The Stanford professor John Ioannidis is one of the most cited scientists of all times, the godfather of meta-analysis, and the great man who proved “Why Most Public Research Findings are False”” and “Why Almost Everything You Hear About Medicine Is Wrong”. Which surely must include his own light-hearted opinions on COVID-19, but this is another story.
Well not quite, because the main reason why covidiots all over the world worshipped Ioannidis’ misguided assessment of COVID-19 was his immense citation index. Which apparently is, presumably to no fault of Ioannidis’, partially due to papermill citations. And it was his achievement of making meta-analyses cool which prompted the Chinese papermills to set up their business, with Ioannidis in turn soon decrying the “unnecessary, misleading, and conflicted” meta-analyses these mills smuggled into allegedly peer reviewed journals.
Ioannidis, apparently unaware where many of his citations come from, describes himself as:
“Highly Cited Researcher according to Clarivate in both Clinical Medicine and in Social Sciences. Citation indices: h=226, m=8 (Google Scholar). Current citation rate: >6,500 new citations per month (among the 10 scientists worldwide who are currently the most commonly cited).”
Yet Smut Clyde managed to somehow connect the excessive citations some Ioannidis’ papers received to papermills, the references inserted arbitrarily. And Smut even discovered a certain Dr Liu, who may have evolved from a humble oncologist at the China Medical University in Shenyang to the owner of a flourishing papermill business, his services progressing from relatively cheap phony meta-analyses to highly profitable fake experimental studies.

Ghostwriter Stories of an Antiquary *
By Smut Clyde
* Apologies to M.R. James. James’ scholarly protagonists were always encountering eldritch horrors in the course of their pursuit of incunabula, mezzotints, monkish decretals, and runic or uncial manuscripts, so they would not be at all surprised at the sort of thing that goes on in modern academic publishing.
Zintzaras & Ioannidis (2005a, 2005b) addressed the genomic research problem of merging chromosomal gene-locus maps from multiple research projects of varying reliability. To this end they devised nonparametric heterogeneity tests, of the general bootstrap / jack-knife style, to avoid the danger of mixing non-apples in with the apples.
- HEGESMA: genome search meta-analysis and heterogeneity testing. Bioinformatics, 2005a
- Heterogeneity testing in meta-analysis of genome searches. Genet Epidemiol. 2005b
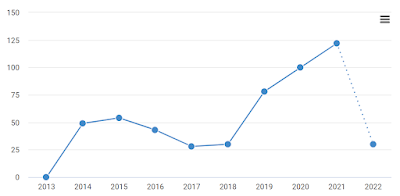
Though narrow of niche, these are solid papers. Yet one can wonder how they came to be cited 409 and 613 times respectively: disturbed from peaceful obscurity by a 2013-2015 spike in citations, before slowly sinking back towards the baseline.
For later reference, it is worth listing some of the topics that Zantzaras & Ioannidis did not touch upon. Publication bias and funnel plots; fixed and random-effect models; the I² test; Cochran’s Q-statistic; Raindrops on roses and whiskers on kittens… So what to make of the extensive genre of studies that cite 2005a or 2005b, or both, as the most relevant authorities on precisely these topics? Is it possible that through no fault of Zintzaras & Ioannidis, their work was incorporated into a papermill template, accruing hundreds of spurious citations?
Let me provide some much-needed context. If, back in the heydays of European empires, you were a tourist or a novice archaeologist in Egypt, you might employ a dragoman to ease your passage through the alien culture, and to handle all the necessary but unaccustomed formalities and transactions: haggling in bazaars, disbursing baksheesh, dispersing beggars and second-hand scarab salesmen, camel insurance, Anubis authentication, translating curses written on a tomb wall in Second-Kingdom hieratic. I learned this from a thorough immersion in Agatha Christie detection novels.
 Academic publishing has a comparable vocational niche. If your career needs a paper in a scientific journal, there are helpful companies to write the ritual Submission Letters, handle all the negotiations with the Editors, squash your syntax into English, and rearrange your prose to fit the Procrustean conventions of the genre (using “rearrange” in a broad sense that includes “writing the whole paper for you and making up the experimental results”). In short, they escort you through the foreign culture of “research”, using their collegial and mutually-beneficial connections with the concierges of selected journals to ensure favourable peer-reviews as part of the package. As a way of signing their work, these facilitation companies liked to finish their edited manuscripts with formulaic apotropaic Acknowledgement declarations: template-driven courtesies to colleagues and the nonexistent ‘reviewers’.
Academic publishing has a comparable vocational niche. If your career needs a paper in a scientific journal, there are helpful companies to write the ritual Submission Letters, handle all the negotiations with the Editors, squash your syntax into English, and rearrange your prose to fit the Procrustean conventions of the genre (using “rearrange” in a broad sense that includes “writing the whole paper for you and making up the experimental results”). In short, they escort you through the foreign culture of “research”, using their collegial and mutually-beneficial connections with the concierges of selected journals to ensure favourable peer-reviews as part of the package. As a way of signing their work, these facilitation companies liked to finish their edited manuscripts with formulaic apotropaic Acknowledgement declarations: template-driven courtesies to colleagues and the nonexistent ‘reviewers’.
- “We would like to acknowledge the reviewers for their helpful comments on this paper”
- “We would like to acknowledge the helpful comments on this paper received from our reviewers.“

This is the background to the 2015 and 2017a/b Extinction Events when various journals from the Springer stable depublished hundreds of papers, reluctantly accepting that their peer-review process had been replaced by a cabal of semi-sentient rubber-stamps. Notably, Tumor Biology and Molecular Neurobiology. The retraction notes were repetitive to the point of self-plagiarism:
“The Publisher and Editor retract this article in accordance with the recommendations of the Committee on Publication Ethics (COPE). After a thorough investigation we have strong reason to believe that the peer review process was compromised.”
Which is not to say that the papers were all ghost-written. Though some general themes recurred, as if manuscripts had come off an assembly line, the retractions were not a homogeneous collection, and many were probably the work of the nominal authors. Observers were careful to point out that some of the papers might even have been valid, and that authors were not always at fault here. They should have known what they were paying for with the guarantees of publication, but partly they were paying the dragomans not to tell them the details.
Retraction Watch quotes BioMedCentral’s retraction notices:
“It was not possible to determine beyond doubt that the authors of this particular article were aware of any third party attempts to manipulate peer review of their manuscript.”†
But getting at last to the point: A substantial fraction of these regrettable events were meta-analyses.
 Here I am following in the path of Guillaume Filion, who dredged up a genre of meta-analytic papers in 2014 that were distinguished by their reliance on a junk-science database, and appeared to have been composed by filling in the slots in a Mad-Libs sheet. The slight deviations from the script did not accumulate (unlike the successive errors in mis-copied medieval manuscripts), ruling out the possibility of sequential plagiarism, and preventing Filion from arranging them in order of copy-paste like a stemmatic tree of mis-copied medieval manuscripts. Which is sad. The world needs more stemmatic trees.
Here I am following in the path of Guillaume Filion, who dredged up a genre of meta-analytic papers in 2014 that were distinguished by their reliance on a junk-science database, and appeared to have been composed by filling in the slots in a Mad-Libs sheet. The slight deviations from the script did not accumulate (unlike the successive errors in mis-copied medieval manuscripts), ruling out the possibility of sequential plagiarism, and preventing Filion from arranging them in order of copy-paste like a stemmatic tree of mis-copied medieval manuscripts. Which is sad. The world needs more stemmatic trees.

What struck Filion, as well as these similarities, was the invocation of a “Begger’s Funnel Plot” — a wholly fictitious construct, confounding the actual concepts of “Begg’s funnel plot” and “Egger’s linear regression test of publication bias”. He flagged these mass-productions on the no-longer-extant PubMed Commons discussion board (for this was before the discovery of PubPeer) as well as blogging about them. Then Charles Seife expanded on the story for Scientific American, with a list of 100 suspicious examples, so publishers couldn’t ignore the warnings any longer and the retractions rained down like the hammer of Thor.
My own spreadsheet of 100 150 200 250 examples subsumes Filion’s discovery and the Begger’s Funnel Plot solecism, and overlaps with Seife’s. I extended it in two directions: by searching for the fulsome Acknowledgements favoured by the dragoman company and its coterminous papermill, but also for unjustified citations of Zintzaras & Ioannidis (2005a, 2005b). “Unjustified” because this corpus is not about gene-locus meta-studies, and Zintzaras & Ioannidis would not have been included in the papers by anyone who had read them.
As I skillfully foreshadowed many paragraphs ago, issues of publication bias and funnel plots do not apply when merging chromosome maps; nor random- vs. fixed-effect models to accommodate heterogeneity. Yet these two doyens of meta-analysis are invoked on precisely those topics… also on I² tests and Cochran’s Q-statistic (which is only mentioned fleetingly in Zintzaras & Ioannidis 2005b, to replace it with a nonparametric rank-order generalisation).

The citations are copied from a script, but more to the point they are wrong, and the work of someone ignorant of the sources they’re citing. The earliest examples appeared in 2010, 2011 and 2012 — authored by Jia-Li Liu from the Department of Oncology in the Fourth Affiliated Hospital of China Medical University (Shenyang, Liaoning Province).†
- Liu JL, Liang Y, Wang ZN, Zhou X, Xing LL. (2010). “Cyclooxygenase-2 polymorphisms and susceptibility to gastric carcinoma: A meta-analysis”. World J Gastroenterol.
- Yuan Liang , Jia-Li Liu , Yan Wu , Zhen-Yong Zhang , Rong Wu (2011). “Cyclooxygenase-2 polymorphisms and susceptibility to esophageal cancer: a meta-analysis”. Tohoku Journal of Experimental Medicine.
- L Zhang , J-L Liu , Y-J Zhang , H Wang (2012). “Association between HLA-B*27 polymorphisms and ankylosing spondylitis in Han populations: a meta-analysis”. Clinical & Experimental Rheumatology.
Now this is interesting (for sufficiently broad definitions of ‘interest’) as other early contributors to the corpus credit Dr Jia-Li Liu for revisions and suggestions. Or rather, he was credited by the initial version of the boilerplate Acknowledgement used by the hired ghostwriter / editing service. Then that Acknowledgement was revised to make it clear that J.-L. Liu was thanking himself for the quality of his work:
- “We would like to thank J.L. Liu (Department of Oncology, the Fourth Affiliated Hospital of China Medical University) for his valuable contribution and kind revision of the manuscript.” (Zhou et al 2011)
- “We would like to thank Jia-Li Liu (MedChina medical information service Co., LTD) for his valuable contribution and kindly revising the manuscript.” (Tian et al 2012)
- “We would like to acknowledge the helpful comments on this paper received from reviewers and Dr. Jiali Liu (MedChina Medical Information Service Co., Ltd.).” (Li et al 2012)
You might very well think that Dr Liu had become a scholarly dragoman, finding more satisfaction from massaging his colleagues’ essays or concepts for publication (and providing the results) than he ever found from oncology. With a name like “MedChina”, you might even surmise that Dr Liu’s editing service / papermill was set up under the aegis of China Medical University as an official policy of helping the clinical staff forge papers, but I could not possibly comment. I can say, though, that the list of MedChina customers was initially dominated by clinicians at CMU’s affiliated hospitals in Shenyang, Liaoning.
Seife arrived at similar suspicions about the role of MedChina, in his 2014 article:
“In November Scientific American asked a Chinese-speaking reporter to contact MedChina, which offers dozens of scientific “topics for sale” and scientific journal “article transfer” agreements. Posing as a person shopping for a scientific authorship, the reporter spoke with a MedChina representative who explained that the papers were already more or less accepted to peer-reviewed journals; apparently, all that was needed was a little editing and revising. The price depends, in part, on the impact factor of the target journal and whether the paper is experimental or meta-analytic. In this case, the MedChina rep offered authorship of a meta-analysis linking a protein to papillary thyroid cancer slated to be published in a journal with an impact factor of 3.353. The cost: 93,000 RMB—about $15,000.”
Hindawi journals published some relevant retraction notices, detailed enough to earn praise from Retraction Watch, in contrast to the gnomic notes provided by Springer. Their Integrity staff had already noticed that some of the meta-analyses falling under the rubric of the pattern identified by Filion had been channeled through Hindawi journals. Less subjectively, the documents submitted for publication were the work of MedChina, according to the Properties fields of the metadata. This was terrible OpSec and I have to wonder what sort of sloppy tradecraft are they teaching in schools these days? Here one such Hindawi retraction notice:
“The original files submitted by the authors were edited by MedChina, a company previously alleged to be involved in the sale of articles [3]. The authors could not be contacted.”
So from 2013, here is Dr Liu (of “China Medical University / MedChina Medical Service Centre”), inviting native English speakers to work on translations. $50 to do all the work on each manuscript while the customer pays $15000.
“Since we have the problem that our papers submitted to SCI journals have often been rejected because of our language problems, such as flaws on punctuation, word usage, spelling and grammar, vague statements and sentence structure, verb and subject consistency, and other language issues, native English speakers are needed urgently for us. You may help us with the job of language polishing toward our manuscripts in order to reach the basic publishing standards required by SCI journals.
The candidates should meet the following requirements:
- (1) Native English speakers from USA, Canada, Australia and European countries;
- (2) Excellent English reading and writing skills;
- (3) Familiar with medical research and the basic requirements of SCI journal;
- (4) Rich experience of writing and publishing biomedical SCI papers is a plus.
Salary:
It is paid according to the number of manuscripts you have processed for us. For each piece of manuscript, we will pay you USD 50. If you have finished excellent work, we may raise the payment up 50% or more, as appropriate.“
If my reconstruction is correct, MedChina would open a disposable burner email account at 126.com or 163.com on each customer’s behalf to handle manuscript submission, unless they preferred to deal with the publisher in person using their own e-mail accounts. To help the company staff keep track of clients and papers, we see corresponding-author account names of the form “cmu$h_XX@$$$.com”: cmu1h_yf, cmu4h_yf, etc. ‘dl1h_yg@tom.com’ was someone at the First Affiliated Hospital of Dalian Medical University, ‘shjt_lfh@126.com’ was Shanghai Jiaotong University… for as the customer base grew, the pattern expanded to accommodate other educational centres and their associated hospitals.
There were repeat customers: ‘cmu4h_jyz@163.net’ was corresponding author on four of these bagatelles (all in 2014). ‘wangheling1127@126.com’ on six papers (all in 2014). ‘hongshang100@gmail.com’ was an especially loyal customer, with seven papers (2011 to 2015). A meta-analysis might be satisfying in the short term but a few hours later you’re hungry again.
MedChina began by targeting low-profile, low-expectation journals: Asian Pacific Journal of Cancer Prevention, from a Korean publisher; Genetics and Molecular Research, from Brazil, reliably hospitable to papermills; Experimental and Therapeutic Medicine, from Spandidos, say no more.
They branched out, though, in search of higher Impact Factors, and Springer journals soon appear in the spreadsheet — along with Gene from Elsevier; PLoS One; DNA & Cell Biology and Genetic Testing & Molecular Biomarkers, both from Mary Ann Liebert.
One lesson of all this is how easy it is to write a meta-analysis. There are protocols and packages for the number-crunching part. The initial scouring of the literature is an automated database query. The hard part is to decide which papers from the query response are suitable as grist to the mill, and rating them for quality, before copying means and standard deviations into the sausage machine. The text can be unchanging boilerplate, from the Statistical Analysis section down to the obligatory mea-culpa admission of the study’s limitations and weaknesses.
[right] “Genetic variants of CYP2D6 gene and cancer risk: a HuGE systematic review and meta-analysis” (Zhou et al 2012).
No wonder Ioannidis inveighed against all the dilettantes and time-wasting triflers who were grinding out unnecessary, pointless reviews and misusing the tools he had created for that purpose. Isn’t it ironic?
“Currently, there is massive production of unnecessary, misleading, and conflicted systematic reviews and meta‐analyses. Instead of promoting evidence‐based medicine and health care, these instruments often serve mostly as easily produced publishable units or marketing tools.”
Ioannidis, “The Mass Production of Redundant, Misleading, and Conflicted Systematic Reviews and Meta-analyses” (2016)
So the question comes to mind, is a meta-analysis problematic just because some nameless scrivener at a papermill slapped it together on behalf of the nominal author? It depends how much rigour in the quality-filtering you can expect from anonymous hacks who don’t even read the sources cited in the Methods script, and from a company with a business model that’s all about churning out product. Can we trust declarations that “funnel plots and Egger’s linear-regression test showed no evidence of publication bias” when the papermill is paid not to write conclusions like “The extent of publication bias means that this paper isn’t worth reading”?
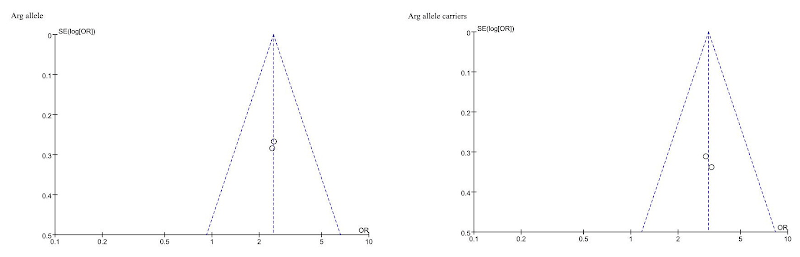
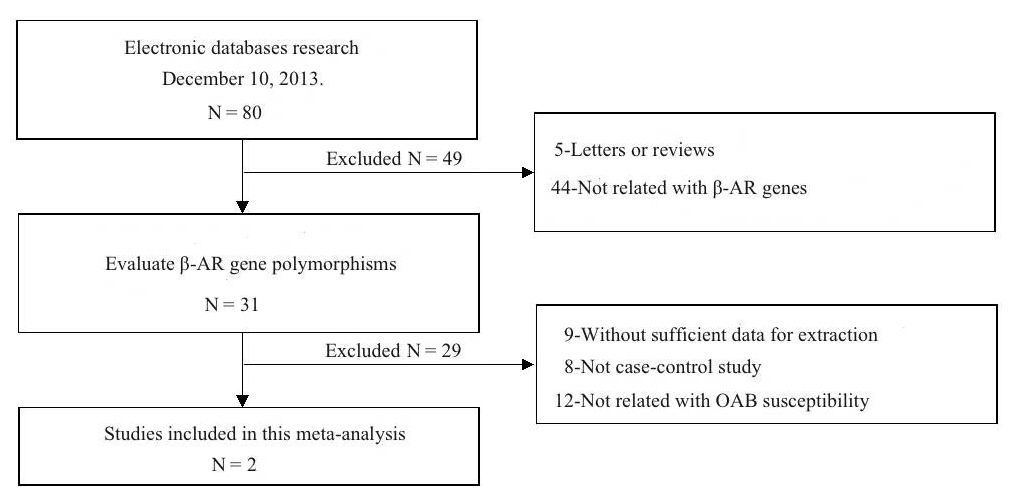
Connoisseurs of high-concept absurdity may appreciate “Association between polymorphism of β3-adrenoceptor gene and overactive bladder: a meta-analysis” (Qu et al 2015), where this style of flimflam achieves a kind of apotheosis. We learn that “The search strategy resulted in the retrieval of 80 potentially relevant studies. According to the inclusion criteria, 2 studies (Ferreira et al., 2011; Honda et al., 2013) were included in the meta-analysis and 78 were excluded“.
What follows is a solemn discussion of “sensitivity analysis”, testing the stability of the conclusions by “sequential omission of individual studies”… that is, by ignoring each paper in turn and performing a “meta-analysis” on whichever paper was left. The authors ruled out the possibility of publication bias by fitting funnels and regression lines to two data points. This is not even the only case of the scriveners solemnly applying the machinery of meta-analysis to a pair of studies: a second data-point is “Association between B7-H1 expression and bladder cancer: a meta-analysis” (Wang et al 2015), but there are others.
So I am going out on a limb here to propose that analytical rigour was not in fact a high priority within the papermill.
Not content to remain in the meta-analytic niche, Dr Liu and MedChina were already diversifying in 2014 and expanding to other arenas of biomedical fraud (“The price depends, in part, on the impact factor of the target journal and whether the paper is experimental or meta-analytic“). We see that in the repeat customers, where some of the regulars signed their email accounts to junk papers that don’t hew to the meta-analytic template, although they have the hallmarks of a papermill provenance, and the same fulsome Acknowledgements.
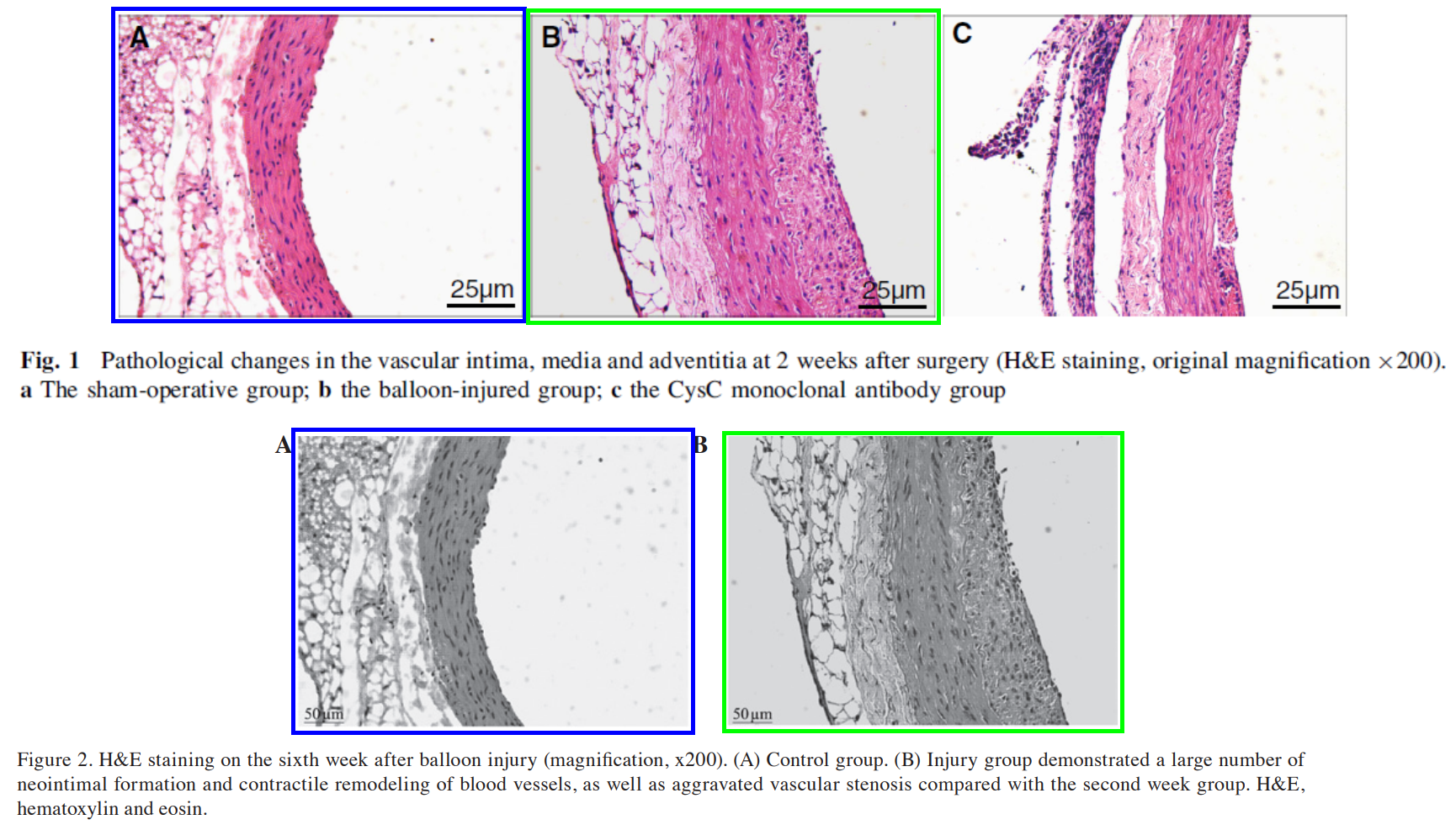
One possible example is “The role of cystatin C in vascular remodeling of balloon-injured abdominal aorta of rabbits” (Wu et al 2014), retracted from Molecular Biology Reports for the usual reasons in the 2015 Extinction Event (“strong reason to believe that the peer review process was compromised“). The Figures were recycled from “Expression and significance of α-SMA and PCNA in the vascular adventitia of balloon-injured rat aorta” (Wu & Lu 2013), extruded through a Spandidos journal. But if MedChina wrote the 2014 version and expedited its publication (“We would like to acknowledge the helpful comments on this paper received from our reviewers“), it is likely that the 2013 version has the same origins.
More informatively, Yu-Xia Zhao (“cmu4h_zyx@yeah.net”) progressed from
- “Diagnostic performance of serum macrophage inhibitory cytokine-1 in pancreatic cancer: a meta-analysis and meta-regression analysis” (Chen et al 2014a) (retracted)
- “Relationships Between p16 Gene Promoter Methylation and Clinicopathologic Features of Colorectal Cancer: A Meta-Analysis of 27 Cohort Studies” (Chen et al 2014b) (retracted)
- “Genetic polymorphisms in the CYP1A1 and CYP1B1 genes and susceptibility to bladder cancer: a meta-analysis” (Chen et al 2014c) (retracted)
to
- “PARP-1 rs3219073 Polymorphism May Contribute to Susceptibility to Lung Cancer” (Wang et al 2014)
- “Single-nucleotide polymorphisms of LIG1 associated with risk of lung cancer” (YZ Chen 2014) (retracted)
- “Relationships between genetic polymorphisms in inflammation-related factor gene and the pathogenesis of nasopharyngeal cancer” (Qu et al 2014) (retracted).
There is a ‘genetic polymorphism’ theme emerging here. I remind anyone who’s still paying attention that the meta-Meshuga was only one component of the depublication tsunami of 2015/2017. Other broad themes can be discerned within the diversification, but they lack obvious flags like “Begger’s plot” and “Zintzaras & Ioannidis” to help codify the scripts generating them.
In fact I would not be at all surprised if Dr Liu’s MedChina venture was the nascent, embryonic form of the later and much larger “Contractor papermill“. That follows the same paradigm, guiding the customer to the publication finishing line by contributing as much or as little is needed: from polishing a nearly-complete manuscript and filling a few gaps with hand-drawn flow-cytometry plots, to providing the whole package, needing only the customer’s co-authorship.
And some of the boilerplate Acknowledgements used by Dr Liu to sign his compositions are continuous with the ones currently gracing Contractor productions:
“We would like to acknowledge the reviewers for their helpful comments on this paper”
This is why the whole entertaining historical episode offers more than mere antiquarian interest. The assembly lines did not stop in 2015 or 2017, for they had already re-tooled to provide clients with a broader product range. Even the papermill hallmarks of Begger’s Plots and boilerplate and citational solecisms continued through to 2021 (though to be scrupulously fair, some of the later instances that I’ve listed in the spreadsheet will be independent ones, with authors merely plagiarising from the papermill’s products rather than paying them). Meanwhile, after expressing concern and chagrin that scammers had taken advantage of their generous naivety, journal editors and publishers went straight back to their old incaution.
Yes, depublication purges occurred, from Springer; and from Mary Ann Liebert, whose retraction notices credited Filion, and were admirably detailed if not entirely lucid. But these only accounted for a fraction of MedChina’s output; the majority of its products didn’t leave such an egregious trail of corrupted peer-review, so they survived without scathe and continue to stink up the literature. Many lack the Zintzaras-Ioannidis thumbprint, and although their Statistical Analysis sections are script-driven and stylised, the repetitious, boilerplate lists of references are at least relevant, and harder to criticise. As for other journals, the editors studiously ignored all the heads-ups they received. That Sci.Am. list includes many PLoS papers, for instance.
The last word belongs to “Jigisha Patel, BioMed Central’s‡ associate editorial director for research integrity”:
“I wasn’t aware there was a market out there for authorship“.
* * * * * * * * * * * * * * * * * * *
† I haven’t mentioned the BMC-journal contribution to that 2015 Extinction Event, even though that publisher’s 43 retractions for peer-review-corruption included a number of meta-analyses. The thing is, they have different hallmarks of an assembly-line provenance, and were seemingly written by a competitor to MedChina, targeting different journals, so there was no place for them in the narrative. What can I say? Meta-analyses are easy to cobble together. They are training wheels for papermills.
‡ Dr Liu seems to have held a second affiliation at Department of Oncology, Renji Hospital, Shanghai Jiao Tong University.

Donate to Smut Clyde!
If you liked Smut Clyde’s work, you can leave here a small tip of 10 NZD (USD 7). Or several of small tips, just increase the amount as you like (2x=NZD 20; 5x=NZD 50). Your donation will go straight to Smut Clyde’s beer fund.
NZ$10.00








Unfortunately, most of what is published nowadays is garbage, particularly those articles published in peer-reviewed journals where the reviewers are supposedly anonymous but we all know all the background games taken place to publish a paper and in high-impact journals. Peer- review should be named and public and publishing an article should be free too and all access should be given to original data when required and possible.
LikeLike
Pingback: The papermills of my mind – For Better Science