPR of Melo’s paper contrasting with my mass spec results and post it here at Leonid Schneider’s blog in this post (if he agrees) as soon as I can in one or two weeks. Please, be aware.
LikeLike
Thomas Thundat now engaged with me in a brief exchange regarding the paper from his lab which “Frank” presented as supporting Melo et al Nature 2015.
Thundat spoke of another paper “from a different group” he saw, “about real patient samples” (which is not published yet) as well as “some very exciting results with mouse” he was recently shown by his visitors from US, which he then admitted were “preliminary”.
I also asked Thundat about his use of the discontinued GPC1 antibody by Thermo Fisher. To this he relied:
I asked if this means that he now does not see GPC1 as a stand-alone reliable exosomal marker for pancreatic cancer (as postulated by Melo et al in Nature), but received no reply yet.
LikeLike
Dear Mr. Schneider,
What an insider I am! The investigation was carried out by an external independent committee. This information is in the memorandum. Anyway, again that’s a question i3S is likely to answer. Since I joined this discussion I saw you raising quite a number of pertinent questions. Why not seeking for the answers with the right sources?
Even though I am not a senior scientist as some of the users who have been commenting, I do know that original research data, such as lab books and other laboratory material, is the property of the research institution where the work takes place. Former lab members do not carry it along when they move to a new lab… We get back to my first post about who should be doing what.
LikeLiked by 1 person
I asked I3S for the full report. MD Anderson doesn’t talk to me, but this of course doesn’t mean they would treat University of Porto the same way.
Meanwhile Frank, whatever you hear at work in the labs and corridors, can be very useful.
LikeLike
Whatever I learn that is non-speculative, you can be sure I will be happy to share.
LikeLike
“Don’t tell me it is hard to publish. Nowadays, with literally hundreds of journals around, publishing is the easiest thing in the world.” It wasn’t what you said? I was just trying to show you that thinks are not that simple (you know, since you come from a different field and such), particularly with “negative results”. Also, I tried to show you that you don’t need to publish to be able to dispute published findings. Which is true by the way, and a healthy practice in my opinion.
Regarding the paper of find, well, like you said “It’s not a replication of the Nature paper experiments, at all.”. The authors don´t look at all into pancreatic cancer cells as Ana did and the relation to pancreatic cancer is a major point here. To my knowledge, no one is saying that GPC1 couldn’t possibly be a cancer biomarker, they’re just saying that some of the evidence regarding the particular findings in Sonia’s Melo paper raises (legitimate) concern. If indeed there are problems with the disputed results, it’s the paper as a whole that is at stake. If only a part of it is true, well, they should have published only that part, don’t you think?
Since confidence is the building bloc in scientific system, I feel that at this point is up to the authors to further clarify their findings.
I think everyone realizes that we have to be careful in addressing these issues, and that there is a very real danger of harming peoples careers. But in this case this researcher (and her PI’s) had every chance of explaining and clarifying the issues raised (which again are legitimate) and they didn’t. Not in any convincing way in my opinion. And fact that the results of this recent investigation were not made public just adds more suspicion to the whole case. I guess one can feel that things in social networks can and often do turn rather inquisitorial (although I don’t’ think this is the case), but I find it much more worrying that issues regarding research practices and scientific misconduct are dealt with in such opaque way.
LikeLike
Dear Citizen Hugo,
Don’t worry about the Dear Frank thing! Overall, I fully agree with you. I wish the scientific publishing system were less opaque. And it’s not limited to fraudulent papers. I wish all authors would be treated equally and have the same access to publishing on top ranked journals, for instance. That said, it has been a useful system in helping humanity come all the way from stone-age up to where we are now. Even though some people act like if they didn’t realize the stone-age is well beyond (couldn’t resist the joke ;-)) It’s also certainly a system that is abused by some and can certainly be improved. Count me in for that positive and progressive discussion.
About publishing negative results. You surely can’t publish them in the same Journals you publish positive results. But if you browse through some of the low impact factor Journals you will see they contain many reports on negative results which, by the way, are equally important to publish. What one can’t do is to jump from the absence of reports to a straightforward accusation of fraud! And this is what is happening with the Nature paper as far as I can tell. The example of quick reproduction of c-erbB2 results provided in a previous post, is most likely the exception rather than the rule. And yes, the paper I provided does not include pancreas, but it includes breast, which Ana Pedro included in her analysis. Ana Pedro claims you can’t detect GPC1 in exosomes from breast cancer. These authors confirm you can.
In this particular case, which I am following now with some passion (I don’t deny it), I see too many loose allegations and very few tangible facts. Facts are the ones related with the retraction of the Nature Genetics paper. Duplicated and rotated images, which were claimed to result from accidental/lousy/whatever handling of the images. I can’t judge beyond that official conclusion, but I would have loved to see the results of an investigation conducted by the Institution where the work took place. The original report from i3S? Most likely a laconic piece of text… But as far as I could find this seems to be standard procedure in Academia.
From that point on everything is very shaky. Allegations and more allegations without any tangible facts. For instance, I just started looking at some of the allegations in pubpeer related with one of Melo’s Cancer Cell papers. For many of the allegations one can’t simply decide, because one would need access to the original blots and other material. But some other claims are straightforward rubbish!
That said, I pick on your comment about people’s careers. It’s a bit like the death penalty; you can always kill a guilty person, but you can’t bring an innocent person back to life. I may be dramatizing, I know.
LikeLike
Absence of evidence is not evidence of absence. We have apparently two facts: nobody published any paper either supporting nor refuting Melo et al 2015.
Of course it is possible that many labs worldwide hide their confirmative results for reasons unclear, that would be most unfortunate for Dr. Melo and Prof Kalluri.
But why would anyone hold back contradictory results and avoid publishing them?
Well, “Frank”, I am not sure about your professional status (you know I have my theories about your identity), would you dare to go publicly in your common narrow research field against a key MD Anderson researcher, who started a business together with Eric Lander (head of Broad institute) himself, in order to monetise that very Nature paper? You are even commenting anonymously here!
LikeLike
Dear Mr. Schneider,
I feel we are already getting in circles here. Yet, I will one last time address your comments. One by one.
– No papers have been published. Maybe there was not enough time for that? To me, a cohort of individuals diagnosed with pancreatic cancer, a cohort of individuals diagnosed with premalignant pancreatic disease and a cohort of healthy people, with suitable blood samples available for exosome isolation and GPC1 detection, is not a common asset. Certainly not available to many research groups. On top of that, validation in this case may mean a prospective study, which means many months, probably one or two years of follow-up. Anyway, you insist in ignoring the paper I mentioned in a previous post. It confirms the Nature paper results that GPC1 can be detected in exosomes from breast cancer, and that GPC1 presence in exosomes can distinguish between cancer-derived and non-cancer-derived samples.
– Going publicly. I know nothing about the business you mentioned in your post, but going publicly against anything is something you do on solid grounds, not based on speculation and hearsay arguments.
– I am not revealing my identity. Is that important? Is the validity of arguments posted in this thread dependent on the identity of who posts? That would be an enormous surprise to me!
LikeLike
Frank,
I don’t claim I cannot detect GPC1 in breast cancer exosomes, I claim I cannot detect GPC1 in pancreatic cancer exosomes. But wait for my fully analysis.
LikeLike
Sorry, I should have begin with “Dear Frank”
LikeLike
Just to be sure that my post will be seen (I posted it first as a reply to Frank, hidden in the middle of the comments):
Regarding the publication in Nanoscale:
There are no results showing specificity for the anti-GPC1 antibody they are using. Without any documentation showing antibody specificity in their cell model, the data is in my eyes worthless.
Furthermore the anti-CD63 antibody (Abcam, clone SPM110) they are using is non-existing. This is
quite essential as the anti-CD63 marker is the only one they use to prove that they actually are looking
at exosomes! They include anti-CD24 and -EGFR as markers of exosomes, but these are not the best markers to identify exosomes in general. Detection of CD63, CD81 and CD9 is the best way to analyze exosome samples and see if the isolated vesicles are true exosomes. Abcam has a long list of anti-CD63, some of them showing a single band without smearing. They also fail to mention that these proteins are detected under non-reducing conditions. When you see western blots for anti-CD63 with a nice distinct band without smearing, or when they do not run it under non-reducing conditions, you can assume that the antibody staining is unspecific, as most of the antibodies are.
As long as it is uncertain which anti-CD63 antibody they are using I do not trust this Nanoscale paper.
LikeLiked by 1 person
Dear Frank.
I am not talking about a conspiracy theory, just the fact that most life science publications apparently are not trustworthy of several reasons. One of them are use of unspecific antibodies, which is a serious problem. That’s a fact. When labs have put serious effort into reproducing data published in high impact life science journals, most of the data are not reproducible. When Amgen checked almost 60 landmark cancer studies, they were able to reproduce the main findings in only 11% of the papers. I know they even went to the individual labs and observed the other scientist repeating the experiments in their home lab, usually without success. As far as I know non of the papers have been corrected or retracted.
There is no conspiracy my friend, only a lot of really bad science out there published in the very best journals.
LikeLike
Dear Frank,
Unfortunately, Morty is pretty right: there is a lot of really not reproducible bad science out there published in the very best journals. Probably, one of the major problems is the focus: the starting should always be patients and patients samples.
LikeLike
Hello everybody,
Here is another paper suggesting GPC1 has prognostic value in pancreatic cancer. It was published even before the Nature paper: https://www.ncbi.nlm.nih.gov/pubmed/23270819.
As far as I can understand all three papers use different antibodies against GPC1, but it must be all rubbish..
And here is yet another paper, suggesting clinical value for the expression of GPC1 in esophagus cancer: https://www.ncbi.nlm.nih.gov/pubmed/27310703. Again a different antibody for GPC1. And don’t tell me it’s a different organ. Antibodies are not organ-specific.
You know what I think? It’s probably time you people start proving published results are wrong and fake, instead of simply questioning the validity of what others did. Why don’t you go to your Labs and run the experiments to prove things are wrong? Wouldn’t that be the scientific way?
LikeLike
Frank, you should be ashamed of your behaviour. Especially since you are a scientist and apparently a rather senior one at I3S. The main finding of Melo et al 2015 was not the existence of GPC1 as such but the expression of GPC1 specifically on cancer-derived exosomes. First paper you offer as supporting never once mentions the word “exosomes”. Second one does, but only in the end of discussion when citing Melo et al 2015 Nature paper!
So what you just did Frank, are the classically muddying the water and diversion tactics used in science to squash any attempt of critical discussion. In fact, it’s nothing but trolling. It tells a lot about your own research and teaching style, I am afraid.
LikeLike
Mr. Schneider,
The first paper I posted is about exosomes. Did you care reading it?
Then the discussion diverted to the antibodies used in the papers. Morty brought the antibody issue up. Not me. The last papers I posted are precisely about the antibodies because that is what was at stake now.
I would also appreciate if you would refrain from making comments about me as a person or as a professional. That certainly doesn’t suit well the reputation of independence and professionalism as a Journalist you are seeking to build.
Finally, I insist on actions instead of talk. People should do the experiments. That’s the only way of moving science forward.
LikeLike
Dear “Frank”,
as I pointed out above, the first paper you offered as support for Melo et al, Nature 2015 was not a biological, but an engineering one. It does make a difference, even if not to you. Quote from their paper:
“We targeted a number of biological markers including, CD24, CD63, EGFR and Glypican-1 (GPC1), that are believed to be over-expressed on surface-membrane of exosomes originated from cancer cells”.
They do not test this belief (they also can’t, being field outsiders), but accept it on the basis of Melo’s and Nature’s authority.
This makes it three papers you ferreted out to prove that your I3S colleague’s work was reproducible. Neither of them actually does so, and as a cancer researcher you should have been able to gather that yourself. So with all due respect, what you were doing was either incompetent or malicious. Never mind my reputation here, it is yours which is at stake. Or is this why you are hiding your identity?
Finally, don’t tell others what to do. If you are so keen, use your own big cancer research lab at I3S and Dr. Melo’s advice to reproduce her discoveries.
LikeLike
Dear All,
I think it’s now time I leave this discussion. It was already getting in circles, and now just turned into a direction and style I am not available for. I appreciate some of the arguments we exchanged and acknowledge the pertinency of some of the questions that were made. It was also a learning experience to me. I believe we are all interested in the truth, but I think one can only address it with independence. Approaching such issues with convictions or, worse, with prejudice, will not get people closer to truth. I leave you my email address, and I will be happy to continue tot alk about science by email: kaposi174@gmx.com.
Best regards,
Frank
LikeLike
There is no need to be so sarcastic when you discuss science, folks!
I am really frustrated over the reproducible problem in life science today, and I am very skeptic to the Melo papers due to what seems to be sloppiness and carelessness, in addition to accusations of manipulation of data. However I do not attack people.
LikeLike
Another prize money for Esteller!!!.
He gets the money, his PhD students get fired… It looks a good deal.
http://www.idibell.cat/modul/news/en/941/manel-esteller-josep-baselga-and-joan-massague-receive-the-xxviii-catalonia-international-prize-from-president-puigdemont
LikeLike