Earlier this months, a research misconduct scandal in molecular cell biology broke out in the big news. Yoshinori Watanabe, Japanese researcher of cell division and how cells separate their replicated DNA during mitosis and meiosis, was found guilty of scientific misconduct by his University of Tokyo (read the news here and here). This followed an investigation initiated in the fall of 2016, after anonymous whistleblowers submitted to the university a report accusing 6 Tokyo research groups of data manipulation, first and foremost, Watanabe (I managed to obtain this dossier, and publish it below).
As the outcome of the University of Tokyo investigations, which concluded on May 31st 2017, misconduct was determined in 5 publications from Watanabe’s lab, which appeared between 2008 and 2015 in the elite journals like Science, Nature and Nature Cell Biology. However, the whistleblower dossier lists 7 papers, one of them a paper in Cell from 2015 with duplicated gel images, and a 2011 EMBO Reports paper which contains a western blot which was obviously digitally retouched to remove unwanted bands. Watanabe’s assistant professor Yuji Tanno was also found guilty of misconduct, and indeed his 2015 Science paper with Watanabe looks like a total train wreck of data manipulations. Yet it seems there is a tendency to present Watanabe’s deeds as mere mistakes (though grossly inappropriate ones) by a great genius scientist, who was confused by the complexity of rules on data acquisition and incidentally broke some while producing outstanding and absolutely reliable top-level research. Some of his peers seem to be calling for leniency or at least some understanding for Watanabe. The selected evidence from the whistleblower dossier which I post below suggests that Watanabe knew perfectly well what he was doing, and he did so in order to produce desirable results which his lab experiments failed to deliver, and which he needed in order to impress the choosy elite journals.
The fallen giant
Nature News reported that a decision on Watanabe’s punishment is yet to be made by the University of Tokyo, but:
“As of March, all 15 members of Watanabe’s lab have left, according to a former member who asked not to be named because of the sensitivity of the situation. Watanabe’s major grant — a 5-year, 416-million-yen (US$3.7-million) award from Japan’s science ministry that was due to run until March 2018 — was suspended last March”.
Japan Times provided some background on what happened in Watanabe’s lab:
“But, the report said, inappropriate processing of data was an everyday practice in Watanabe’s laboratory. Watanabe even instructed and taught researchers at his lab how to process data in order to enhance the findings in their papers, the report said.
As for Tanno, the report concluded he was, in part, a victim of Watanabe’s inappropriate instruction, as he had been mentored by the professor since he was a student. Tanno was an assistant professor when the misconduct took place”.
NHK World wrote that the University of Tokyo may be returning Watanabe’s funding, something which no sane European or US university would ever consider, no matter what their scientists did:
“More than 13-million dollars of public funding was used on the 5 papers. The university says it will discuss with the education ministry how much should be paid back”.
Watanabe’s entire career took place at the University of Tokyo. He graduated there in 1984, did PhD 5 years later, was mentored by the dean Masayuki Yamamoto, in 2004 Watanabe became full professor there, aged 43. Now the moustache-wearer brought his academic Alma Mater into the news in a way which may be unacceptable in Japanese culture. And despite his impressive publishing record, he is not yet too big enough to fail, his lab, which faces dissolution, turned the fateful 13 years old.
But how will the scientific community, specifically Watanabe’s international peers react? An update below indicates the cell division researchers are not ready yet to distance themselves from this “giant”, and may even see him as a victim (see. Others are quoted in the Nature News article as if Watanabe’s case is not so bad:
“Watanabe maintained that these manipulations were not intended to deceive and that none undermined the main conclusions of the papers.
Some of Watanabe’s peers who were contacted by Nature accepted that explanation. “I find it impossible to think that these mistakes were dictated by the intention to deceive the scientific community or falsify the data,” says Daniela Cimini, a cell biologist at Virginia Polytechnic Institute and State University in Blacksburg”.
The Whistleblower dossier
Next to the already available education report by the University of Tokyo on the Watanabe case, as well as his own file where the Photoshop master kind of admits some of his allegedly inadvertent errors, I obtained the original whistleblower report from August 29th, 2016. Its available here, the Watanabe information starts on page 11. The dossier is in Japanese, but in most cases the image evidence speaks for itself.
Update: I now obtained also other documents of the same “ordinary_researchers” whistleblower dossier, with evidence on the 6 Tokyo research labs including Watanabe. The files are here, here and here.

These seven Watanabe publications were listed in the whistleblower dossier, I provide some select “illustrations”.
- Yokobayashi S, Watanabe Y. The kinetochore protein Moa1 enables cohesion-mediated monopolar attachment at meiosis I. Cell. 2005 Dec 2;123(5):803-17.
Left image is from the original whistleblower report, also posted on PubPeer. On the right, is Watanabe’s own “correction” which he posted as PubPeer comment one year ago. A duplication of bands is not denied, but this is how Watanabe quotes the editors of Cell on this matter, it fits perfectly to the non-action by Cell on the matter of other Photoshop artists Olivier Voinnet and Maria Pia Cosma at around same time:
“Thank you for your patience while I discussed your request with our editorial team. We went through the material you sent us and it does seem to us that this was a honest mistake in the preparation of panel A of figure 5 of the paper, that bears no impact in the overall conclusions of the paper. In those cases, in particular given also the age of the paper, we would not normally issue a correction, since the benefit of it for the reader is very small. For now, we will keep following the pubpeer post and will re-evaluate the need of a correction if any developments occur.”

2. Yamagishi Y, Sakuno T, Shimura M, Watanabe Y. Heterochromatin links to centromeric protection by recruiting shugoshin. Nature. 2008 Sep 11;455(7210):251-5. doi: 10.1038/nature07217.
The framed image from Yamagishi et al 2008 for Sgo1 Tubulin, as shown in the tweet above, was re-used, but as a stand-in for a totally different experiment, as evidenced by the quantifications which have different SD values. In any case, even if it was the same experiment, how did Watanabe quantify at all, if he only apparently had one single image, or maybe only just one “suitable” one? The University of Tokyo described this example as:
“Fabrication of graph data, fabrication of negative control data with no data input and copy-paste”.
There were other rigged quantifications, with copy-paste of datasets in unrelated context, which Watanabe semi-admitted for Figures 2e, 2f and 3d:
“During preparation of the manuscript, we mistakenly used a different population of cells during quantification, and due to this oversight we did not register the latter cells […]. the results of the quantification are closely similar and our conclusions are unaffected by this error. We have submitted a request to the journal for correction of the strain list.”
Left image is from the original whistleblower report, also posted on PubPeer. On the right, is Watanabe’s own “correction”. Yellow signal indicates the total absence of any background in the western blot panel HP1a outside the two remaining bands. This can only be explained by digital erasure of the other bands, together with the entire background of image, to avoid detection. This is Watanabe’s explanation:
“We have submitted a request to the journal for correction of the strain list. In Fig 4a, western blot panel of HP1α was improperly processed”.
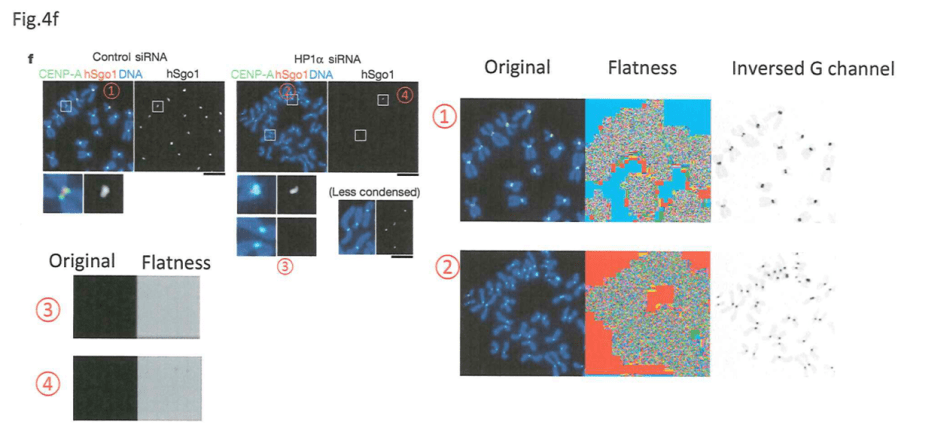
3. Yamagishi Y, Honda T, Tanno Y, Watanabe Y. Two histone marks establish the inner centromere and chromosome bi-orientation. Science. 2010 Oct 8;330(6001):239-43. doi: 10.1126/science.1194498.
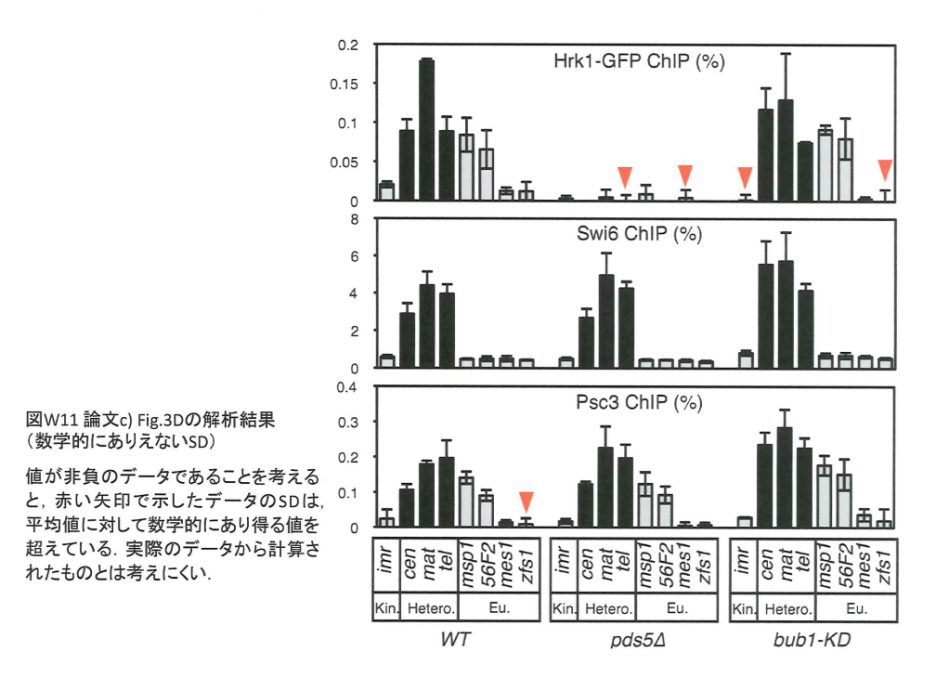
In this regard, Watanabe admitted data rigging for the adjacent Figure 3A in his “Corrections” file:
“In Fig. 3A, two ChIP datasets (ChIP1 and ChIP2) were improperly combined into a single graph. In the corrected Fig. 3A, ChIP1 and ChIP2 are presented separately. These corrections do not affect the description of the results of the specific experiment or the conclusions of the article as a whole. We apologize for any confusion that this error may have caused. Errors
1) Bir1(WT) and H3-pT3/H3(WT) derive only from ChIP1 but not ChIP2.
2) Experiment has not been performed, and should have been indicated as N.D. (not determined).
3) There is a copy error in Swi6(swi6Δ)”.
4. Ishiguro T, Tanaka K, Sakuno T, Watanabe Y. Shugoshin-PP2A counteracts casein-kinase-1-dependent cleavage of Rec8 by separase. Nat Cell Biol. 2010 May;12(5):500-6. doi: 10.1038/ncb2052. Epub 2010 Apr 11.
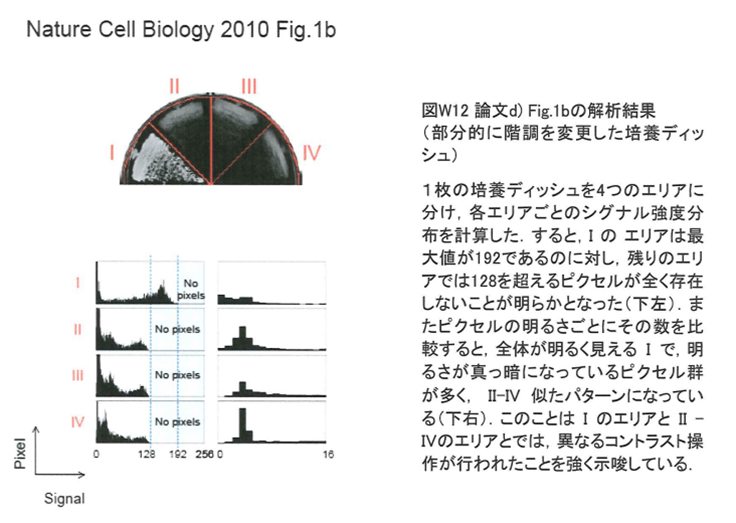
Another example for selective contrast adjustment inside an image, present in the whistleblower dossier, was also uploaded to PubPeer, for Supplementary Figure S1. Here, two yeast dishes labelled with arrow were subjected to a digital signal intensity reduction.
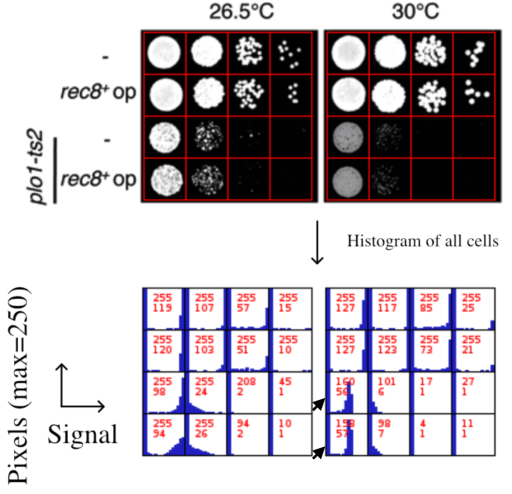
Watanabe commented, with an explanation which beggars belief, especially for a black-and-white image:
“These plates contain Phloxine B, a drug which stains dead cells in red so that the colonies containing dead cells became a bit darker when the pictures were converted to black/while. We apologize for any confusion that this omission of explanation may have caused”.
5. Tada K, Susumu H, Sakuno T, Watanabe Y. Condensin association with histone H2A shapes mitotic chromosomes. Nature. 2011 Jun 1;474(7352):477-83. doi: 10.1038/nature10179.
In this paper, the whistleblower highlighted image manipulation in several western blot panels. Parts of western blot panels were digitally erased, as evidenced by the rectangular monochrome patches lacking any background pattern (left image). On the right, the University of Tokyo demonstrates how one of these panels was adjusted to remove several bands (translations mine).
There was even more blot doctoring, especially in Supplementary figures, as the image on the left below shows, from the whistleblower dossier. On the right, is Watanabe’s Corrections file, where he shows the original gel images, with the extinct bands present:
Watanabe’s explanation, together with a proud “We have received an affirmative reply from Nature“:
“In Figs. 3e, 3g, 5a and S16, the western blot panels were not properly processed. We show the corrected panels, which have been reconstructed from the raw data files, are shown here. These corrections do not affect the description of the results of the specific experiment or the conclusions of the article as a whole. We apologize for any confusion that this error may have caused”.
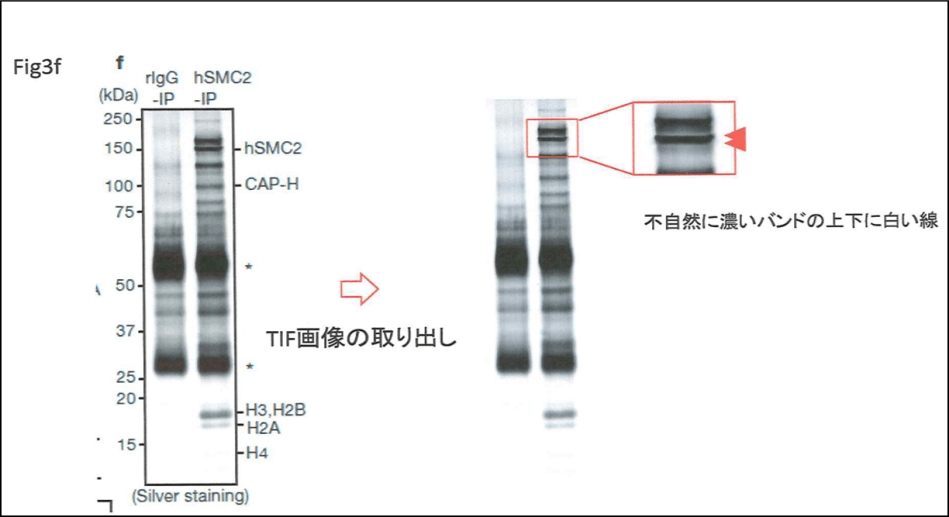
Of course just removing gel bands is no challenge for the Photoshop master Watanabe. He is so skilled, he can even make bands appear where there are none, like in this Figure 3f. A band for the protein hSMC2 was apparently stuck onto the gel image, where a control immunoprecipitation was compared to that with a anti-hSMC2 antibody. Does this affect “the conclusions of the article as a whole”? Probably depends on whom you ask, a cell biologist, or a journal editor-in-chief.
6. Kagami A, Sakuno T, Yamagishi Y, Ishiguro T, Tsukahara T, Shirahige K, Tanaka K, Watanabe Y. Acetylation regulates monopolar attachment at multiple levels during meiosis I in fission yeast. EMBO Rep. 2011 Oct 28;12(11):1189-95. doi: 10.1038/embor.2011.188.
Which brings us to the next Watanabe paper, Kagami et al 2011, published in EMBO Reports. This is the only one corrected so far, Watanabe even proudly displayed a letter from EMBO Press Editor-in-Chief Bernd Pulverer, where a minor correction was offered to him on May 1st 2017. The correction indeed appeared on July 3rd. This is what it looked like:
“In Figure 2A, due to over-contrasting of the top two Western blot panels (AcPsm3), there is a loss of signal, in particular in the top right panel. Further, the right two panels were derived from the same blot, which was stripped and reprobed. The left two panels were derived from different blots of the same sample. This limits the quantitative information content of the published figure, but it does not change the conclusions derived from the experiment. In the corrected figure, the two left panels are now assembled from the same blot that was stripped and reprobed, and the contrast is properly adjusted in all panels. All these blots were processed in parallel in the course of the same experiment”.
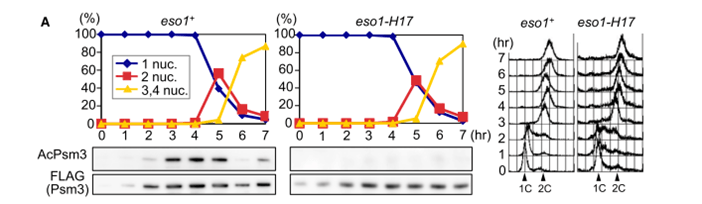
This was the real problem with that western blot, according to whistleblower dossier: the bands in the left AcPsm3 panel were obviously digitally erased, not just blanked out by “over-contrasting”:
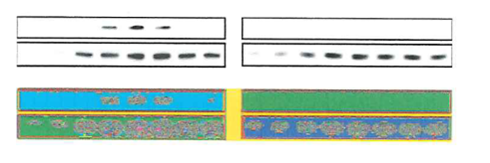
Back in May 2017, Pulverer write to Watanabe:
“Regarding fig 2A, we conclude that the contrast is overly accentuated in the Western blot panels in the left part of the figure, which leads to a loss of linear signal, limiting the quantitative information that can be derived from the images.
You also rightly noted that there is no apparent signal in the top right hand Western blot panel of fig 2A. This may be due either to the complete absence of experimental data or images that were over-contrasted to the point where all signal is lost at the resolution of the published image. Either way, panels with no detectable data are not informative. In this case, our view is that the problem is most likely either due to a mistake or to poor image processing, rather than intentional falsification[…]
We recommend issuing a corrigendum primarily to point to the fact that the top right panel lacks signal and to show the We recommend issuing a corrigendum primarily to point to the fact that the top right panel lacks signal and to show the correct panels. As part of this corrigendum, we would also invite you to note that the Western blots in fig 2A were derived from the same samples, run at the same time, but that these were concatenated to assemble the published figure. This resulted in differential contrast and exposures in the published figures, limiting the quantitative information content of the resulted in differential contrast and exposures in the published figures, limiting the quantitative information content of the figure, but not per se the conclusions derived”.
So the acetylation of the protein Psm3 is not transient and restricted to the event of cell division (as originally suggested by Watanabe in his EMBO Reports paper), but, considering the erased bands and the loading controls, probably present at all times, possibly even in similar levels. Is it too pedantic to claim that this may be a significant change of story? Pulverer reiterated in his letter however:
“Thus, our view is that the basic conclusions of this figure, and therefore the paper as a whole, stand”.
What is the point of peer review then, if the editors know it better anyway? I confronted Pulverer with the whistleblower’s evidence, after some back-and-forth emails he declared:
“I have to admit that at the resolution of the image you sent we see no ‘blue patches'”.
Well, I am slightly colour blind, but I see blue quite well, and hope my readers do also. Pulverer then provided to me this statement to be quoted:
“We received what the authors claimed to be source data for the top left panel. We also received three other blots that it is claimed show the same experiment – they show overall similar effects (time points 4 and 5 show relatively enhanced acetylation).
We compared the SD to the published figure and saw sufficient similarities to convince us this was the same data.
We applied contrast changes across the whole image and concluded that the weak bands could be ‘contrasted out’ without selective deleting them. This require extreme and non-linear contrast changes and it is in our view a case of severe beautification.
We concluded that the overall effect was apparent in the source data and three replicates and that the conclusions therefore stand as far as we can judge based on the evidence provided.
We also conclude that a corrigendum is necessary to show the unmanipulated figure”.
Watanabe is incidentally member of EMBO Reports Advisory Board. Meanwhile, Watanabe is scheduled as invited speaker at the EMBO conference on meiosis, taking place this month in Croatia.
Update 11.08.2017. Iva Tolić, one of the organisers of the EMBO Meiosis meeting, wrote to me when I asked if Watanabe will be appearing as scheduled:
“We are still considering this issue. The investigations of the University of Tokyo are not concluded and final statements have not yet been released. We share your passion for keeping science true, but we also think that Dr. Watanabe should be allowed to present his case”.

7. Tanno Y, Susumu H, Kawamura M, Sugimura H, Honda T, Watanabe Y. The inner centromere-shugoshin network prevents chromosomal instability. Science. 2015 Sep 11;349(6253):1237-40. doi: 10.1126/science.aaa2655.
Maybe Watanabe can present his Science paper from 2015 as a case to his defence. It is so excessively manipulated that the EMBO workshop participants will sure have a field day in Croatia debating it. Good thing that University of Tokyo’s misconduct verdict is apparently not valid for Watanabe’s peers (it is only the disciplinary measures which are “not concluded” yet). But here for example is the Supplementary Figure S16 from Tanno et al 2015, as highlighted by the whistleblower dossier. It has same monochrome patches proving erasure of data.

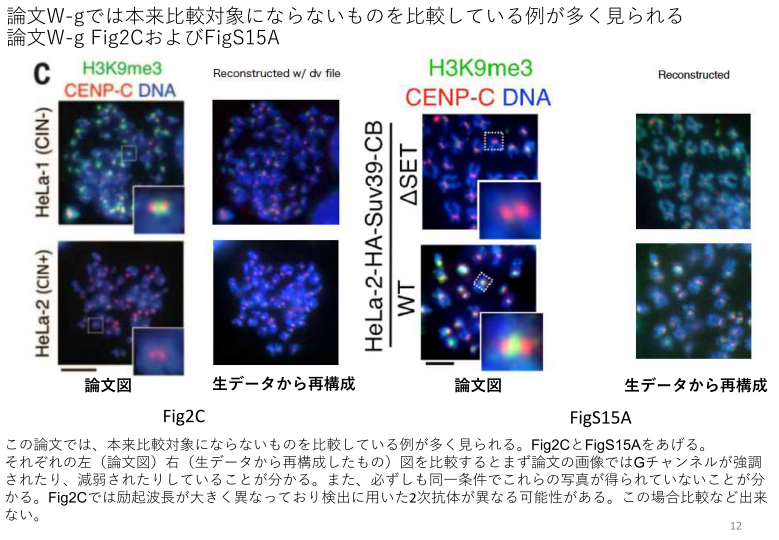
And again Watanabe was playing with different settings inside same experiment. Masses of immunofluorescence data is completely unusable after it was found out that these images were deliberately acquired under different microscopy settings. For Watanabe merely a tool to normalise the signal, a method we saw already with Sonia Melo and her colleagues:
“The differences in exposure time compensate for differences in the intensity of staining between independent non-treated samples”.
This is no way to do science, a graduate student learns fast enough the importance of producing all images which are to be compared with each other under exactly same experimental and image acquisition conditions. To assume Watanabe did not know this is ridiculous. Of course he knew he is manipulating data, but he needed results to present to Science.
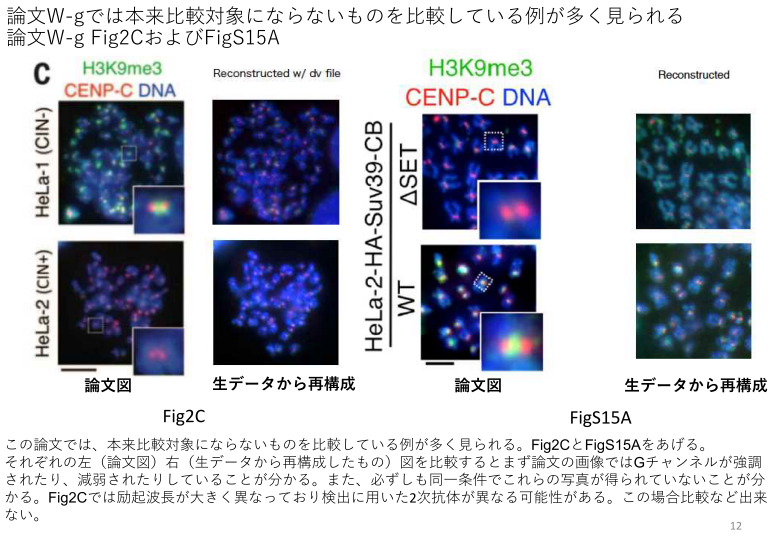
Watanabe’s commentary:
“In figs. 2C, S13C and S15A, the representative images for comparison were captured under different imaging conditions. In fig. S8B, S11F, S12B, the images used for compared were not adjusted using identical settings. In fig. S8A, the dot blots were mislabeled and not properly adjusted for contrast. We show corrected images or explanation here. Given that quantifications were performed using the original images, we believe that the changes in the representative images do not affect the conclusions of the paper. However, as this paper has a number of errors, we are contacting the journal to inquire whether extensive correction or retraction is more appropriate. We apologize for any confusion that these errors may have caused”.
This last paper was where Watannabe apparently became so secure in his data manipulation impunity that there probably hardly is any figure unaffected by fraud. Of course also here the quantifications were rigged. In Figure 4A, all error bars look exactly the same, and so do some datasets, labelled with arrows.
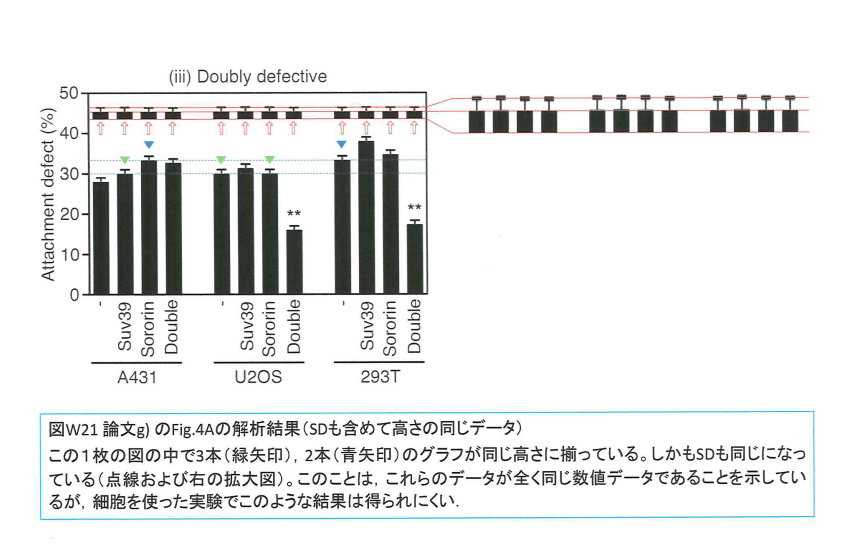
The Tanno et al Science 2015 will probably be retracted, even Watanabe seems to have accepted its fate. What about his other manipulated works? And are his cell biology peers prepared to forgive him, because he is supposed to be a “giant”?
Update 8.09.2017. The paper Tanno et al 2015 was retracted by Science on September 5th.
Update 2.05.2018. Watanabe was sacked by University of Tokyo in April, according to news reports. Press release here.

Donate!
If you are interested to support my work, you can leave here a small tip of $5. Or several of small tips, just increase the amount as you like (2x=€10; 5x=€25). Your generous patronage of my journalism will be most appreciated!
€5.00











How such an individual can be considered a giant of anything other than data fabrication beggars belief. The usual defence “work has been replicated” is not a defence. Most science has to be incremental – regardless of the hype of novelty, etc. touted by journals and so contaminating papers and authors – journals are entirely responsible for this. Consequently, it is pretty easy to take a reasonable guess as to what the unknown is. Then you can make up the data, claim a great first, and become a giant.
So once again, a cheat exposes how the reward system that requires publishing in ‘glam’ journals distorts science. Sacking or busted back to untenured postdoc would be appropriate actions for this scale of fabrication, but this is not going to happen.
LikeLike
Dear Ferniglab,
How many times we are watching the same movie, again and again? The names might change…either Watanabe, or Voinnet….or Melo…but the move is always the same.
I was expulsed from the lab where I did my postdoc….because I did not like this movie….honestly I think scientists can do much better, openly, much more effectual to society and with fully integrity….and still earn good money….
LikeLike
Hi Leonid
Isn’t it in Japan all abut saving face? What do these relevations about his “deeds” mean in this regard?
Second, and more fundamental, if certain “corrections” and “errors” “do not affect the conclusions of the papaer”, then why were they included in there in the first place?
Inquiring minds want to know.
Oh, btw, I can only hope he gets a warm welcome in Croatia (pitch forks and tar+feathers seem appropriate)
Cheers oliver
LikeLike
Pingback: Boletim de Notícias: SBPC apoia Diretas Já | Direto da Ciência
So crazy world!!
FYI. The software MOTUIN (http://motuin.org) may have some help to spot those forgery. It seems to me that PUBPEER has some users using it to report “bad science”, for example https://pubpeer.com/publications/0953E23528C31ADF4F3FF85EEB77AC#undefined
LikeLike