Human Brain Project is supposed to be half-way onto simulating the human brain in a supercomputer, the €1 Billion EU project withstood all criticisms, protests, ridicule and mutinies, from inside and outside, and even escaped proper evaluation. Its last remaining challenge on the road to full success is the overbearing masculinity in its leadership ranks, which HBP was apparently asked by the EU to act upon after my tweets during a HBP conference. Maybe this was why HBP Executive Director Chris Ebell resigned just this month?
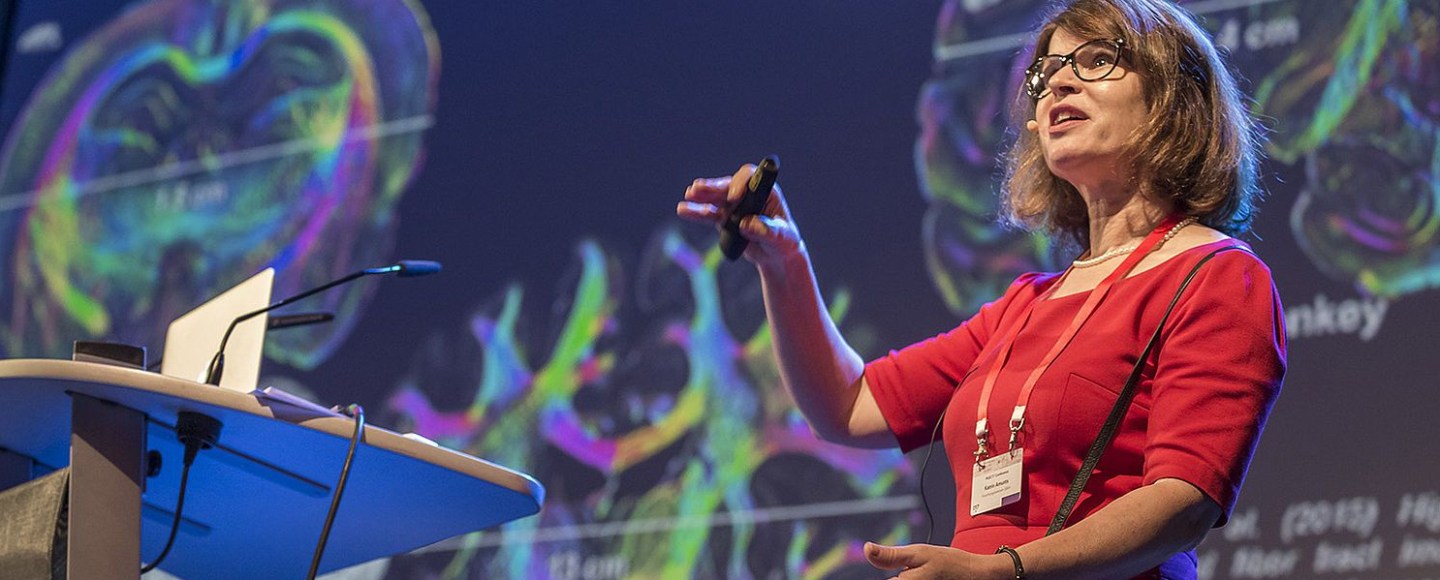
In that regard I was keen to interview the only woman among HBP leaders, the scientific director Katrin Amunts, professor for Brain Research at the University of Düsseldorf and director of the Institute of Neuroscience and Medicine in Forschungszentrum Jülich (FTJ). In November 2017, I published an interview with Thomas Lippert, one of the HBP leaders and director of the Institute for Advanced Simulation at FZJ. I then invited Amunts, who on 20 November 2017 agreed to do an interview by email. Eventually, I gave up, and published a non-interview, where my questions were completed with text bits I picked up on the HBP website. Right away, Amunts worte to me, offering to complete the interview. I received her replies soon after, and publish them here.
What crystallises now: HBP does not pretend to simulate the entire human brain in a computer anymore, Henry Markram’s main selling point of a “brain-in-a-box“, complete with consciousness, was discarded as science fiction, Markram himself seems utterly sidelined and outside all decision making process. The word “simulation” received a new meaning, a modest one, it is about mapping the brain now, and trying to simulate some very general principles of neurobiology. This is what the Amunts interview sounds like, of course we do not know what HBP executives keep telling the gullible EU bureaucrats.
Here is therefore a proper, long read interview with HBP scientific director Katrin Amunts.
Amunts was born in the communist Eastern Germany (GDR), she learned Russian and went to study medicine in 1981 at the Pirogov Institute in Moscow, in USSR. Afterwards, she stayed for a PhD at the Federal Centre for Mental Health of the Academy of Sciences of USSR, her thesis was titled “Quantitative analysis of the cytoarchitecture of area 4 of human brain cortex in ontogenesis“. The associated Moscow Institute of Psychiatry featured a huge collection of human brain samples, due to the special legal situation in Soviet Union.
The Moscow graduate defended her thesis just as the socialism in Eastern Europe was collapsing and her own country ceased to exist, the Soviet Union soon followed suit. In 1993, after a brief stay in Berlin, Amunts moved to the West, to work with the neuroscience professor Karl Zilles, head of C. & O. Vogt Institute at the University of Düsseldorf . In 2013, she was appointed as Zilles’ successor there.
There are more details on Amunts work in brain cytoarchitecture in my previous article.
Now, the interview, my questions are labelled with LS, Katrin Amunts’ replies with KA. I adjusted the scholarly paper formatting and incorporated the provided hyperlinks into the text. I highlighted parts of Amunts’ replies I deemed particularly important for the understanding of HBP. The interview begins.
LS: Your own research platform is about human brain structure. In this video, you explain the approach: thin-sectioning of donated human brains, in order to scan those and assemble a 3D computer model.
Yet this says nothing about the actual neuronal structure and connectivity of the brain on the cellular and intracellular basis. How do you intend to map all the neurons at HBP? Your site says: “3D-PLI allows to map nerve fibers and fiber tracts in whole postmortem brains with a resolution of a few micrometers“. In this interview, you say it is 20 micrometers (and that you are confident the brain can be decoded by HBP). That is not really close enough to properly distinguish individual neural fibres, never mind synapses. How would you comment on these limitations? How will you fill the huge predictable gaps from the scanning data?
KA: Let me first emphasize that understanding the brain is more than mapping and analyzing cells. The brain is organized on different scales in space and time, including molecules, cells, microcircuits, up to large scale networks with temporal changes in the range of milliseconds, but also long-term changes over the whole lifespan. To understand human brain organization means to integrate these different aspects of brain organization into a comprehensive concept. I started my own research on the level of cellular architecture (=cytoarchitecture), which is a natural and good reference for relating other modalities to, as nerve cells are central for brain function. Differences in architecture suggest differences in the functional involvement of an area. This hypothesis has been proven many times. Therefore, a most accurate representation of the brain map with regard to cell architecture and the segregation of brain areas is a prerequisite for a better understanding of the relationships between structure, function and behavior.
Our initial model BigBrain has a resolution of 20 micrometer. Recently we have significantly improved the resolution of this type of mapping: We can now go down to mapping a brain at the resolution of 1 micrometer; this is sufficient to resolve the individual cells to a certain level of detail and to link such info to single cell modelling. Regarding 3D-PLI, the resolution does go down to 1,3 micrometer, see e.g. here: Zeineh, …, Amunts & Zilles, Cerebral Cortex, 2017. This resolution allows to distinguish nerve fibers in many cases, although it is not a method to image synapses, where you need methods like Electron Microscopy. In contrast to the latter, however, 3D-PLI is capable to cover the whole human brain. I.e., both methods have their strengths, but also disadvantages.
At all levels of brain research, there are gaps between the data obtained on different spatial and temporal scales. Not one method alone can provide the complete picture; it is in each case just information about one or a few aspects of brain organization.
Working out ways of relating the very large data sets, that are gathered on different scales and on different aspects, to each other is precisely what I would see as one major challenge in neuroscience and that is what we are trying to do in the HBP. One way to approach this challenge is a multimodal and multiscale 3D brain atlas. Such atlas must be able to integrate data from different labs in different countries since no single lab is capable to acquire comprehensive data on all aspects of brain organization. A lot of work is needed to curate and standardize data, to make them compatible for multimodal and multiscale models, and to visualize it.
This development requires collaboration with experts from Neuroinformatics as they build a platform where the data is available in useful ways, can be shared and analyzed, and where further data can be incorporated and linked across levels to better understand the multiscale organization of the brain and bridge the gaps. Results at different levels and from different sources can be combined at a large scale, and our data exchange infrastructure FENIX allows to do this even on a raw data basis up to Petabytes of data. This bridging of gaps means to gain a better understanding of how phenomena on one scale translate to what we observe on the other scales.
We tackle this problem systematically. Our method 3D-PLI that you asked about is a good example of this: Structural connectivity is currently mapped on very different spatial scales. With Electron Microscopy, 2-Photon Microscopy and other extremely high-resolution methods you get a “ground truth” on a nanometer level, and identify the actual connections of a very small number of cells in a volume. But you can only do this extremely detailed mapping for rather small volumes of brain tissue, and long-range connections cannot be mapped to such a level of detail in the human brain. There is currently no realistic way to map a whole human brain on this level. Another widely used method in Connectivity research, Diffusion Tensor Imaging lies on the opposite end of the spatial scale. Its resolution is a little bit below the millimeter range, but it has the advantage to scan the whole brain of living subjects very quickly. Even if these two methods describe the same aspect of brain organization, i.e., the connectome, the scales on which they work are extremely far apart.
In fact, 3D-PLI does not rival EMs and 2-Photon subcellular resolution or DTIs speed; what makes it so interesting is that in the tradeoff between resolution and speed it stands between these two extremes. It is quite highly resolved, albeit not to the level where you would see synapses. And it is fast enough that you can map whole brains, although it takes much longer than with DTI (and of course is limited to post mortem brains). Relating and comparing data obtained from all three levels of investigation methodology can help us understand how the slowly obtained “ground truth” in terms of wiring relates to what you see in the faster obtained, but of course more coarse-grained images for whole brains. In HBP experts for these methods are currently working together striving to bridge the gaps between the different scales of connectivity.
One general clarification is required at this stage, as several of your questions relate to the problem of gaps in the current data and the idea of completeness in mapping:
I hope we can agree that maps, models and simulations are necessarily a reduction, both regarding the spatial scale and the modalities you choose. When we talk about brain modelling and simulation, this does not mean that we create a 1-to-1 virtual duplicate of “the brain”.
Instead there are many different models for different questions, and the evaluation criteria should relate to the usefulness of the model, not an unattainable idea of perfect and “complete” representation. I have tried to emphasize this ever since becoming Scientific Director of HBP, see for example the interview I gave to the Laborjournal in 2016 or our Roadmap Paper in Neuron the same year. So our goals are to provide models and simulations, as well as open tools and databases, which are useful for scientific inquiry or clinical applications, and we deal with the gaps just like any other project either by collecting missing experimental data where possible, or by trying to understand the relation of what we see on different levels through theoretical approaches.
LS: Your own website says: “This model will not only help reveal the neurobiological basics of mental capacities, but will also enable characterization of their individual facets and underlying mechanisms.” Could you explain how a map of a brain, if it was ever achieved with the tools available, can give insights into how the brain actually works?
KA: It seems quite obvious to me that the many maps of the brain that have been developed over previous decades, however imperfect they may have been and still are today, have been an important basis for advancing our understanding of “how the brain works”, insofar as you try to understand how the complex structure of the brain relates to the functional side.
In that sense we are continuing a long tradition of brain science, just on a new scale, embracing modern means of digitalization and high-performance computing to deal with today’s large and extremely high-resolution data sets. I would find it hard to imagine that an understanding can be gained without this as basis, it would be like theorizing in a vacuum.
Maps like JuBrain and the BigBrain model are being used by the community for functional studies to better understand the topography of their findings, to increase spatial precision, to exclude hypotheses or to open new lines of thinking. They can shed light to interesting relations across scales of organization. (See e.g. Gomez et al.: Microstructural proliferation in human cortex is coupled with the development of face processing (Gomez, …, Amunts, Zilles & Grill-Spector, Science, 2017).
![csm_PASC17_035_8ac4a386cd[1]](https://forbetterscience.com/wp-content/uploads/2018/09/csm_pasc17_035_8ac4a386cd1-e1536306017639.jpg?w=950)
LS: The HBP approach is to map all the neurons of the brain of a human and a mouse. But what about glia cells, the astroglia and oligodendroglia? Those are active and necessary cells of a brain (astroglia are even electrophysiologically active), without which the brain will not be able to function at all, or, on the smaller scale, to process the signals correctly. Which department at HBP addresses the glial networks, and how is this work being integrated?
KA: First of all, it should be emphasized that mapping the neurons is very far from a complete description of the HBPs approach. Mapping and analyzing neurons in certain brain regions is a part of work packages in mainly two Subprojects, Human and Mouse Brain Organization (for more details see here). What the cell level atlas provides is the reference framework in which the other modalities, e.g. connectivity data on the different levels of granularity/resolution, receptor density and brain parcellation, functional connectivity etc. are being integrated.
It is true that the role of glial cells is an important area of research, and that integration of these data into brain models and simulations is desirable. The Human Brain Atlas and the Neuroinformatics platform with all their tools are built in such a way that data on glial cells can be easily integrated.
You are right in that we started our project with a focus on neurons. If you have to pick a cell type to start with for a brain model, the neurons are without a doubt the first and obvious choice, both because of their role and the vastly larger amount of data that we have on them.
Still we have work in HBP on glia that is aiming to lay the necessary groundwork. There is a task that specifically focuses on modelling glia, which is led by Dr. Marja-Leena Linne at the University of Tampere in Finland. She is in frequent exchange with the glia community, external and internal of the HBP, about the development of in-silico and theory models of glial cells (astrocytes but also microglia) and new results in terms of astrocytes’ influences on vascular systems, synapses, neuronal networks, animal behavior and cognition. All this has been and is currently applied in the development of both theory and simulation models of astrocytes.
As part of that work they recently published a survey of altogether 105 existing models for astrocyte-neuron interactions. The models have been categorized, evaluated and some of them have been re-implemented for reproduction (Manninen, Havela & Linne, Frontiers in Comp Neurosc, 2018). They also developed new simulation models for astrocytes’ influences on short- and long-term plasticity (Havela,… Linne, International Conference on Intelligent Computing (ICIC 2017), 2017). The complexity of these types of models have been reduced using novel methodology, with an aim to develop novel theory models of astrocytes (Lehtimäki, …, Linne, IFAC-PapersOnLine (20th IFAC World Congress), 2017).
Such models will readily serve in large-scale neural network simulations in the future. Further research relating to glia is done at the experimental level at Universidad Madrid, and also in the Blue Brain Project, for example.
Our aim is that the infrastructure of the HBP is used more and more also by researchers in the field of glial cells, to integrate and share results of their research with other researchers.
LS: You suggested in a presentation here the brain would never need or use replacement parts.
I understand this was a simplification for didactic purposes, since neural and glial cells are generated from the pool of neural stem cells throughout lifetime, even in old age. How does HBP intend to incorporate the neurogenesis aspect in the mapping process? Which HBP partners are involved in this?
KA: Yes, that was meant in contrast to techniques like hip-replacements; it was not meant to say that there was no cellular replacement throughout lifetime.
To your point, neurogenesis is a phenomenon that by definition would not be part of the actual mapping, but it could be taken into account in later models. It is an incoming topic of growing importance, but as it is a relatively recent area of research with relatively limited clear data, all in all it seems premature to include it in the current modelling efforts. It is not feasible to do everything at once in one initiative; the multiplicity of aspects is just too large.
But we put in place a platform with allows data or models on phenomena like neurogenesis to be added as it comes. So while we do not provide the comprehensive picture of neurogenesis ourselves, someone who works on this topic might use these tools to analyze his or her data and find correlations to other modalities through our platform. And from the technological side an integration of new insights about neurogenesis will be feasible, as the platform grows.
Just to reiterate, we are not claiming to cover all aspects of research on brain organization by ourselves. No single project can do that. Rather, the main focus is the construction of a lasting infrastructure for neuroscience, a technological system that is built for brain science and by brain scientists, with computational resources to systematically link models, experimental and theoretical neuroscience, and Big Data and simulation methods. We call it the HBP Joint Platform.
Our approach is to “co-design” the infrastructure through so-called use cases, where neuroscientists collaborate with experts that are building the Joint Platform of the HBP with its tools, models and data. In this manner we help to provide a basis to make it feasible to put the presently available data as well as future data into a coherent framework across scales, and linking across modalities so that scientists can find the data, analyze and exchange them, and make use of it for their own research with the help of the platform. Use cases will come more and more also from the outside of the project, helping to develop the Joint Platform of the HBP in a user, i.e., researcher’s driven way.
LS: After some initial uncertainty, it does look like one of the main goals of Human Brain Project is to simulate the human brain in a supercomputer. You yourself said in an interview in 2017: “By then, we should have computers that enable us to keep an entire human brain model in the memory on a cellular level and analyse it. In 20 years, I should think we’ll have realistic simulations of a human brain at nerve cell level that factor in the most important boundary conditions“. Your FZ Jülich colleague and HBP partner Prof Thomas Lippert did say in an interview with me that brain simulation should be just as doable as simulating weather or climate. How would this simulation happen, which biological input will the supercomputer need? Can HBP achieve the goal of collecting all this input?
KA: Hopefully what I have answered so far makes this a bit clearer.
Simulation is used as a tool for specific research questions. To cover a broad range of research, different simulation approaches are being used to various levels of abstraction. In any case, simulation is a very powerful tool considering the multi-scale organization of the brain. There are some aspects of brain organization, which are, e.g., for technical reasons, not accessible by empirical research at all.
In brain research, we are dealing with an overwhelmingly complex nonlinear system. Nonlinearity makes it very hard to predict the behavior of the system, even when you have a lot of experimental data. That is a reason why researchers in other fields that involve nonlinear systems have turned to simulations. Simulation has therefore a very important place in the project.
Regarding my quote, I would say that we are on a good way to do what I described there. As mentioned for modelling, we are currently using technology that allows us to map an entire brain at a 1 micrometer resolution, which is well below nerve cell size, and we are also collaborating with computer experts to really work with this extremely large-scale data. This requires, for example, new High-Performance Computing systems that have enhanced memory, or interactive supercomputing. Such collaboration is in line with the new European strategy for Exascale computers. These new systems would have the needed computing power to simulate, for example, large networks of nerve cells with the NEST simulator (Neural Simulation Tool). This simulator has recently been optimized to scale on these Exascale systems for brain size simulations.
Other types of simulation need different biological input – it starts from the very detailed simulations with NEURON, which is used in the Blue Brain Project, the Brain spiking neural network simulator to very macroscale simulation engines like The Virtual Brain, which simulates larger brain dynamics in whole-brain models. Simulation is also key in neurorobotics, where experts are implementing closed-loop experiments in physical or virtual robots, as a way for neuroscientists to test their models.
So simulation is and will be used and advanced in the project, what input and level of detail is needed depends on the question, and guiding should be again considerations of usefulness.
To understand larger brain dynamics, for example in Epilepsy seizures, it is e.g. not necessary to have everything modelled down to the cell or synapse. For certain questions like this you can use simulations and models that reflect features of the complex network architecture and dynamics to a reasonable level of abstraction.
As with the experimental methods, there is also much to be gained from bringing these different levels of simulations together.
And Thomas Lippert’s precise quote was“Such a model-based computational science approach has been successfully applied in many other fields of research: examples are simulations of models of materials, models of elementary particles or models of weather and climate. Computational brain research is not special in this respect.” This does not imply that brain simulation is easy, rather it states that the general approach to simulation is the same for any physical system.
LS: In this interview, you also mentioned that HBP works on understanding what consciousness is. Which tools are being applied for this? How can the brain atlas contribute?
Also, where do you see consciousness first appear in the evolution? Is it a unique human feature, or an intrinsic function of every complex biological neural system, even if it is not even a central brain as such (see cephalopods)? If the latter, how can any brain be ever simulated in our supercomputers?
KA: There are multiple groups in the HBP that carry out work on these questions across the Subprojects 3 (Systems & Cognitive Neuroscience), 4 (Theoretical Neuroscience) and 12 (Ethics and Society). Some are more interested in the clinical side, like disorders of consciousness, others more in fundamental and conceptual questions. The nice thing is that they are very much working together through the project, and collaborate with the very active community of consciousness research. This also led them to organize the first HBP International Conference on “Understanding Consciousness” in June 2018 in Barcelona. There are many different tools and experimental paradigms being used. You can find some links to further info here. The high resolution and multimodal atlas as a reference can be helpful for locating activities or what is often called neural correlates of consciousness more precisely.
About the evolutionary question: It seems quite clear to me that it [consciousness, -LS] is not a unique human feature, but developed earlier in evolution. But I could really only speculate on your question on cephalopods. There is still a lot to do towards a deep understanding of consciousness.
To the last part of your question: I am not sure I agree that simulating the physical system of the brain and its dynamics (which is also all that we really can address with experimental neuroscience methods) actually depends on this. We can answer many questions without this, for example, simulating brain function during visuo-motor integration. Still, consciousness poses highly interesting questions and I am glad that progress is being made in this area.
LS: In this regard, I presume also HBP is aware that the brain does not compute the way our (super-)computers compute. Otherwise there would not be a huge focus on developing neuromorphic computing in HBP. I am puzzled hence how HBP plans at the same time to simulate neuronal functions of an entire mammalian brain in a normal computer, and try to emulate the most basic neuromorphic information processing function by developing new technology quasi from scratch. It does look a bit like putting a cart ahead of a horse, but how do you see this?
KA: Neuromorphic computing is developing as a technology worldwide, because it offers very intriguing features for certain applications, including brain research. The HBP is the first research infrastructure, which is embedding two of the worldwide leading systems, SpiNNaker and BrainScaleS. You can read about them here and here. Both systems have been developed over several years.
Among the strengths are the potential for lower energy consumption, resilience and possibilities of coming closer to biological real-time by sacrificing some of the accuracy compared to supercomputing calculations. That makes them very interesting for research into phenomena like learning, or for applications in areas like robotic control. Specialists developing neuromorphic devices sometimes use HPC to simulate processes that can then be transferred to neuromorphic chips. Comparing the speed and energy consumption of compute jobs between both systems helps to better to understand the strengths and weaknesses of both systems. I also should emphasize that the relation between the development of neuromorphic technologies and the neuroscience in HBP does not have this one-way character that your question seems to imply.
It goes both ways, in that the development of these chips is also driven and inspired by working in collaborative projects with the neuroscientists in HBP. Our experts on neuromorphic computing learn from neuroscientists how to improve their architecture, for example by integrating knowledge about information processing in neurons with their dendritic branches.
At the same time, we observe that High Performance Computing is advancing rapidly and in innovative directions as well. To combine the strengths of both computing strategies into one would be extremely valuable. The idea of modular supercomputer does foresee that and allows dividing complex task into parts that are distributed onto the architecture that can do them most efficiently. See figure 5 here:

LS: HBP’s goal is still also to simulate brain diseases in order to find potential cures. E.g.:
Which diseases for example, and how would this happen? Could you give some neuroscience insights in the approach?
KA: There are several approaches, so these are just some examples. One that is quite advanced would the use of macroscale simulation with The Virtual Brain to understand seizure propagation in Epilepsy, and to support locating the epileptogenic zone in surgery preparation. The challenge in planning these surgeries is always to localize this zone as precisely as possible. A pilot study has shown that individual computational models of brain connectivity of patients could be helpful to improve outcome. So now the approach is being tested in a larger project called EPINOV, short for “Improving EPilepsy surgery management and progNOsis using Virtual brain technology”. The Virtual Brain technology is also applied to better understand recovery after stroke, for example. Some of the recent papers: Proix et al Nature Communications 2018, Proix et al Brain, 2017, Jirsa et al, NeuroImage, 2017.
NEST is a more detailed, cell-level simulator, and among many applications it has for example been used to investigate aberrant network behavior in diseases like Parkinson, see e.g. here (Bahuguna et al, Frontiers in Comp Neurosc, 2017).
On the subcellular level of molecular modelling we have a project on Modelling Drug Discovery that builds on our infrastructure. Such simulations are interesting in order to analyze the binding of drugs to a receptor and to understand the conformational changes; this is an important step towards identification and testing of potential drugs.
It should also be mentioned that Big Data approaches in the project are equally important, Alzheimers and Epilepsy are for example diseases that are also investigated with these approaches on our research infrastructure.
LS: HBP apparently considers mouse brain as a simple enough model to fall back on if human brain simulation cannot be achieved. Yet neuroscientist Prof Wim Crusio commented in this regard:
“If I would have to evaluate as reviewer a research proposal aiming to simulate a Drosophila brain on a €1bn budget over 10 years, I’d nix them for being overly optimistic. If the aim were to simulate a mouse brain, I’d inquire as to what they had been smoking when they wrote that application and recommend they start taking their meds again. Simulating a human brain? Comparing that with simulating materials and such? I’m afraid I’d actually be left speechless, although probably rolling on the floor howling with laughter…“
How would you, as neuroscientist peer, go and dispel Prof Crusio’s concerns?
KA: First, the premise in the first sentence is incorrect. The mouse brain activities are not something to “fall back on”. Such experiments may be used for specific questions, when the similarities between both species are close enough.
In order to really know the differences and similarities between different species, one has to address it by research, feature by feature, on all levels in space and time. Therefore, we do interspecies comparison.
The mouse is very important as the most widely used animal model, both in basic and preclinical research. It is supplemented by studies in rat, but also other animals. You can read about the different elements of HBPs work and how they relate to each other on our website, e. g. here and here.
And the answer to Prof. Crusios comment relates to what I said before on simulation. There are different simulation approaches for specific questions. I am convinced that simulation methods are an important tool for brain science and for understanding the complex relation between its organization and dynamics, and they supplement empirical work, theory, and data analytics.
You don’t need a perfect duplicate before simulation methods become useful. The idea is to make increasingly refined simulation, in a productive loop, with experiment, theory and modelling.
On another note, from my own work I just think it is vital that better ways are established for dealing with the vast amounts of data that we have and that are continuously gathered in neuroscience. We are already running into serious issues of structuring, analyzing and connecting the large datasets that are collected. These issues as of yet are unresolved. Simulation is one part of our larger concept on this matter of establishing a computing infrastructure for neuroscientists. We have laid this all out in a roadmap paper in November 2016 that you can find here: (Amunts, Ebell, Muller, Telefont, Knoll & Lippert, Neuron, 2016).
LS: Another neuroscientist, Dr Mark Humphries, is wondering what the scientific rationale of Blue Brain is. His concerns precisely are that the Blue Brain uses two-week old rats as source of data, also that non-primate prefrontal cortices lack the so-called “granular layer IV“. What exactly does Blue Brain bring to the HBP to understand an adult human brain? Can you comment as both neuroscientist and HBP scientific director?
KA: As you know the HBP was refocused. This article was published in August 2016, when the process was still ongoing, so a lot of what is written there is at least dated. But even then, the assertion that the cortical column was supposed to be the core of HBP is just not accurate, as the project had a much broader setup from the beginning.
Blue Brain remains an important contributor to the HBP’s Brain Simulation Platform, with know-how and technology as well as data that is highly interesting, not least for comparative animal and human neuroscience. Blue Brain is also very advanced, for example, when it comes to the methodical side of the model building process, and collaborates with other groups in the project, so the experience and expertise of this partner is very valuable in HBP.
The Blue Brain Project started before the HBP and besides some work package funding from the HBP grant it is mainly funded by the Swiss government.


Donate!
If you are interested to support my work, you can leave here a small tip of $5. Or several of small tips, just increase the amount as you like (2x=€10; 5x=€25). Your generous patronage of my journalism will be most appreciated!
€5.00


Just to reiterate, we are not claiming to cover all aspects of research on brain organization by ourselves. No single project can do that. Rather, the main focus is the construction of a lasting infrastructure for neuroscience
There does seem to be a complete lack of research goals. Just “Now you have given us 1 billion euros, we will make a database.”
LikeLiked by 1 person
exactly my impression as well. Now the infrastructure is the “fall back”. Well, if it is true I would suggest to put someone in charge that is an expert in data integration and IT infrastructures rather than ms. Amunts.
I would also be interesting to know if the commission subscribes to this concept. There is already a lot of money invested in “e-infrastructures” why one more ? especially such an expensive one.
I also have an impression that Leonid is more into the whole Biology/Neuro stuff so she put a lot of BS about data and IT in her answers hoping that he does not understand/question those parts.
One is sure, however, the HBP is a big success. And always will be. For everyone. Maybe not so for taxpayers but other than that.
LikeLiked by 1 person
Big success? Not as big as it seems. Especially when we are talking about 3D human brain map at 1 micrometer resolution. As far as I know, nobody was able to show it, even a small part of it, never mind the whole human brain.
LikeLike
I like her answers, and it is nice that she answers you. The project goals are so complicated, and so far away, that the risk is very large that the project only becomes a money tool for many groups. But if science is good in anything, it is in improving techniques, and the bold goals and heavy financing should allow a substantial improvement of the methods of brain analysis. I don’t know if Katrin Amunts is a good leader, but her thinking is suitable for one.
LikeLike
Coming to this article a week late because I don’t use RSS feeds. I am glad she wrote back to you. The answers are telling me that this is a catch-all project which can be used to concievably fund any sort of neuroscience research and promote collaboration between laboratories. Obviously it is good for any sort of research to be able to interoperate with other research. The fuzziness of the answers betrays some basic problems with the project’s stated goal of simulating a brain, but I don’t see an outright waste of money here.
LikeLike
The bait-&-switch element is the big concern. The consortium did not rock up to the EU with a research proposal to build a large all-purpose database. If they had done that, would they have been awarded 1 billion euro? Would the EU have considered that to be an outright waste of money?
We do not know, because the consortium began with different goals — ambitious and widely-ridiculed goals — which they abandoned once the moneypipe was connected (relying on institutional inertia to keep it flowing).
LikeLike
Claims of HBP to simulate animal brain or parts of it are over blown,, even with a basic understanding of biology. Example of hype
https://www.theweek.in/news/sci-tech/2018/11/12/The-world-most-powerful-artificial-brain-powered-up.html
.
LikeLike
Pingback: Brain Drain, Moscow to Düsseldorf – For Better Science
Pingback: Who owns the brains in Moscow, Amunts lawyer’s version – For Better Science
Pingback: Graphene Flagship deploys Stripy Stellacci to fight the Coronavirus – For Better Science