My earlier article about strange image irregularities in the publications of the German mitochondria researcher Roland Lill seem to have motivated this pre-emeritus biochemistry professor of the University of Marburg to come to PubPeer and address the issues. While in his earlier statements he simply waved off all concerns of western blot band duplications, this time and with other papers he admitted those, while presenting the original Western Blot scans. Together with the first author on two such papers, Janneke Balk, Lill explained why copy-pasting western blot bands, sometimes on top of other gel images, had nothing at all sinister in it, but used to be somewhat of a normal research practice 10-15 years ago. And in some cases, gel bands can naturally duplicate themselves.
Any advice on research integrity from the side of a Senator of the German Research Foundation (DFG) and of the German Academy of Sciences Leopoldina is certainly most valuable, this is why I will present here his explanations, so the younger generation of scientists can learn about correct figure preparation, including the proper use of gel band copy-paste function. The past evidence was forwarded to me by a reader of my site. I will also offer Professor Lill and you for debate another example from his past publication, where a western blot was duplicated in different context. This was forwarded me from yet another reader of my site.
Another duplication
This recently spotted duplication appeared in this paper
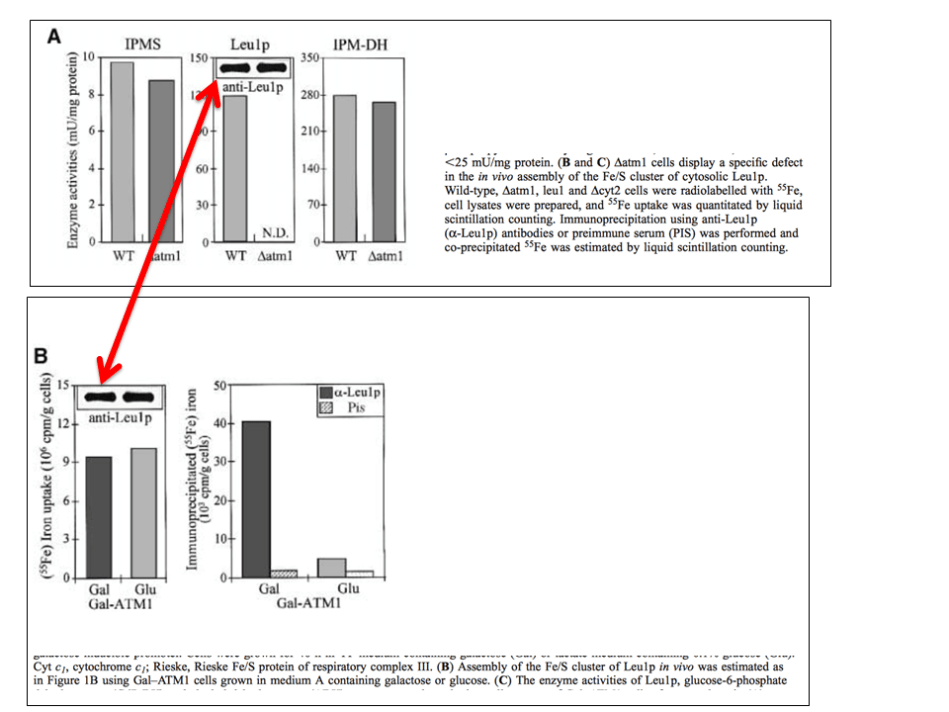
Prof Lill will sure soon explain how it happened. For some reason however, he is still not interested to discuss the issues of apparent band duplications in his publications with Heike Lange, who is now a tenured researcher at the CNRS Institut de Biologie Moléculaire des Plantes (IBMP) in Strasbourg, France (read more about Lange here). This is because he examined one case (Lange et al EMBO Reports 2000) at “larger magnification” and decreed on PubPeer that those bands are not duplicated. Case closed, no need to dig up original gel scans, as he did for some other publications. Not even a darker version of the same figure, to see some background and its integrity. Move along everyone, nothing to see here. Heike Lange’s papers are off limits.
Lill Lesson for Young Scientists: A diligent scientist uses the tool of contrast overexposure to remove all background traces on a western blot, to avoid unnecessary accusations where bands appear suspiciously too similar.

It’s OK to copy-paste
After my article appeared, Lill and his other former PhD student Janneke Balk moved to PubPeer to explain an apparent duplication in their common paper Balk et al MCB 2005. It turned out that the bands were duplicated and stitched together on purpose, with a scientific rationale in mind, because the authors decided to use various transgenic yeast strain samples as wildtype controls of each other. In the paper, they however declared that the genetically unmodified strain W303-1A was used as wildtype throughout (see my comments here). There is a good reason to use clearly defined widltype controls, to avoid artefacts and to ensure reproducibility. Misrepresenting this information is somewhat unconventional, just as copy-pasting gel bands to save on lab work is. Balk, seconded by Lill, declared:
“To minimize the number of yeast cell cultures, yeast cells overexpressing CIA1 (from the GAL promoter, indicated by ) doubled up as cells with wild-type (wt) levels of NBP35 expression, and vice versa. This is why the duplicated anti-HA bands appear cross-wise in the figure. […]
Looking back at the results after so many years and now with knowledge of the helpful guidance on digital image manipulation from Rossner & Yamada 2005 (J Cell Biol. 166, 1 – 11), the ‘wt’ signals in C should have been separated from the ‘up’ lanes. Or I should have loaded the samples twice on the gel, in anticipation of the figure layout”.
Thing is, Balk and Lill knew how to present separate gel lanes properly already then. They made a clear separation between two panels “cell extract” and “a-HA” of the same figure, but not between the duplicated bands, which were supposed to look as if they were run next to each other, and of course not copy-pasted. They also made the upper and lower gels of each panel seem like they belong to the same physical protein gel, which they did not.
Lill Lesson for Young Scientists: When you have better things to do with your time, you are allowed to short-cut on experimenting and simply reuse gel bands to produce new gel images. One single experiment can be used to generate many different figures, which are just as good as actual experiments.
Anything goes
Balk now works as research group leader at the John Innes Centre in Norwich, UK. Its director Dale Sanders was very impressed by her creative way to generate figures, and declared the matter closed. I would love to know what Sanders made of Balk’s next explanation, for hers and Lill’s paper Balk et al EMBO J 2004, but sadly he decided to call me an “internet troll” whose writings are not worth any of his time because they are not “peer reviewed”.
The surprise was: Balk and Lill admitted on PubPeer to have digitally excised bands and stuck them one by one on top of another gel image, even twice, an image which was already used in the same figure. This is why same background appears in the figure three times: once with its original gel bands (Aac2p), and twice with the gel bands superimposed on top of it (Sec61p and Pol30p). Even Balk and Lill could not explain they did this, as Balk wrote:
“The way the Sec61p and Pol30p lower panels were put together is indeed not standard practice. I don’t know why it was done like that”.
It seems not were bothered, because the authors provided their original autoradiography gel films from the labels on which it looks like the bands were indeed what they were claimed to be.

Balk explained on PubPeer:
“For the figure, the immune signals were rearranged to match the blot for Nar1, Pgk1 and Aac2, by digitally cutting out the bands, and pasting these on top of the panel for Aac2. While our presentation did not raise any concerns at the time, we emphasise that all conclusions from the paper are verified once more based on the original data”.
Indeed, if the peer reviewers didn’t mind (or simply didn’t notice, which probably amounts to same), it is not our place to point fingers. An anonymous expert “Hoya Guppyi” declared to have been “asked to review figure 4B of Balk et al” (asked by whom? PubPeer? University of Marburg? Dale Sanders? Prof Lill? All of them?) and attested:
“In my opinion, although the way the figure was arranged did not follow best practice, especially by today’s standards, there is no evidence that the authors intended to mislead readers or to misrepresent the results they obtained in these western blot experiments”.
I noticed however that the handwritten labels on the original western blot autoradiography film appear to have been made in two different hand writings, detectable by the way the number 1 or the lower-case letters p and l appear in horizontal and vertical labellings. Balk however immediately denied this.

Such films are X-ray films, after acquiring an autoradiography image, the film is developed and labelled with a pen, a permanent marker. This is why one can fully exclude any manipulation of X-ray films labels in the Lill lab, because such pen labelling can only be removed without a trace with an alcohol solution, before a new one can be scribbled. Which is totally out of question, both technically and ethically.
Lill Lesson for Young Scientists: Even the most outrageous figure manipulation in a published paper most likely has an utterly innocent explanation. After all, if it didn’t raise any concerns with peer reviewers, any latter criticism thereof is utterly inappropriate. It is the aggressive calling out of Photoshop-enthusiastic scientists which is the real act of research misconduct.
Self-duplicating bands
According to Lill, there are however occasions where Western blot bands do get duplicated, all by themselves. A set of bands in the paper Csere et al FEBS Letters 1998 looked very similar in two panels of a figure.

In this regard, Lill explained on PubPeer (highlights mine):
“The suspicion is that the two left-most lanes of panels A and B have been duplicated. There are two possible interpretation scenarios which both would technically be absolutely OK. Decision for one of the two scenarios would need a look at the original autoradiogram(s). They are no more available (the last author Dr. Kispal died in a tragic car accident in 2003). Note that in the days of the experiment (19 years ago) the figures were produced using thermoprinter printouts that were glued into Powerpoint drawings. Scenario 1. Both parts of the experiments in A and B were run on the same gel side-by-side with the two leftmost lanes as the shared standard lanes (-/+ PK) (frequently done for such import experiments). For didactic presentation of the two experimental parts, the respective right parts of A and B were fused to the two leftmost lanes. What does not really fit to this view is the lack of a visible cut between the two left lanes (= standard import) and the respective right parts. With the poor image processing techniques available at the time, such cuts should be easily visible. Scenario 2. Both parts represent individual gel runs. If the same person ran similar samples with the same mitochondria preparation and preproteins on the same gel apparatus, high similarity of band shapes are usual in these types of experiments (which represented a routine experiment for the lab in those days). This interpretation is supported by the fact that the lower parts of the two leftmost bands in A and B are clearly not identical. As stated above, both scenarios for interpreting this experiment are technically absolutely OK”.
I never heard of anything like this ever happening outside Prof Lill’s Marburg lab. The whole issue of gel band shape similarities is exactly because this basically never happens by chance. This is how people generally spot irregularities and data manipulations, because usually gel bands never duplicate themselves, though of course statistically everything is possible (nuclear missiles transformed into a sperm whale and bowl of petunias, for example). Yet Lill certainly spoke out of his many decades of lab experience when he said “high similarity of band shapes are usual”. It is all very strange.
I tried here to enhance the signal of the low resolution figure, and hope Prof Lill will find this useful for tracking down the cause of band similarities.

Lill Lesson for Young Scientists: Sometimes gel bands duplicate themselves, when an a gel run is repeated. To avoid being falsely accused of data manipulation, never run similar samples on the same gel apparatus. Ideally, ask a colleague to run the gel for you, in a different lab.
Almost the same
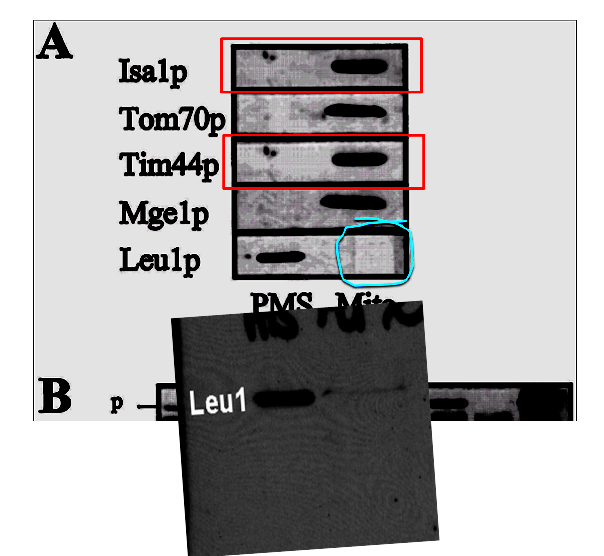
In fact, in yet another case Prof Lill was possibly confused by these recurrent band similarities in his lab when similar samples are loaded on different gels. A Western blot panel in Kaut et al JBC 2000 contained a mysterious box, highlighted in blue. There was also a duplication of a panel, labelled in red.

Lill agreed that the panels Isa1p and Tim44p are the same, and provided what he described as original gel scan for Leu1p, proving that the originally submitted figure contains no “square” feature (highlights mine):
“This experiment aimed at identifying the subcellular localization of the yeast Isa1 protein, and found the protein in mitochondria. The result has been reproduced by many labs for various organisms (see, e.g., Jensen & Culotta 2000 MCB 20:3918-27; Kim et al. 2010 BBRC 392:467-72; Long et al. 2011 Mol Microbiol. 81:1403-18; Sheftel et al., 2012 MCB 23:1157-66). The claim here is that the bands of Isa1 and Tim44 are duplicated which seems to be the case. I checked the accepted version of the manuscript (available on the JBC website http://www.jbc.org/content/early/2000/03/15/jbc.M909502199) [dead link, JBC message “content not found”, -LS/ correction: reader pointed to a functional link, -LS]. It contains two clearly distinct (and correct; see below) bands for these two proteins. Why a different version than the accepted one was published by JBC is unknown to me. I have contacted JBC to request a correction. Original data (scans) found on an old CD shows Tom70, Tim44, CC1HL, Mge1 (simultaneously stained on a single Western blot) in the mitochondria (right) fraction, another blot shows Isa1 in the mitochondria fraction, while Leu1 is in the PMS (post-mitochondrial supernatant; left). All bands match the accepted version of the figure. The Leu1 part does not show the criticized “square”. Even if you look carefully in the published version (poor image quality), it is not a perfect square. Thus, this irregularity may arise from a reproduction problem”.
However, while Lill’s freshly provided original scans for other bands seem to match, that for Leu1 does not. Here I made a comparison, the original highlighted figure and the Leu1 gel autoradiography Lill provided as correct original. The bands are distinctly differently shaped, and what happened to the weak second one? What if the stored autoradiography film somehow deteriorated over time, while the published figure preserved the original presence of the bands?

Lill Lesson for Young Scientists: An autoradiography can change its appearance over time. Your bands might not look the same after a decade of storage. Also, do not trust journals, they are known to occasionally sabotage the data integrity of papers they publish, by modifying figures in secret, so that duplications and artefacts of alleged manipulation suddenly appear.

Donate!
If you are interested to support my work, you can leave here a small tip of $5. Or several of small tips, just increase the amount as you like (2x=€10; 5x=€25). Your generous patronage of my journalism will be most appreciated!
€5.00






Interesting read. 👧
LikeLike
The link to the accepted version of the jbc manuscrpt is ok.
http://www.jbc.org/content/early/2000/03/15/jbc.M909502199
Just took away the parentheses.
Nice piece
LikeLike
I agree that WB results can be copied and pasted and that sometimes bands get be naturally duplicated
Nevertheless I think always the original WB’s should be shown and as well the criteria used for the copy and paste should be fully explained as the WB is naturally a tricky technique with many factors influencing the final outcome
LikeLike
I disagree with your first sentence. It is not ok in my opinion.
Full WB scans should always accompany a publication. Several journals request these data for submission.
WB is tricky sometimes. But it is too easy to copy and paste bands. It takes more time but correcting the experimental setup is the only way to go.
Regards, Christian
LikeLike
Perhaps you are right. Maybe it should be put in the publication a figure of one full WB as a representative one, submit the remaining WBs as supplementary data and mark in the WB figure the bands that were selected and mention why
LikeLike
Dear Leonid
You didn’t by chance miss the /sarc tag on this post?
Cheers, Oliver
LikeLiked by 1 person
I find this post absolutely OK.
LikeLike
J Biol Chem. 2011 Dec 2;286(48):41205-16. doi: 10.1074/jbc.M111.296152. Epub 2011 Oct 10.
Specialized function of yeast Isa1 and Isa2 proteins in the maturation of mitochondrial [4Fe-4S] proteins.
Mühlenhoff U1, Richter N, Pines O, Pierik AJ, Lill R.
Author information
1
Institut für Zytobiologie und Zytopathologie, Philipps-Universität Marburg, Robert-Koch Strasse 6, 35032 Marburg, Germany.
2017 correction for figures 2E and 7B.
http://www.jbc.org/content/292/43/17979.short
LikeLike
J Biol Chem. 2000 May 26;275(21):15955-61.
Isa1p is a component of the mitochondrial machinery for maturation of cellular iron-sulfur proteins and requires conserved cysteine residues for function.
Kaut A1, Lange H, Diekert K, Kispal G, Lill R.
Author information
1
Institut für Zytobiologie und Zytopathologie der Philipps-Universität Marburg, Robert-Koch-Strasse 5, 35033 Marburg, Germany.
2017 correction for figures 2A and 3B.
http://www.jbc.org/content/292/43/17980.short
LikeLike
Pingback: Oncogene EiC Justin Stebbing, a hypocrite of research integrity? – For Better Science
Pingback: Salk Gandalf Tony Hunter get AACR prize for magic western blots – For Better Science
Pingback: Lill space-time-blot anomaly in Marburg – For Better Science
Pingback: DFG and Marburg drop misconduct investigation of Roland Lill papers – For Better Science
Pingback: Leopoldina and Tiwari’s scamferences, or what’s the point of Academies – For Better Science
Pingback: Bad Choices in Dresden III – For Better Science