The Olivier Voinnet affair, the biggest research fraud scandal in modern plant science, is officially over. It is time to forgive, to forget and to move on. Let us not look back on those retracted, corrected and let’s-pretend-fraud-evidence-isn’t-there Voinnet papers and focus on what’s really important: science. The pure, unadulterated academic science, which is obviously first and foremost about placing beautiful papers in impactful journals and helping yourself and your friends to public money, while making sure that traitors, snitches and envious haters are “drummed out of science”, as Voinnet himself once threatened it towards his whistleblower, Vicki Vance.
Everyone moved on. Plant scientists summarily put the Voinnet embarrassment behind them, some see my past Voinnet reporting to be a kind of obsessive stalking. French CNRS, Voinnet’s original employer, declared in a more recent case of CNRS chief biologist Catherine Jessus the Photoshop data manipulations to be good scientific practice and called all French scientists to vigilance against whistleblowers (read here). Swiss federal university ETH must be quite happy with having elegantly avoided to sack their now CNRS-delegated professor despite the massive misconduct findings: just in the last two years Voinnet published papers in elite journals The Plant Cell, Molecular Cell, and PNAS . The journal Nature Plants even explained why Voinnet and his associates are welcome to submit their papers again. Despite new evidence of manipulated data still popping up, the journals drew a line and are not interested anymore. Especially the elite journal Nature Genetics, whom I sent in June (as a reminder) a dossier of four Voinnet’s papers in need of some editorial attention. Not much happened, which may or may not have to do with the fact that the editor of Nature Genetics Myles Axton has some strange data in his own last-author paper in the journal Development. Incidentally, same journal which found itself unable to retract a Jessus paper having established massive data manipulations in it.

Tainted editor?
The material in Axton’s Development paper may not be of Voinnet “quality”, but is still troublesome and in need of an explanation. This is the paper:
Andrew D. Renault, Xiao-Hua Zhang, Luke S. Alphey, Lisa M. Frenz, David M. Glover, Robert D. C. Saunders, J. Myles Axton
giant nuclei is essential in the cell cycle transition from meiosis to mitosis
Development 2003 130: 2997-3005; doi: 10.1242/dev.00501
And this is the evidence one of my readers alerted me to (it is also available on PubPeer). Because of the very well hidden digital gel splicing, it is difficult to trust the scientific results as they are presented.

So the scientist responsible for that is in charge of overlooking, pardon, of overseeing his journal’s policies regarding Voinnet and other cheaters. Axton at least claims to understand the concept of image integrity, he even used the example of Voinnet’s image duplications in a lecture in Australia, in November 2014. Here a slide from that talk:
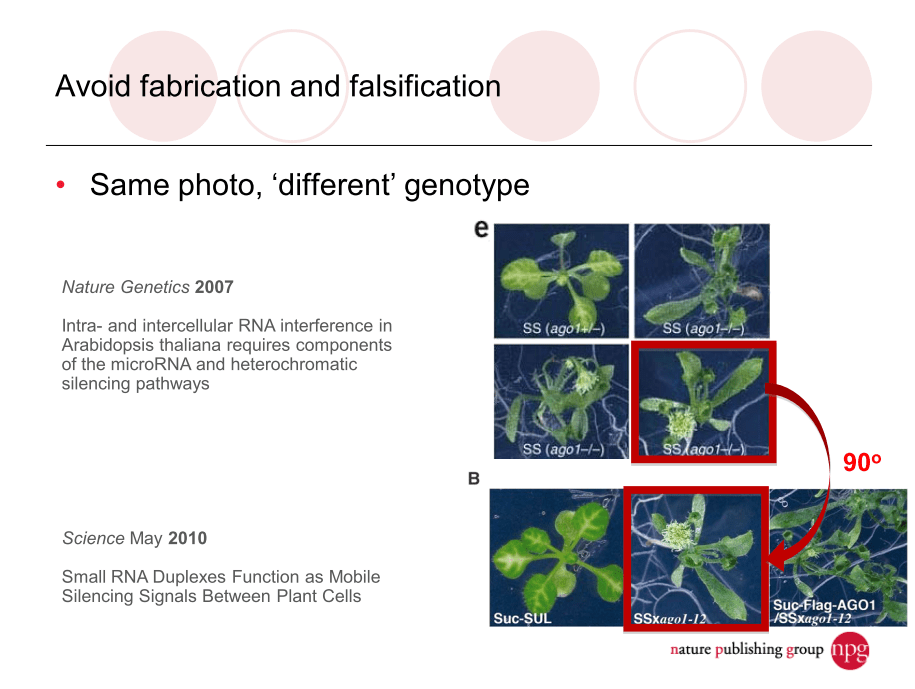
Back in 2014, Axton was apparently under impression the main issue in the Voinnet scandal was image copyright, and because Voinnet re-used an older figure from Nature Genetics in Science 3 years later, no action was taken by the former journal. The note attached to the presentation, likely Axton’s cue to himself, reads:
“The same authors turned this little weed through 90decC and it appeared three years later in Science, requiring a correction to cite the original paper.
In fact Science does not allow publication of any material already published. If your lab is open to collegial criticism and runs on transparency and objective evidence
(Hypothesis, evidence, method for everything) it is more resistant to this sort of careless error than a lab focused only on getting results. Mistakes are common and caught by the same openness and skepticism that protects your lab from more serious misconduct including fraud. High impact means high visibility”.
Voinnet since earned 8 retractions, including the above mentioned Dunoyer et al Science 2010 paper, but not a single retraction in Nature Genetics. The journal issued one correction though, for Dunoyer et al. 2006 . The dossier I shared with Axton and Nature Publishing Group contains further evidence of manipulated data in that paper, and in 3 more, in the same journal.
Anomalies and imperfections
My first email contact with Axton was also about the Voinnet affair, it happened in January 2015. Shortly after Axton educated scientists on image integrity using the Voinnet example. I wrote to Axton about this paper:
Arturo Marí-Ordóñez , Antonin Marchais , Mathilde Etcheverry , Antoine Martin , Vincent Colot , Olivier Voinnet
Reconstructing de novo silencing of an active plant retrotransposon
Nature Genetics (2013) doi: 10.1038/ng.2703
The issue I needed Axton’s help with concerned some strange bands in the Figure 2H, which looked floating over the gel background. There was a PubPeer debate where an “expert” explained that such patterns can arise where the area around of the gel band is selectively contrast-adjusted. Hence, it shows no band copy-pasting, but is rather a compression artefact of differential image processing inside an image. E.g., to enhance a band. Nothing to point fingers at, according to the “expert”. The explanation, which would have any halfway-honest wet lab biologist have their hair stand on end, was however later seemingly endorsed by PubPeer themselves, who apparently operated under the new CNRS code of data integrity conduct. Only the original high-resolution figure could help understand what exactly was done to that gel image. Problem was, the authors were non-responsive, and ETH declared Mari-Ordonez’ doctoral thesis embargoed, so at that time I could not consult it for a better version of that image.
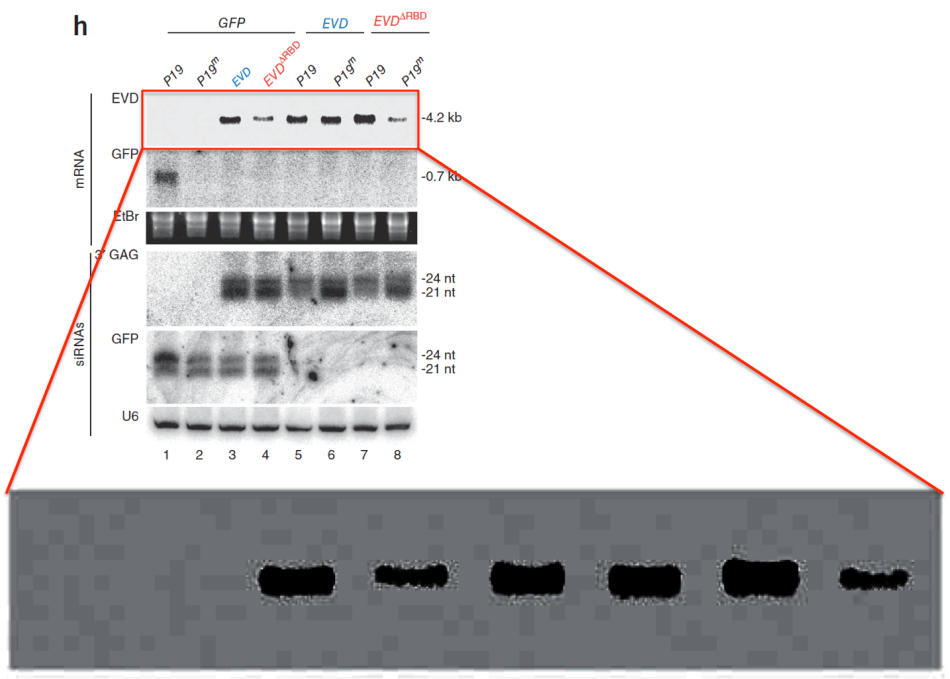
Thus, I asked Axton for the original image. This was what he wrote back to me:
“We maintain that anonymous peer review supervised by a journal with consistent standards is an excellent way to improve a research paper for reuse in the research community. It is also important to publish in a timely manner. This means that every paper contains anomalies and imperfections. Those that affect the main conclusions of the research or the utility of the paper may need to be corrected. It is also important to distinguish these from inconsistencies that have little bearing on the results and utility”.
I asked Axton for the high-resolution version of a published figure once again. Axton refused, with this one line:
“As you know copyright resides with the author. For permission to reprint a figure, please contact http://www.nature.com/reprints/permis sion-requests.html”
Soon after this strange email exchange took place, more evidence on that paper appeared on PubPeer, this time it was clearly duplicated gel lanes. Four years have passed and nothing happened. At that time the paper was slightly over one year old, yet it was too much trouble to ask the authors if they actually held any original data. So things like that were dismissed by Axton, the expert in applied and practical research integrity, as “inconsistencies that have little bearing on the results and utility“:
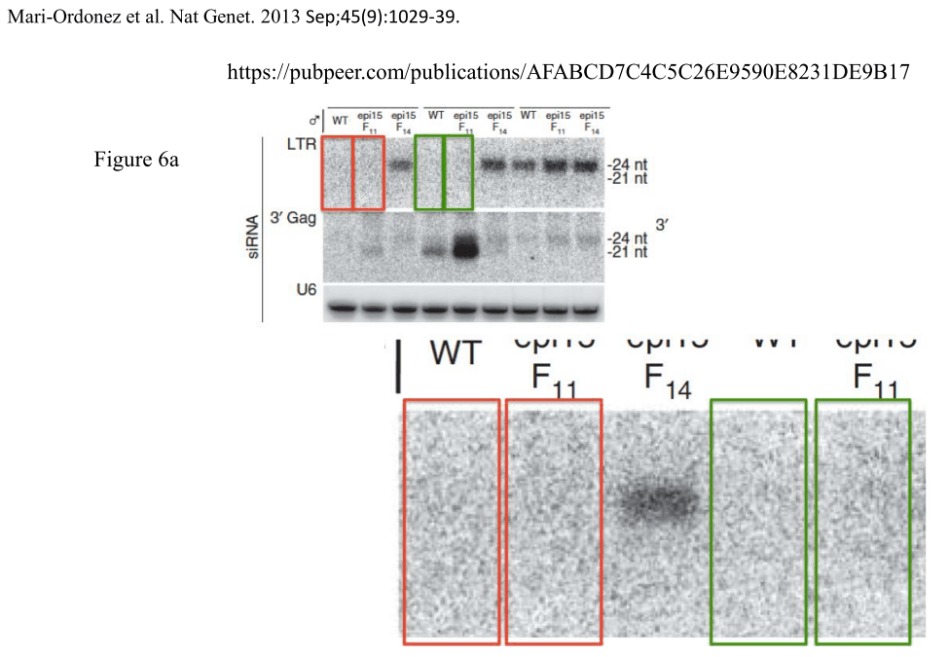
The shown evidence is included in the dossier I received from a reader and shared with Axton, which also contains additional stuff for that paper, for example:

Or loading controls, from Voinnet’s infamous “loading control library”, reused in whatever irrelevant context he saw fit:

Correction to close the case?
All that no more than silly “anomalies and imperfections” for Nature Genetics editors. No reason to do anything about it. The next paper in that journal I present (in no particular chronological order) is Dunoyer et al 2006, previously corrected in 2015.
Patrice Dunoyer, Christophe Himber & Olivier Voinnet
Induction, suppression and requirement of RNA silencing pathways in virulent Agrobacterium tumefaciens infections
Nature Genetics volume 38, pages 258–263 (2006) https://doi.org/10.1038/ng1722
Corrected was the Figure 5, where Voinnet reused a loading control, which in this case at least contained same experimental samples and was explained by stripping and re-probing the membrane. Nothing too bad. This is probably why after that correction, Nature Genetics closed the Voinnet case completely. The PubPeer users could now post anything they want: inappropriately reused loading controls, duplicated gel bands, mixture of both: life at Nature Genetics moved on. The dossier I gave Axton has much, much more on that paper, some of it was already published on my site. Here is additional evidence, not published on my site or PubPeer yet.
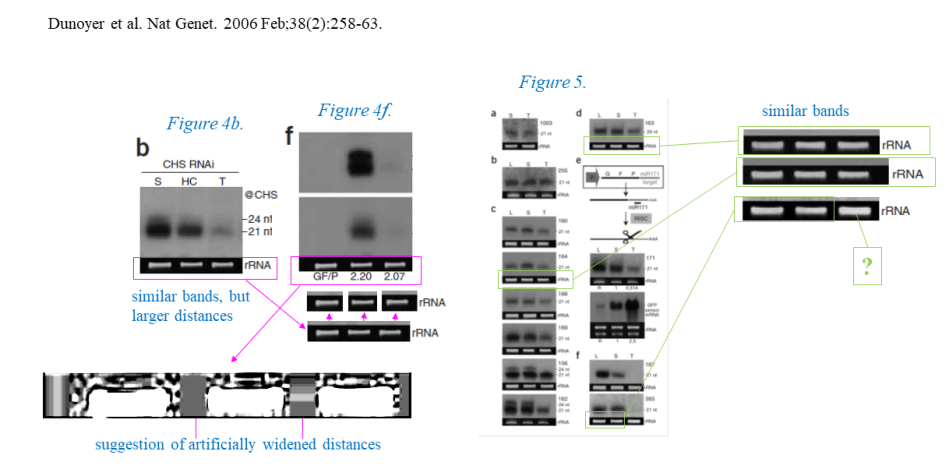
We see duplicated loading controls (creatively done, yet so common with Voinnet it’s almost but not quite boring, maybe this is why the journals stopped caring?). In one case, the bands were duplicated, but then pulled apart to match the gel they are supposed to be the loading control of. Why are loading controls important? Because we can’t trust certain authors on that paper to load equal amounts of samples, or any sample at all where no signal is desired.
The new dossier extends on the evidence I already published, like this extra appearances by same bands in yet another Voinnet paper:
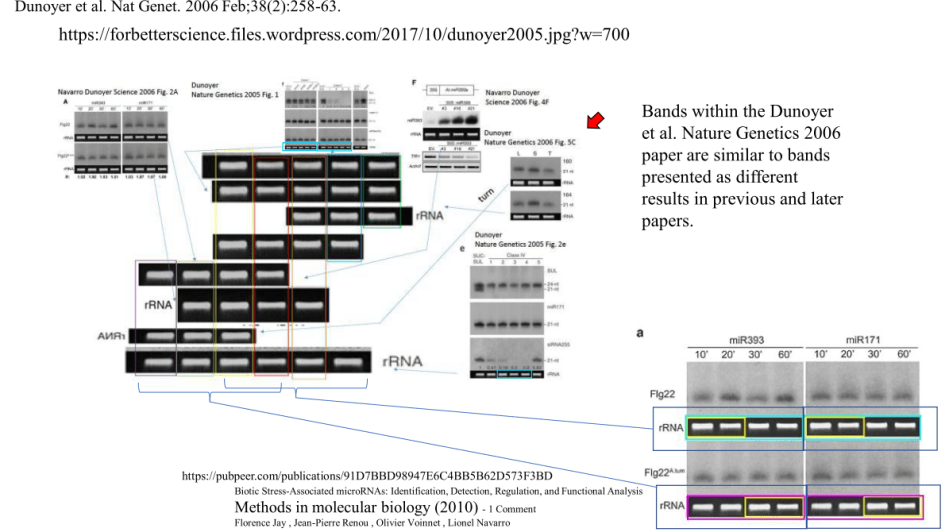
The paper Jay et al Methods in Molecular Biology 2010 contains not only misappropriated loading control data from Navarro et al Science 2006 (corrected) and Dunoyer et al Nature Genetics 2006, but also from Dunoyer et al Nature Genetics 2005.
Patrice Dunoyer, Christophe Himber & Olivier Voinnet
DICER-LIKE 4 is required for RNA interference and produces the 21-nucleotide small interfering RNA component of the plant cell-to-cell silencing signal
Nature Genetics volume 37, pages 1356–1360 (2005) https://doi.org/10.1038/ng1675
Shown is a sad story of a little RNA band which got lost in many papers and figures.
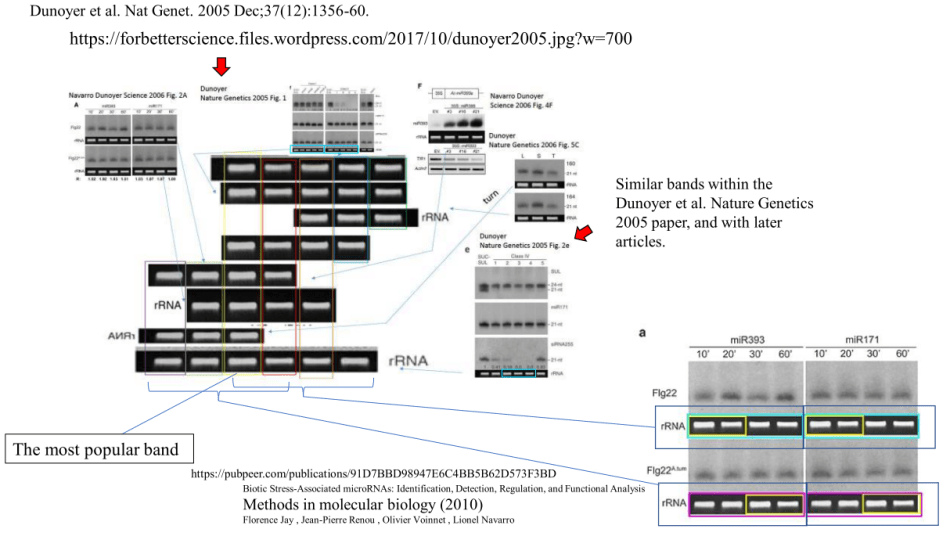
Of course it is not “just” rigged loading controls with Voinnet. We get duplicated gel bands also, in same Dunoyer et al Nature Genetics 2005 paper.

The craziest thing: this old Voinnet Nature Genetics paper from 2005 was NEVER flagged on PubPeer! It is one of his pristine, perfectly reliable ones, because free of any suspected data manipulations (if you trust the scientific claims therein, otherwise see my investigation here). Most likely there is not a single Voinnet paper free of manipulated data. All you need is someone with a good eye to keep taking a good look at Voinnet literature.
As was the case with this Nature Genetics 2007 paper, which does feature prominently on PubPeer. Not that the journal cares. Because much of the material was re-used in the later published and now retracted Science 2010 paper, Nature Genetics can pretend to host unique research data not published anywhere else. Image copyright is the only thing which apparently bothers the Editor-in-Chief Axton in the Voinnet affair. Maybe he must be asked to correct his own Development paper first, to understand the idea behind data integrity.
So here is the last Voinnet masterpiece I show. Also for this paper from 2007 some of the new evidence was already discussed on my site. Now some more, as a slide show even. Enjoy.
Patrice Dunoyer , Christophe Himber , Virginia Ruiz-Ferrer , Abdelmalek Alioua , Olivier Voinnet
Intra- and intercellular RNA interference in Arabidopsis thaliana requires components of the microRNA and heterochromatic silencing pathways
Nature Genetics (2007) doi: 10.1038/ng2081
Voinnet’s right-hand man and regular key co-author Patrice Dunoyer left science and France, to go back home to New Caledonia. Other first authors did rather well for themselves, despite their retractions and embarrassing corrections. Most are professors or otherwise tenured academics, others are well on their way there. In fact, the desire of the journal editors and academic elites to protect the careers of next generation of cheaters, errrr, pardon, of young genius scientists, was the reason why Science refused to retract a Voinnet paper from 2006. Never mind that it was found fraudulent and marked for retraction even by the benevolent ETH investigation. Cell simply stayed out of it. Nature Genetics pretends nothing at all happened. Life goes on.
Update 30.08.2018.
A source informed me that Arturo Marí-Ordóñez has now been made a group leader at the Gregor Mendel Institute in Vienna. He is officially listed as postdoc in Julius Brennecke’s group at Institute of Molecular Biotechnology (IMBA), which is the same building as Mendel Institute. Brennecke was described to be “a good friend of Voinnet”.
Update 4/5.12.2018.
Myles Axton left his position with Springer Nature as Editor-in-Chief of Nature Genetics and since 19.11.2018 works with a different publishing house, Wiley, according to his ORCID profile.
His task at Wiley is to create “a premier, broad-interest, Open Access journal serving researchers in the field of genetics and genomics […] with Editor-in-Chief Myles Axton”

Donate!
If you are interested to support my work, you can leave here a small tip of $5. Or several of small tips, just increase the amount as you like (2x=€10; 5x=€25). Your generous patronage of my journalism will be most appreciated!
€5.00


Is this surprising?
Why put an end in such a symbiotic and mutual profitable relationship?
LikeLike
Research institutions that are serious and do care about science should put pressure on the publishers and reconsider the already over prized subscriptions if they are not changing their policy regarding publishing invalid and manipulated data in which they know is problematic. The only thing they apparently care about is their profit so if they notice a tiny indication that research institutions start to ask about quality and trust then they maybe will do something.
We scientist should contact our home institution and present the evidence presented here and from Pubpeer.
One nice example on the list: Lancet’s infamous editorial: “Paolo Macchiarini is not guilty of scientific misconduct” from 2015.
LikeLike
Most journals are little more than vanity presses: they take money from those who want to publish and from those who want to read what is published. A very few, most notably JBC, take their role as guardians of science seriously. They have established a paradigm we should applaud and support, rather than waste time slagging those who publish anything and retract nothing. Cheaters like Vionnet owe their success to a lack of due diligence by those who pay and support them, and to the indifference and corruption of their peers. Corrupt societies support and champion corrupt researchers, politicians, artists and all manner of opportunistic vermin that learn to hide behind the skirts of nationalism. To the French, Vionnet is like Lester Maddox was to US southerners in the Randy Newman song Rednecks: he may be a fool, but he’s our fool.
LikeLike
Every person will agree that none of the loading controls in the four Voinnet Nature Genetics papers can be trusted. In addition, each of the four papers shows manipulations of the main data figures, which I summarized here
At Pubpeer, for Axton, the NG chieftain, another suspicious picture is shown. It says that it may be a compression artifact, but it looks as if the second band was photoshopped
LikeLike
https://www.ethz.ch/en/news-and-events/eth-news/news/2018/09/ETH-professor-not-guilty-of-manipulation.html
This is grotesque. He is the PI and Senior Author. He is responsible and he is PERSONALLY responsible as he is the PI and Senior Author. Whether he created an ENVIRONMENT that encouraged fraud/manipulation or INSTRUCTED it is secondary to the fact that he is the man in charge. He will have to leave science, he will probably also have to leave as this is becoming too stinky.
LikeLike
Voinnet was under reprimand, in 2015 he and Dunoyer admitted to have manipulated all data alone. Were Voinnet to be caught on faking data again, ETH would have no way to avoid sacking him.
LikeLike
The real question is who is LEGALLY responsible. Did the ETH consider this? Do they want to be legally responsible or would they agree V/D should be?
LikeLike
The ETH 2015 deal was about “you tell everything and fix everything, and we forgive everything and allow you a new start”. The first part of the bargain was not kept, and for that Voinnet accuses his co-workers. Without details, however, accusations between authors are completely meaningless.
LikeLike
Pingback: Olivier Voinnet: not guilty in past, present and future – For Better Science
Pingback: Olivier Voinnet, the new Dreyfus? – For Better Science
Pingback: Western blot loading control libraries and beyond – For Better Science
Pingback: Inspector Voinnet, Wollman in fraud-overdrive, and Farewell to Jessus – For Better Science
Pingback: Did Plant Cell learn from Voinnet Affair? – For Better Science