Bert Vogelstein is God. This Johns Hopkins University professor has cured cancer ages ago, so far only in mice, but not yet in humans. Those who also managed to cure cancer in mice in Vogelstein’s lab become big professors themselves. Like Kenneth Kinzler, who remained at Johns Hopkins, or Heiko Hermeking, who returned to Germany to become professor at the Ludwig Maximilian University (LMU) in Munich.
But some papers by Vogelstein, Kinzler and especially Hermeking’s were questioned on PubPeer. A harassment campaign? Or rogue collaborators and PhD students having been sloppy? Both?
Vogelstein and Kinzler never replied to my email inquiry. Hermeking told me that Vogelstein suspects to be behind those baseless comments his envious peers, whom he intends to take legal action against. And to me personally, Hermeking issued this warning:
“…if you post/publish unfounded, reputation-damaging claims, which hinder me and my colleagues in carrying out our research, there any ways to respond to this since yesterday“.
The reference in this context went obviously to an earlier German Supreme Court decision about hate-postings against politicians. So I went and tried to debunk the PubPeer allegations on Vogelstein’s and Hermeking’s papers. Here we go.
The evidence on PubPeer is mostly years old, starting 2015. But it all came up after Clare Francis reminded Nature’s editor Barbara Marte about an old paper by Vogelstein, Kinzler and Hermeking.
Timothy A. Chan, Heiko Hermeking, Christoph Lengauer , Kenneth W. Kinzler, Bert Vogelstein, 14-3-3Sigma is required to prevent mitotic catastrophe after DNA damage Nature (1999) doi: 10.1038/44188
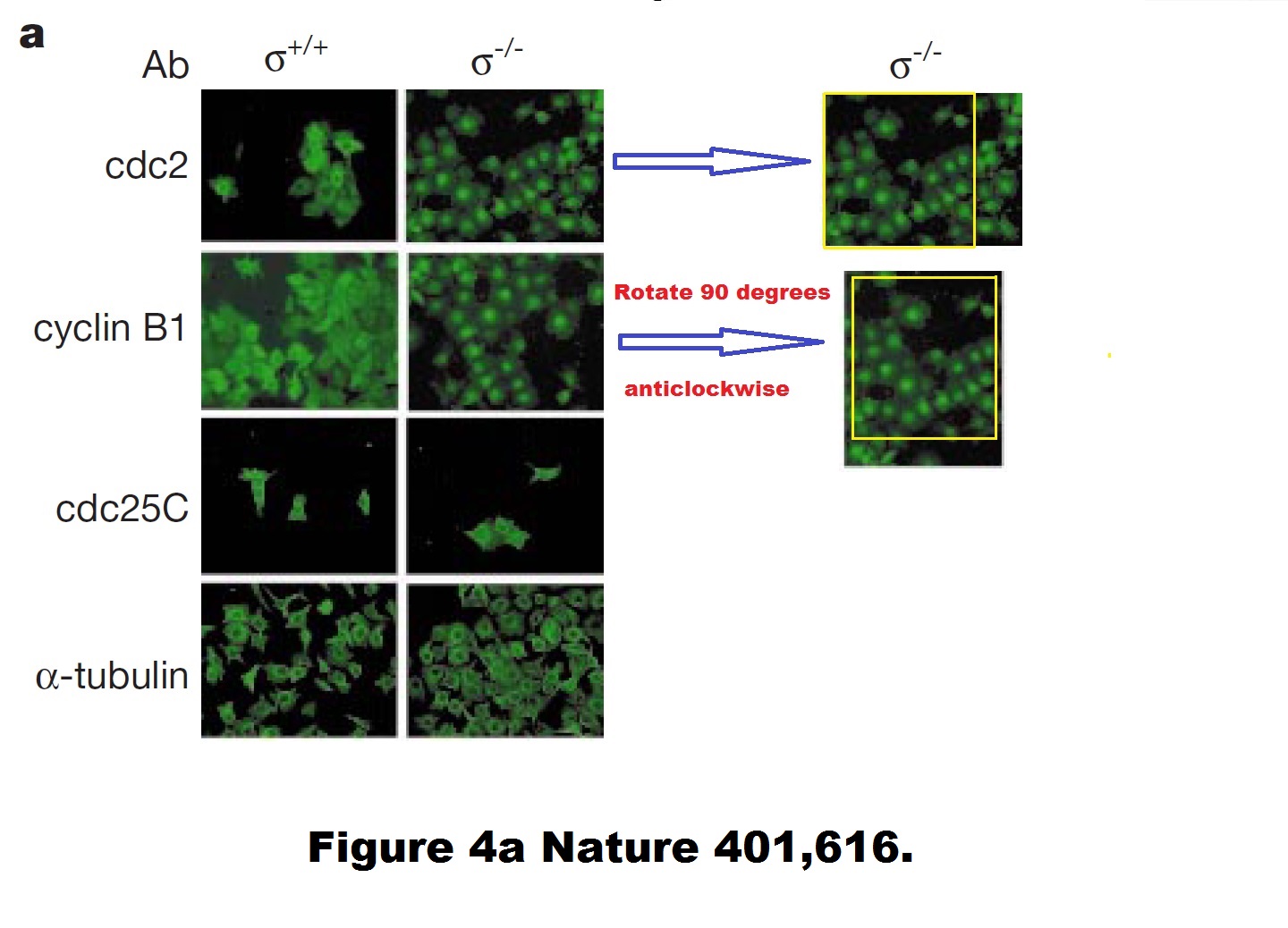
A duplicated and rotated image? Never! Hermeking explained to me:
“…as outlined in the responses to the Nature 1999 paper on PubPeer, CDC2 and cyclin B1 have the same staining pattern, i.e. they co-localize. Since green was always used as the color in the figure (with LSM-IF you get black and white pictures that are recolored), it looks as if the same image was used twice.
Unfortunately, the PhD student used the same image section of a double staining for CDC2 and cyclin B1 and rotated the photograph of the latter 90 degrees. He therefore did not use the same photo twice.“
That PhD student, Hermeking later explained, was Timothy Chan, now head of an oncology centre at Case Western Reserve University in Ohio.
But still, how can two different antibody stainings produce a virtually pixel-identical result? And even if such molecular biology miracle did happen, why are the cdc2/Cyclin B1 stainings different for +/+ cells? Hermeking was kind enough to teach me some basic cell biology:
“As you surely know, cyclin B1 and CDC2 bind directly to each other. Therefore, co-localization is to be expected.
As far as I remember, Mr. Chan was asked by Prof. Vogelstein to check this in the original images and he has confirmed that these are 2 different images. When staining the +/+14-3-3sigma cells, Mr. Chan selected two different sections.“
Such amazing 100% colocalisation of two antibodies for two different proteins, in immunofluorescence no less! Other people would never get an identical staining even when using two different Cyclin B1 antibodies. But then again, such failed scientists won’t ever succeed in Professor Vogelstein’s lab, like Professors Chan and Hermeking succeeded. Science is not for everyone.
There as another issue in that paper. Elisabeth Bik found something:
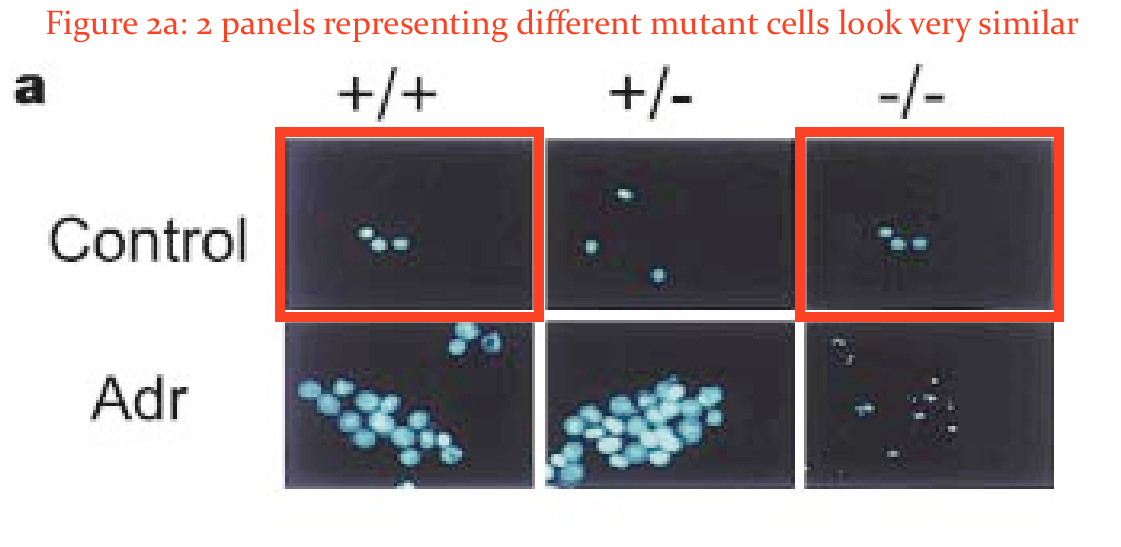
In addition, would the two panels in Figure 2A marked here with red boxes expected to look so similar?”
Hermeking was unimpressed by Bik’s amateurish and unqualified attempts to spot duplications:
“If you zoom a lot into Fig. 2A, you can see, in my opinion, that it’s not the are the same cell nuclei.
These are the results and figures by Timothy A. Chan.“
Regarding that 1999 Nature paper, Hermeking also declared to me:
“I would also like to note that since the comments are anonymous, the PubPeer Forum is unfortunately sometimes used to express personal opinions on the research of others that are not purely factual in nature. Whatever the motivation. This unobjective criticism is also expressed in the discussion of the Nature Paper from 1999 (Chan et al., position 2 in your list) and this is discussed/criticized at length by other participants (I myself have never made a contribution/ an answer in this forum because colleagues advised me against it). In my opinion, it is well known that not all studies on the same topic come to the same conclusion. In this case, an author with a publication on the function of 14-3-3sigma that had a different/different result appears to be using the PubPeer forum to discredit authors and their co-authors who disagree with him. This is the common conclusion which the authors of the Nature 1999 publication came to. The last author, Prof. Bert Vogelstein (Johns Hopkins Medical School Baltimore, USA), already proposed legal action against these comments a few years ago, as they are obviously not objective. So far we have refrained from doing so.“
Legal action? Against whom? Hermeking swiftly warned me that he will sue me for so-called hate-posting, see the quote above.
The pseudonymous Clare Francis, unaware of these threats, kept complaining to Nature. By mistake, the editor Marte hit reply-all instead of forward:
“Just what I needed before the weekend. So she (which I think is a he) is back. I think I handled this paper. 1999. I don’t believe at all that they had contacted us, because as you know I take these things seriously. They do look very much like duplications. PubPeer also refers to papers that couldn’t reproduce the findings. Oh lord…. I doubt it’s on ejp Vic and I will look into it. Barbara“
Marte later announced to Clare Francis and to me to investigate this paper. Good luck!
But Vogelstein doesn’t need Nature. He published two papers in Science which revolutionised cancer research, or so we are expected to think. The one from 7 years ago, Tomasetti & Vogelstein 2015, proclaimed the new wisdom on where cancer really comes from:
“….random errors occurring during DNA replication in normal stem cells are a major contributing factor in cancer development. Remarkably, this “bad luck” component explains a far greater number of cancers than do hereditary and environmental factors.”
The older you get, the more often your stem cells divide, thus the higher probability for cancerous mutation. A lot of scientists laughed at this weird and silly assumption, so to shut them up, Vogelstein issued a follow-up paper, also in Science, Tomasetti et al 2017, where he expanded on his theory, that two thirds of all cancers are indeed caused by bad luck of random genetic mutations only. Nobody will dare to argue with TWO Science papers!
Still, scientists were not prepared to accept the gospel because it was too silly. Both papers were rightly criticised by many peers, some of defunct PubMed Commons criticisms are preserved on PubPeer, here and here.
Now, I don’t know what exactly Vogelstein meant to convey. It seems he is keen to sell personal cancer-risk diagnostics, being involved with many biotech companies, as his conflict of interest statement reveals:
“B.V. is on the scientific advisory boards of Morphotek, Exelixis GP, and Sysmex Inostics, and is a founder of PapGene and Personal Genome Diagnostics. Morphotek, Sysmex Inostics, PapGene, and Personal Genome Diagnostics, as well as other companies, have licensed technologies from JHU on which B.V. is an inventor.”
But I am pretty sure certain other industries will also love Vogelstein’s teachings. See, cancer is actually not really caused by smoking, asbestos, radiation, environmental pollution, workplace hazards etc – it’s just bad luck. Science has spoken, twice.
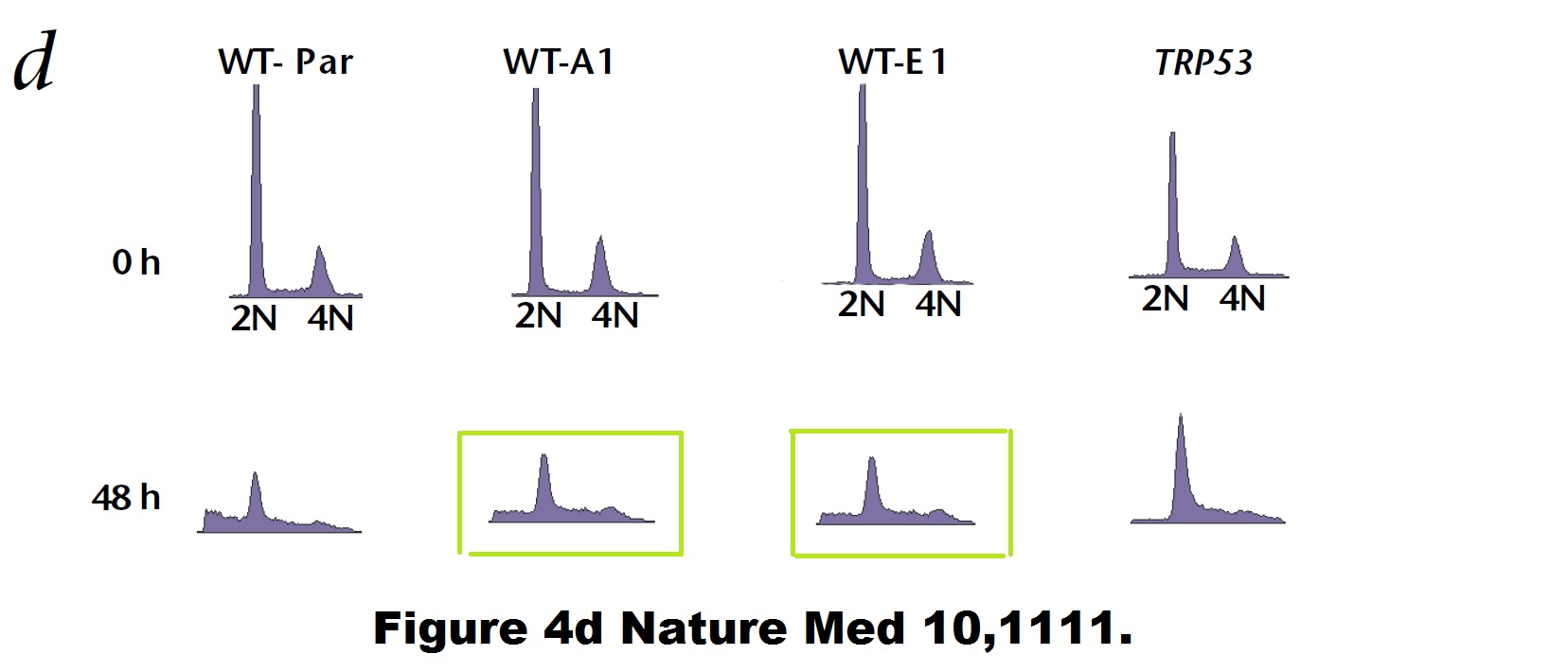
Bad luck is also what befell some of Vogelstein’s papers. Like a duplicated flow cytometry plot here:
Paul M. Hwang , Fred Bunz , Jian Yu , Carlo Rago , Timothy A. Chan , Michael P. Murphy , Geoffry F. Kelso , Robin A. J. Smith , Kenneth W. Kinzler , Bert Vogelstein Ferredoxin reductase affects p53-dependent, 5-fluorouracil–induced apoptosis in colorectal cancer cells Nature Medicine (2001) doi: 10.1038/nm1001-1111
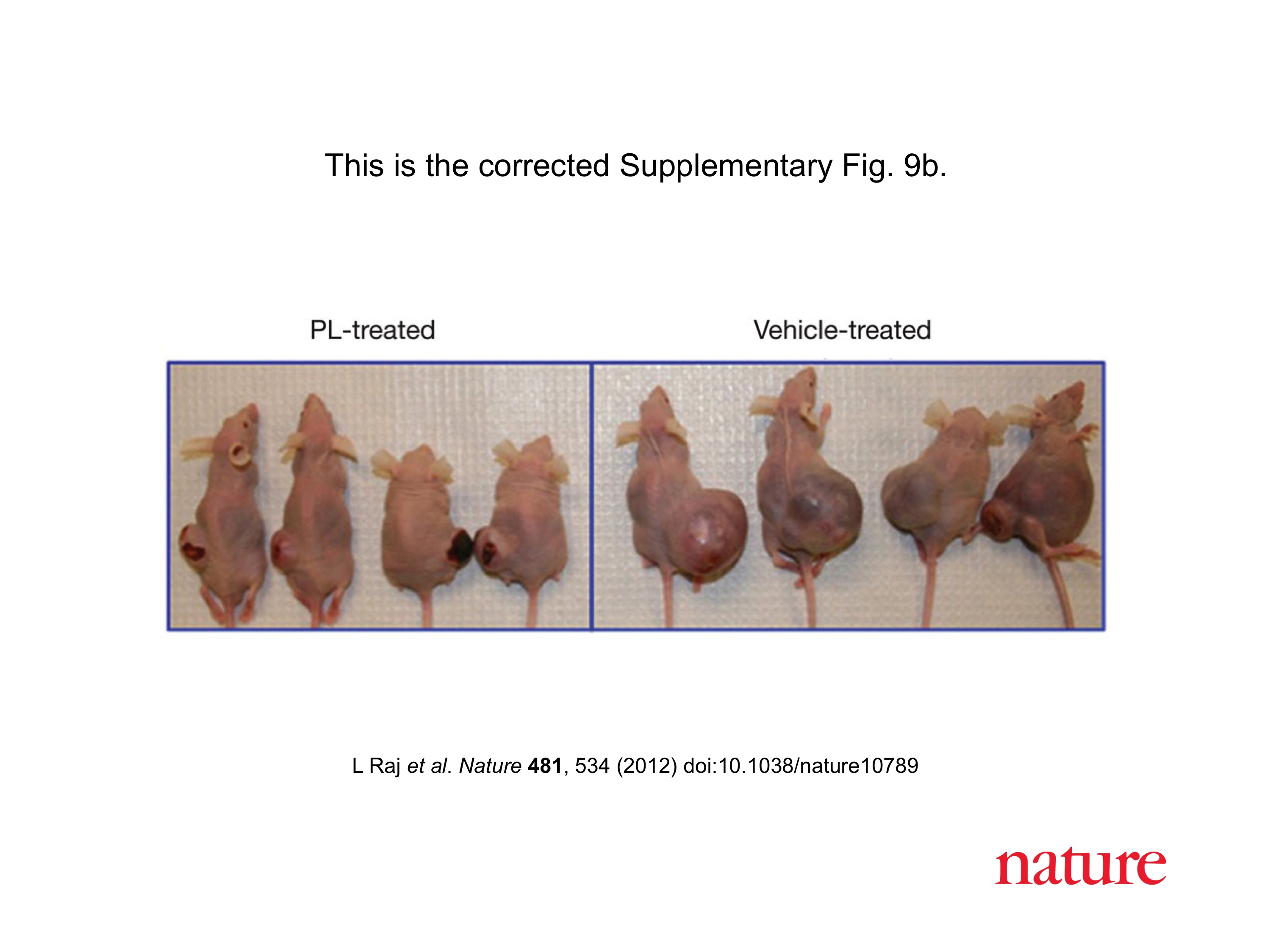
Bad luck was also what these mice had. Apparently, Vogelstein and Kinzler are such big stars of cancer research, that Johns Hopkins University allows them to do to their mice what pedestrian scientists are not allowed to do, due to concerns about animal welfare and cruelty.
B. H. Park , B. Vogelstein , K. W. Kinzler Genetic disruption of PPARdelta decreases the tumorigenicity of human colon cancer cells Proceedings of the National Academy of Sciences (2001) doi: 10.1073/pnas.051630998
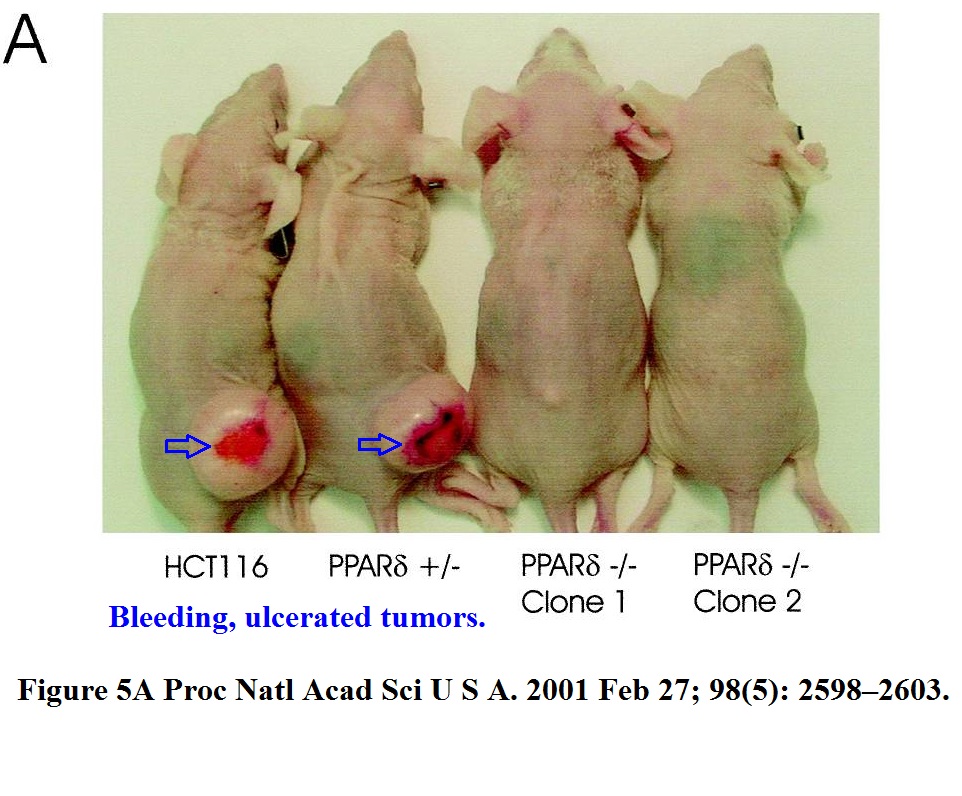
These immune-deficient nude mice suffered a lot, but it was not in vain. They were tortured for a paper which the National Academy of Sciences member Vogelstein “contributed” to PNAS. The study was namely so groundbreaking he chose his own peer reviewers, who apparently also agreed, that yes, the mice got what they deserved.
Let’s look at some strange gels instead.
Yardena Samuels , Luis A. Diaz , Oleg Schmidt-Kittler , Jordan M. Cummins , Laura Delong , Ian Cheong , Carlo Rago , David L. Huso , Christoph Lengauer , Kenneth W. Kinzler , Bert Vogelstein , Victor E. Velculescu Mutant PIK3CA promotes cell growth and invasion of human cancer cells Cancer Cell (2005) doi: 10.1016/j.ccr.2005.05.014
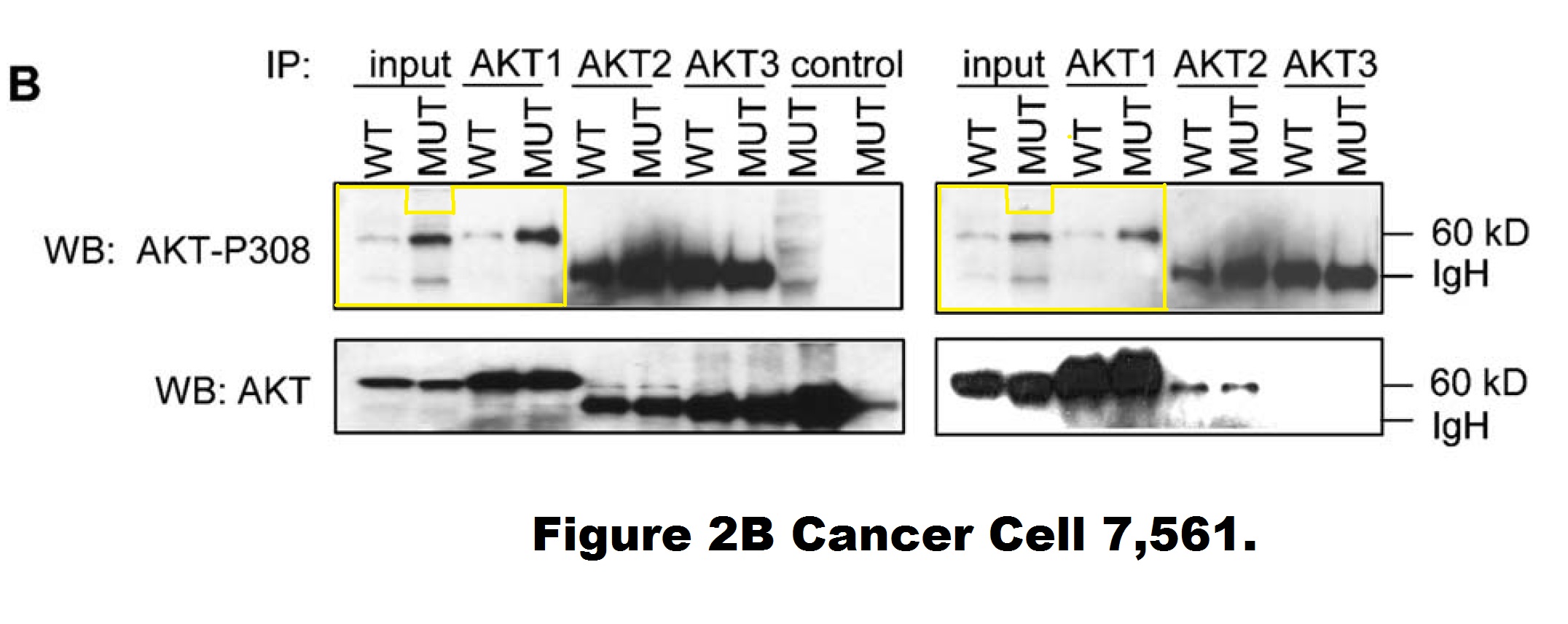
A closer look into this figure reveals that the entire right-hand phosho-AKT gel (showing DLD1 cells) is quite possibly a shorter exposure of the left-hand phosho-AKT gel (showing HCT116 cells). Those are likely two separate image acquisitions of the same blot (of unclear origin). It is difficult to explain as a mistake of oversight, since those are supposed to be different cell lines. Moreover, the HCT116 panel contains two extra two lanes (“control MUT”) on the right, the last one of which appears digitally spliced on. Vogelstein and his academic offspring at Johns Hopkins, the group leaders Kinzler and Velculescu did not reply to my email when asked to explain.
But Hermeking replied. He even provided me with a pdf addressing the PubPeer criticisms of his papers. Here is the file, I will quote from it below:
Once, Vogelstein, Hermeking and their colleagues even had to issue a correction over spliced gels. It was an interesting one, if one looks closely.
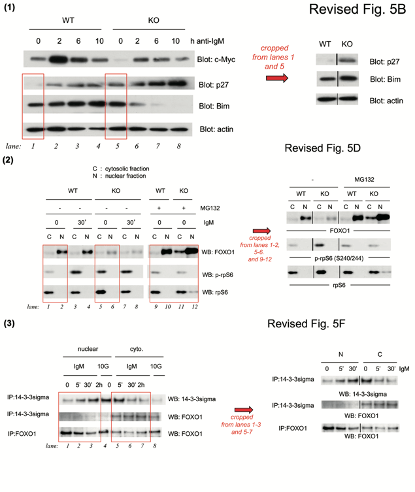
Yu-Wen Su , Zhenyue Hao , Atsushi Hirao , Kazuo Yamamoto , Wen-Jye Lin , Ashley Young , Gordon S Duncan , Hiroki Yoshida , Andrew Wakeham , Philipp A Lang , Kiichi Murakami , Heiko Hermeking , Bert Vogelstein , Pamela Ohashi , Tak W Mak 14-3-3sigma regulates B-cell homeostasis through stabilization of FOXO1 Proceedings of the National Academy of Sciences (2011) doi: 10.1073/pnas.1017729108
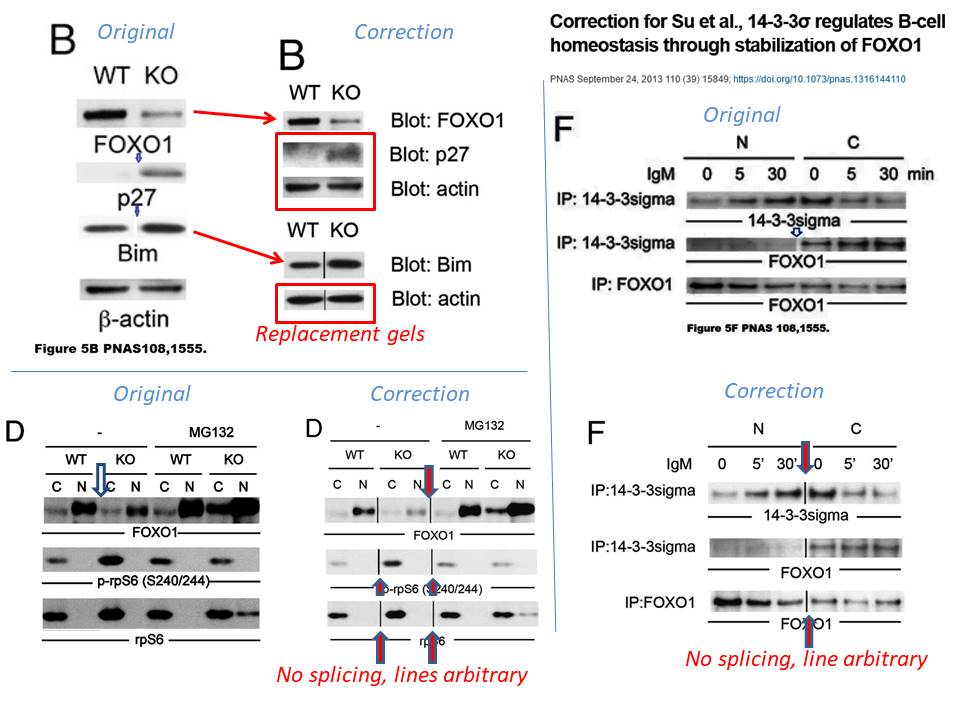
The correction’s text was not exactly honest. First of all, half of the data in Figure 5B was stealthily replaced, it is not clear why those old bands were removed and new results produced from nowhere. Well, it is clear, actually. The authors’ problem was: they could not just admit the irregular gel splicing, because it would render the entire figure meaningless and raise reasonable suspicions of data manipulation. In Figures 5D and 5F, the blots did remain same, but splicing lines were added. Yet not just where that splicing actually occurred, but also in other positions, where the gels were not spliced at all, if my own eyes are to be trusted. But with al these extra divider lines, the trickery looks more or less innocuous, and not at all like data fudging. Clever!
In any case, Hermeking states that these experiments were done by a “postdoc from the lab of Tal Mak in Canada“, i.e. the University of Toronto professor Tak Mak who also contributed this paper to PNAS as Academy member.
Update 10.02.2022. Now Tak Mak sent this postdoc to threaten me with a lawsuit also. This message just arrived, with Mak in cc, from Yuwen Su, now PI at National Health Research Institutes in Taiwan:
“My name is Yuwen Su, and I am the first author who performed the experiments of the PNAS paper you mentioned. First of all, I thank you for the great interest for reading the paper and for points out my correction. The experiments were done when I was a postdoc. fellow in Mak’s laboratory.
The corrected figures were repeatable with the same conclusion. The related original figures were sent to editor when we did the correction. To response your points: (1) The lower panel in the revised 5B was spliced from the same gel in one experiment from lanes 1 and 5 of the original gels. (2) The lines in revised 5D were cropped from lanes 1, 2, 5, 6, and 9 to 12. (3) The lines in revised 5F were cropped from lanes 1, 2, 3, 5, 6, and 7 of the original gels.
I would appreciate that if you withdraw the false allegations about the results. We reserve the right to take legal action, if you keep spread the false statement.“
The above low-resolution 128 kb image was attached. Not even of the full-size scans of original gels, but of a PowerPoint with dividing lines drawn and exported as a blurry image, which is pointless to scrutinise. The published Figure 5 and its correction have a much higher resolution. The paper was 2 years old when the correction was issued, is that all they have left of the raw data? And Su is confused, the revised figure 5B of the published correction is certainly not exactly what her picture shows, legal action or not.
Why do these scientists respond to reasonable criticisms of their works only with threats of lawsuits? Shouldn’t their science speak for itself? Didn’t Mak himself once accuse a colleague of “cheating”, and now he sends his first author to threaten me with legal action for failing to see in the original published figure the splicing lines where they drew them?
In yet another correction, it was also a collaborator. A problematic flow cytometry figure had to be corrected, which Hermeking states was done at the Cornell University, meaning in the lab of Alexander Nikitin.
Chieh-Yang Cheng , Chang-Il Hwang , David C. Corney , Andrea Flesken-Nikitin , Longchang Jiang , Gülfem Meryem Oner , Robert J. Munroe , John C. Schimenti , Heiko Hermeking , Alexander Yu. Nikitin miR-34 cooperates with p53 in suppression of prostate cancer by joint regulation of stem cell compartment Cell Reports (2014) doi: 10.1016/j.celrep.2014.02.023
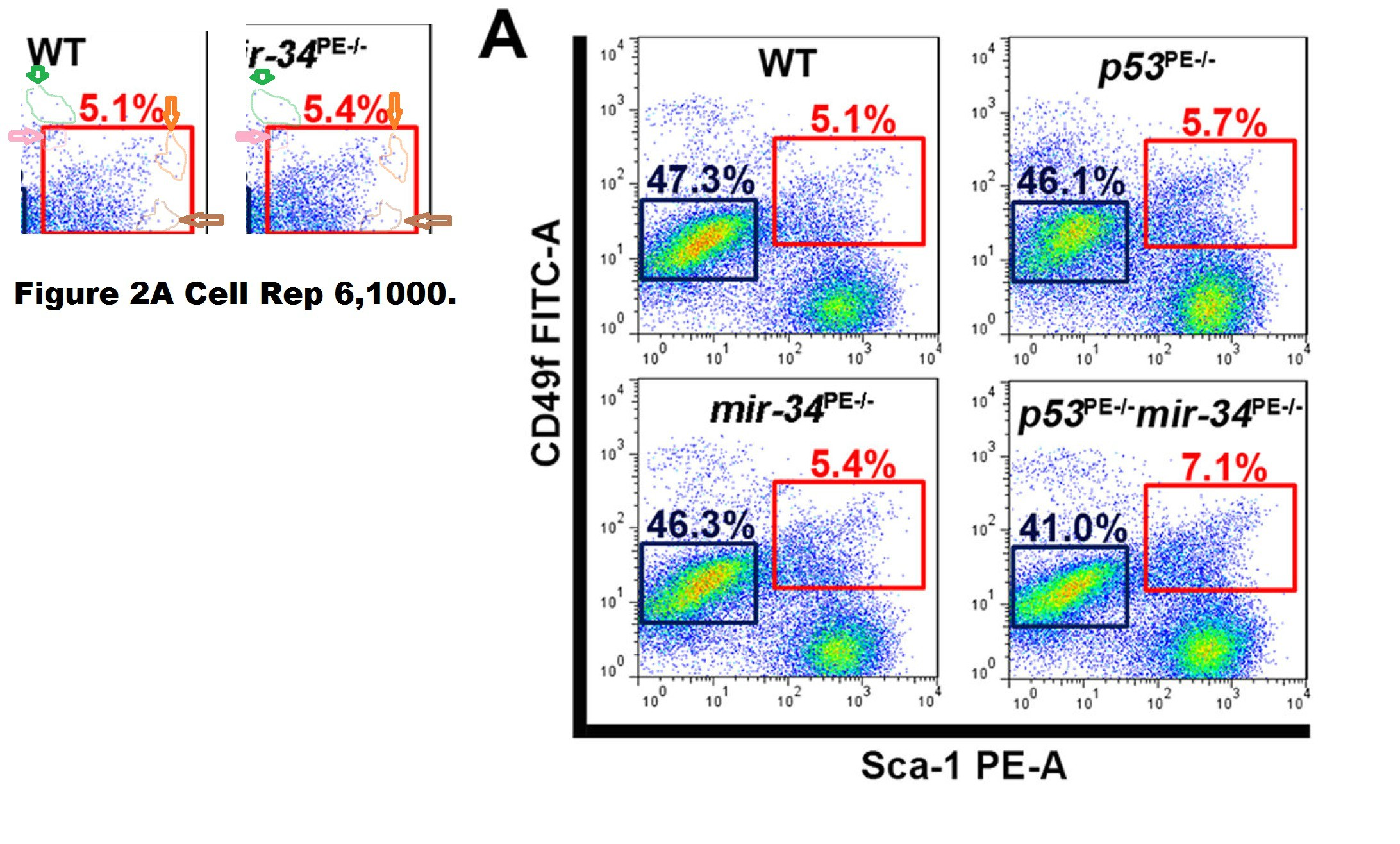
The Correction issued in 2015 declared:

“In Figure 2A of the originally published version of this paper, FACS plots for the wild-type and mir-34PE−/− samples were inadvertently switched during preparation. The corrected Figure 2A is provided below and now appears with the corrected version of the manuscript online. None of the conclusions in our manuscript are affected by this error.“
The “WT” FACS plot was replaced in the correction. Thing is, it was not the case that the mir-34PE−/− plot had been just accidentally copied on top of the WT one. The two FACS plots show the same measurement but plotted slightly differently, i.e., with slightly different gates which changes the overall appearance but retains much of the cell pattern. How can such things happen by mistake?
Nikitin protested that I should not believe my own lying eyes:
“All gate windows are the same in all images. All numbers were accurate in our original iterations of the figure. Unfortunately, during reformatting the figure for another journal, the correct WT image was inadvertently replaced with a wrong one, while the original, accurate numbers remained. During preparation of our erratum, all issues were discussed with journal editor and all documentation to substantiate our explanation was provided.
Your efforts to keep science accurate and reliable are certainly appreciated. However, it seems focusing on mistakes that directly impact data conclusions may better benefit research community.“
But I am trying, Dr Nikitin! Here, look, I wrote about your senior colleague, the Weill Cornell dean Augustine MK Choi:
But how does one know which mistakes affect the conclusions and which don’t? Maybe the authors should publish a disclaimer at the end of each paper which figures are important (if any) and which are just filler illustrations? Pictures with no other purpose but to lighten up the textual monotony of scholarly treatises?
Is the Figure 1b of this paper from Hermeking’s own lab in Munich just decoration or part of the conclusions?
Gerold Untergasser , Heike B Koch , Antje Menssen , Heiko Hermeking Characterization of epithelial senescence by serial analysis of gene expression: identification of genes potentially involved in prostate cancer Cancer Research (2002) issn: 0008-5472

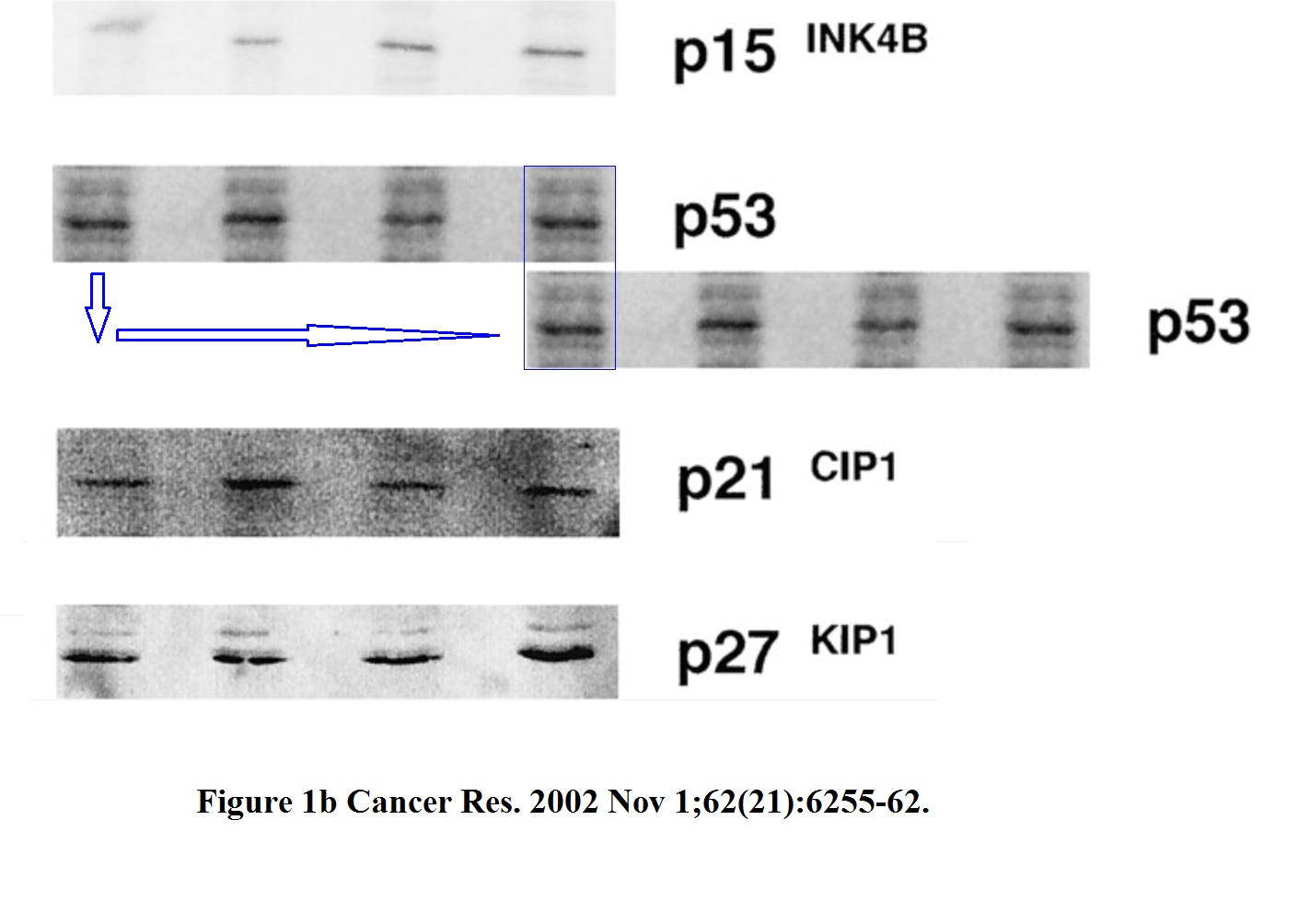
The two highlighted p53 bands (passage 5. and 8.) are too similar, such unexpected similarity in both the both the band shape and the background signal is highly unlikely to arise by chance. The minor pixel differences can be explained by image compression, while the overwhelming similarities cannot be explained at all.
But Hermeking disagrees:
“Gel bands similar, but not identical”
Hermeking declared exactly same for the following paper:
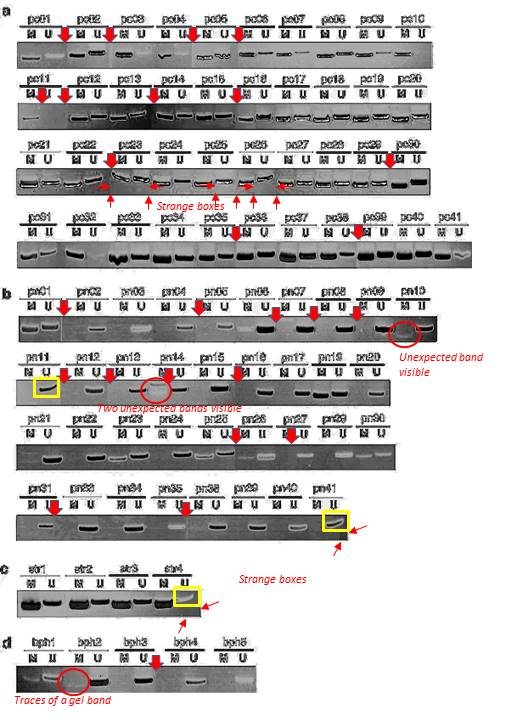
Dimitri Lodygin , Joachim Diebold , Heiko Hermeking Prostate cancer is characterized by epigenetic silencing of 14-3-3sigma expression Oncogene (2004) doi: 10.1038/sj.onc.1208004
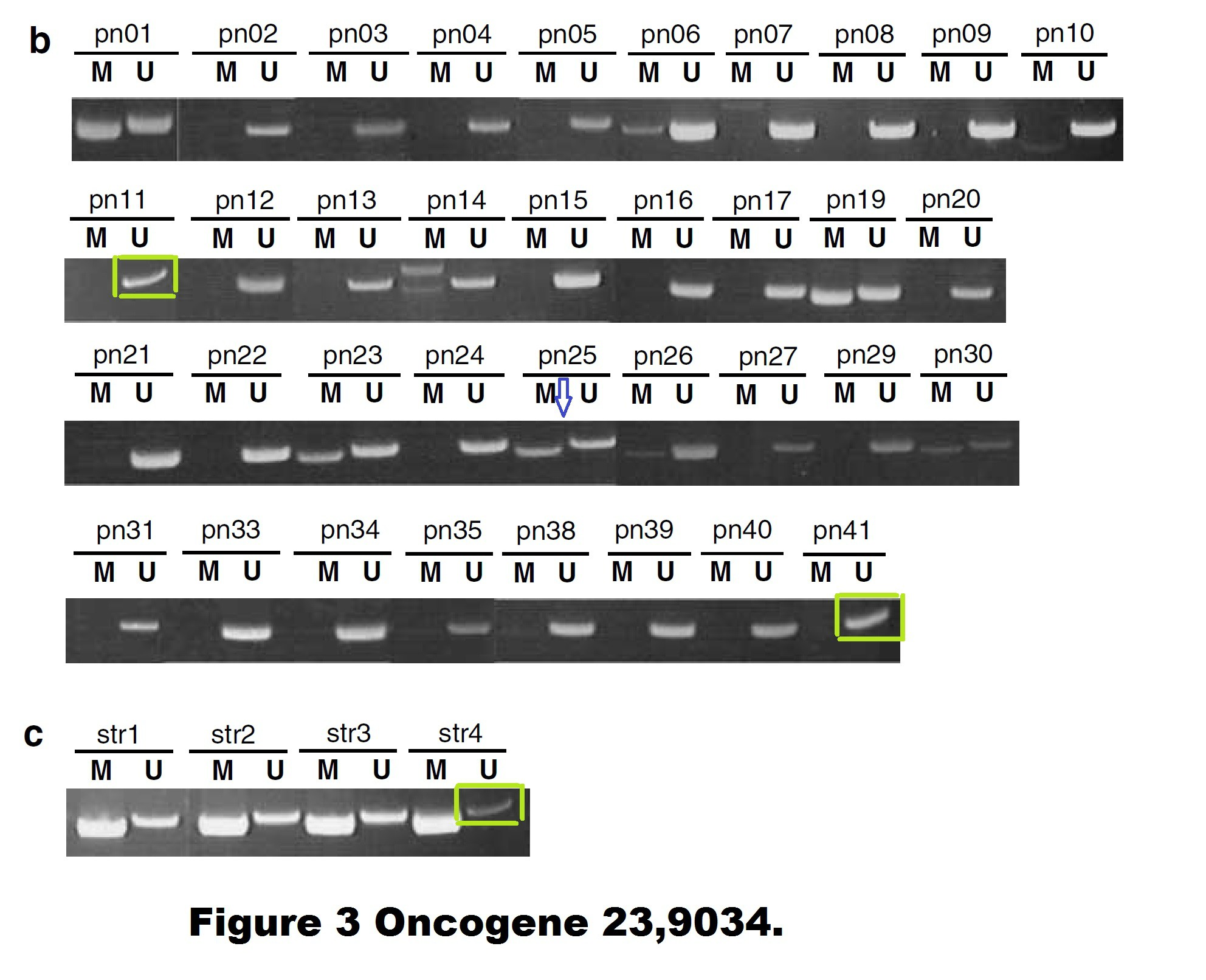
Quite possible indeed that these gel bands, flagged on PubPeer in 2015, are not identical. But the Figure 3 is a disaster in many other ways. The allegedly continuous gels are actually Franken-gels spliced from many pieces. There are strange gel bands of unexpected sizes lurking in the shadows, there seem to be strange box-shaped objects hoovering below some of the gel bands, how can all of that be explained?
I wrote to Hermeking and the first author Dimitri Lodygin, now group leader at the University of Göttingen in Germany. They remained silent.
There are also other issues with that paper. A PubPeer commenter insisted in February 2020 that some gels might be too similar to the gels in another study from the Hermeking lab. It is difficult to check, the resolution of the gels is extremely low, maybe Lodygin and Hermeking will find at least the original figure files they submitted to the journal?
Dimitri Lodygin , Alexey Epanchintsev , Antje Menssen , Joachim Diebold , Heiko Hermeking Functional epigenomics identifies genes frequently silenced in prostate cancer Cancer Research (2005) doi: 10.1158/0008-5472.can-04-4407

There is also this, flagged already in 2015, according to Hermeking the bands are again “similar, but not identical“:
These SFRP1 bands do look quite unexpectedly similar though. And the rest of the figure is a bit strange, too. Again those strange box-shaped shadows lacking all background, difficult to explain with image compression alone… In one case, a RASSf1A band looks as if it is inside a box.
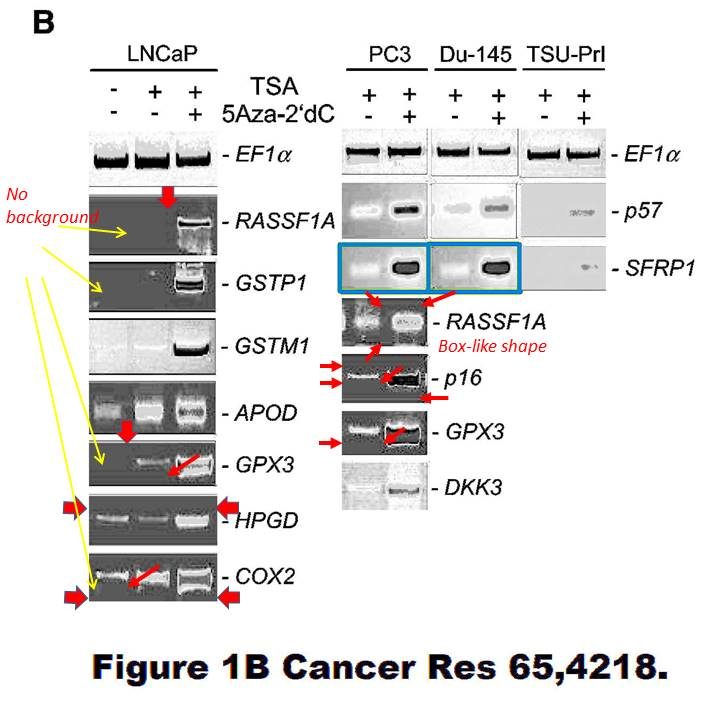
Again, maybe the authors have higher resolution versions of those gel images? Maybe the authors should check all their gel figures?
Dimitri Lodygin , Amir S Yazdi , Christian A Sander , Thomas Herzinger , Heiko Hermeking Analysis of 14-3-3sigma expression in hyperproliferative skin diseases reveals selective loss associated with CpG-methylation in basal cell carcinoma Oncogene (2003) doi: 10.1038/sj.onc.1206854
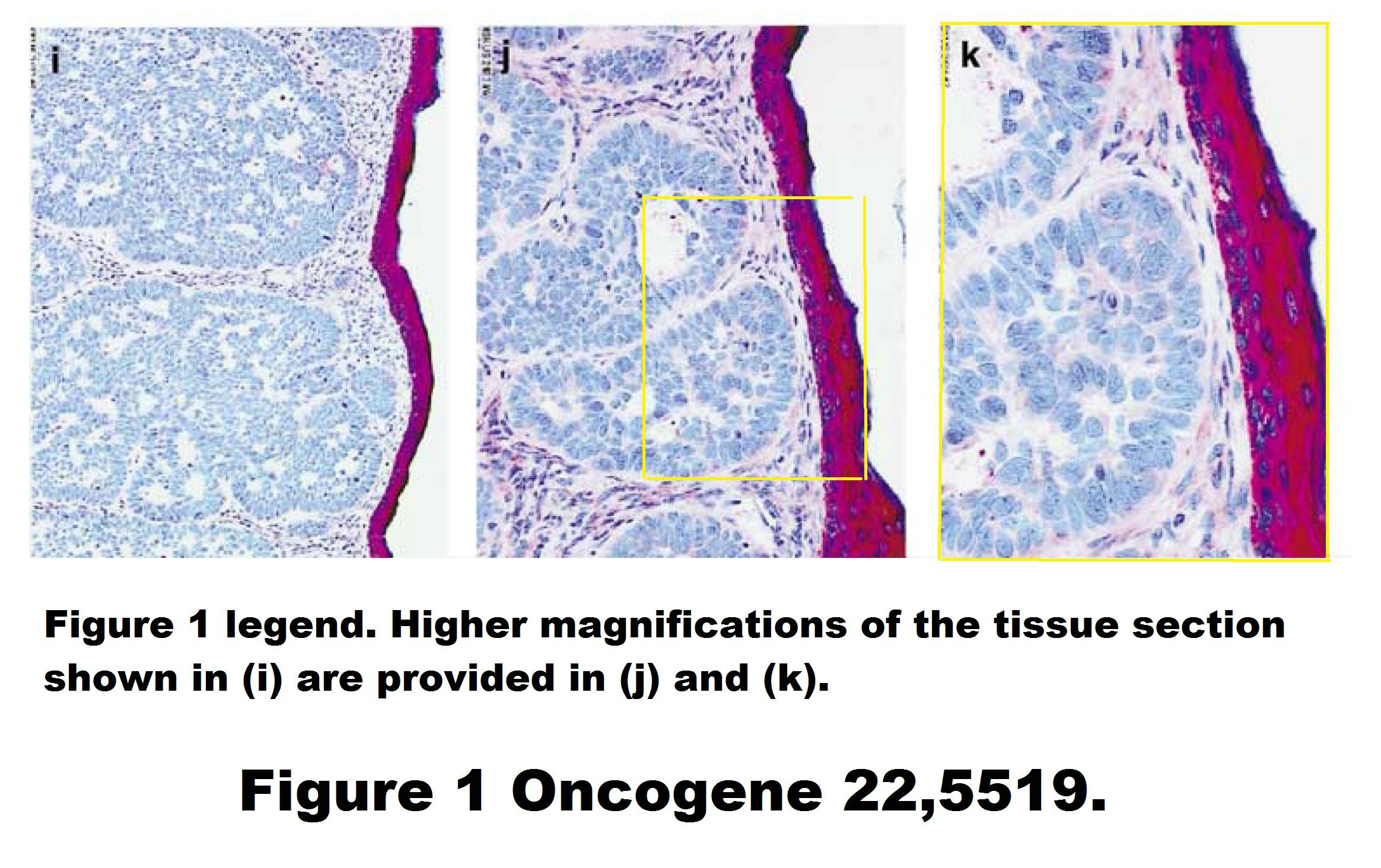
However, it seems that in the only gel figure of that paper, there is some possible splicing. If true, it was extremely difficult to find.
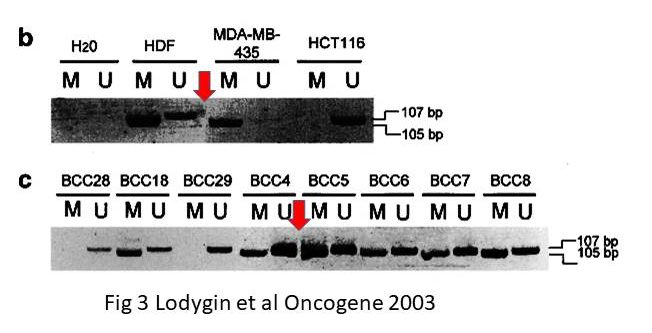
In an old collaborative paper, where Hermeking is one of the middle authors, a PubPeer user alleged in 2015 some gel band duplications:
A. T. Ferguson , E. Evron , C. B. Umbricht , T. K. Pandita , T. A. Chan , H. Hermeking , J. R. Marks , A. R. Lambers , P. A. Futreal , M. R. Stampfer , S. Sukumar High frequency of hypermethylation at the 14-3-3 sigma locus leads to gene silencing in breast cancer Proceedings of the National Academy of Sciences (2000) doi: 10.1073/pnas.100566997
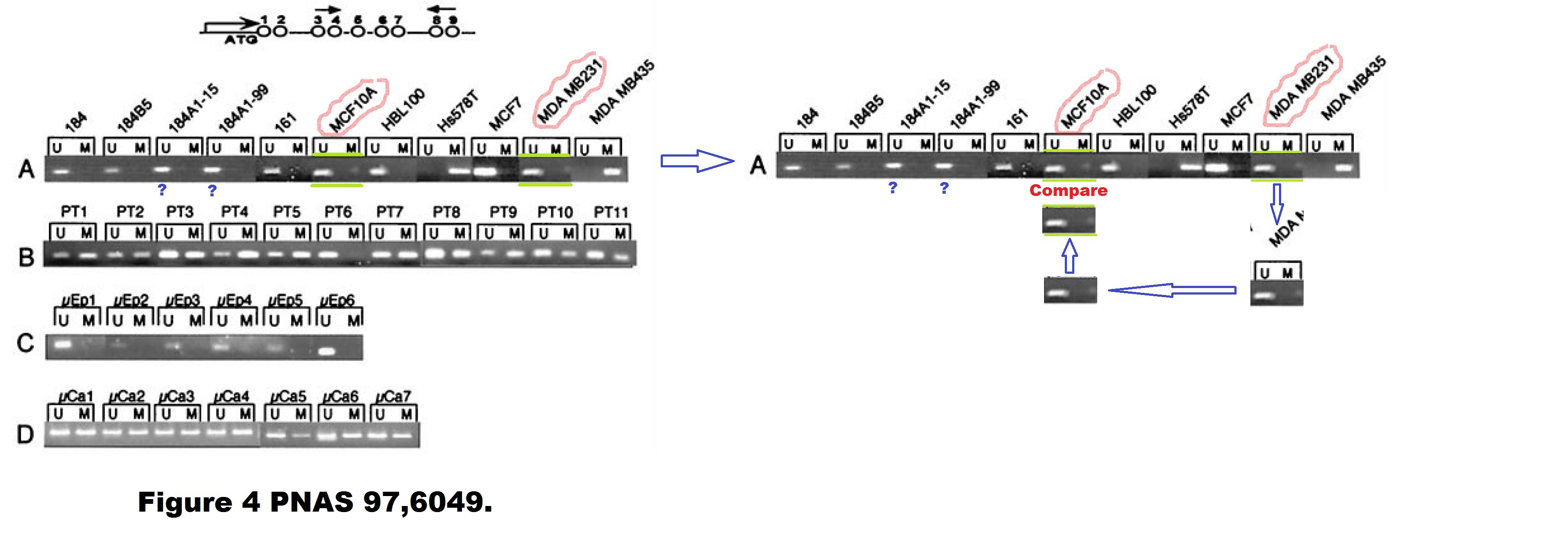
Professor Hermeking had no comments on that one. All four allegedly continuous gels of this figure are most obviously spliced from various pieces, allow me to visualise it:
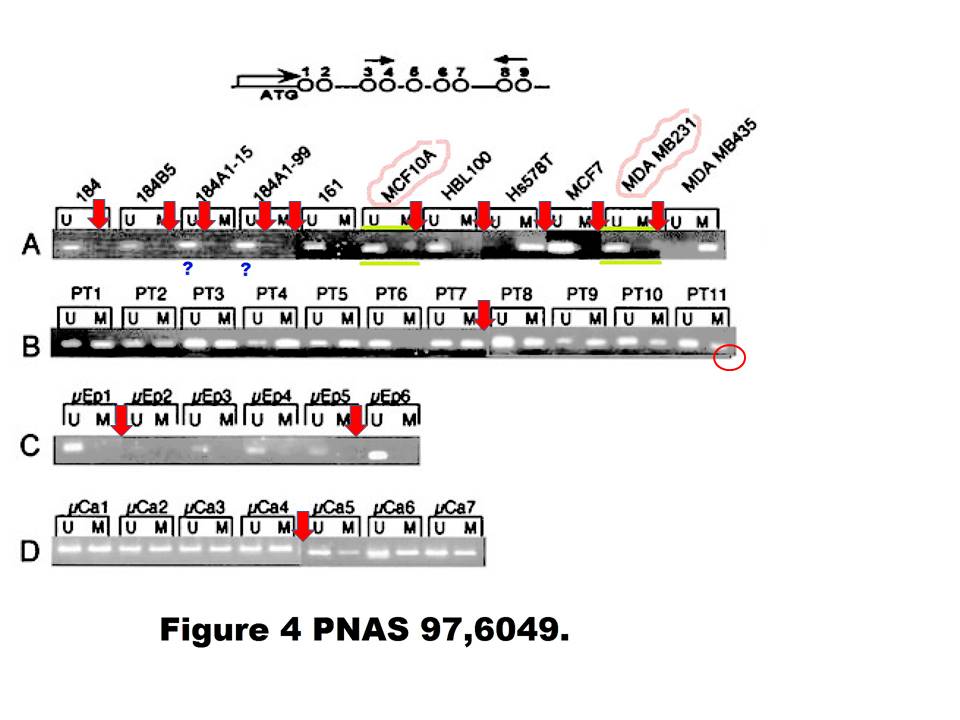
Under these circumstances, the case for the two suspected duplications is much stronger than the case against. There is even a corner (red circle) where one additional gel image was digitally overlaid on top of the original gel image. This is not splicing as such anymore, but something much more problematic. I am puzzled why Professor Hermeking is reluctant to find and name the culprit here.
Here are two old Hermeking papers where it is not exactly clear what the scientific rationale for the (undeclared) gel lane splicing was.
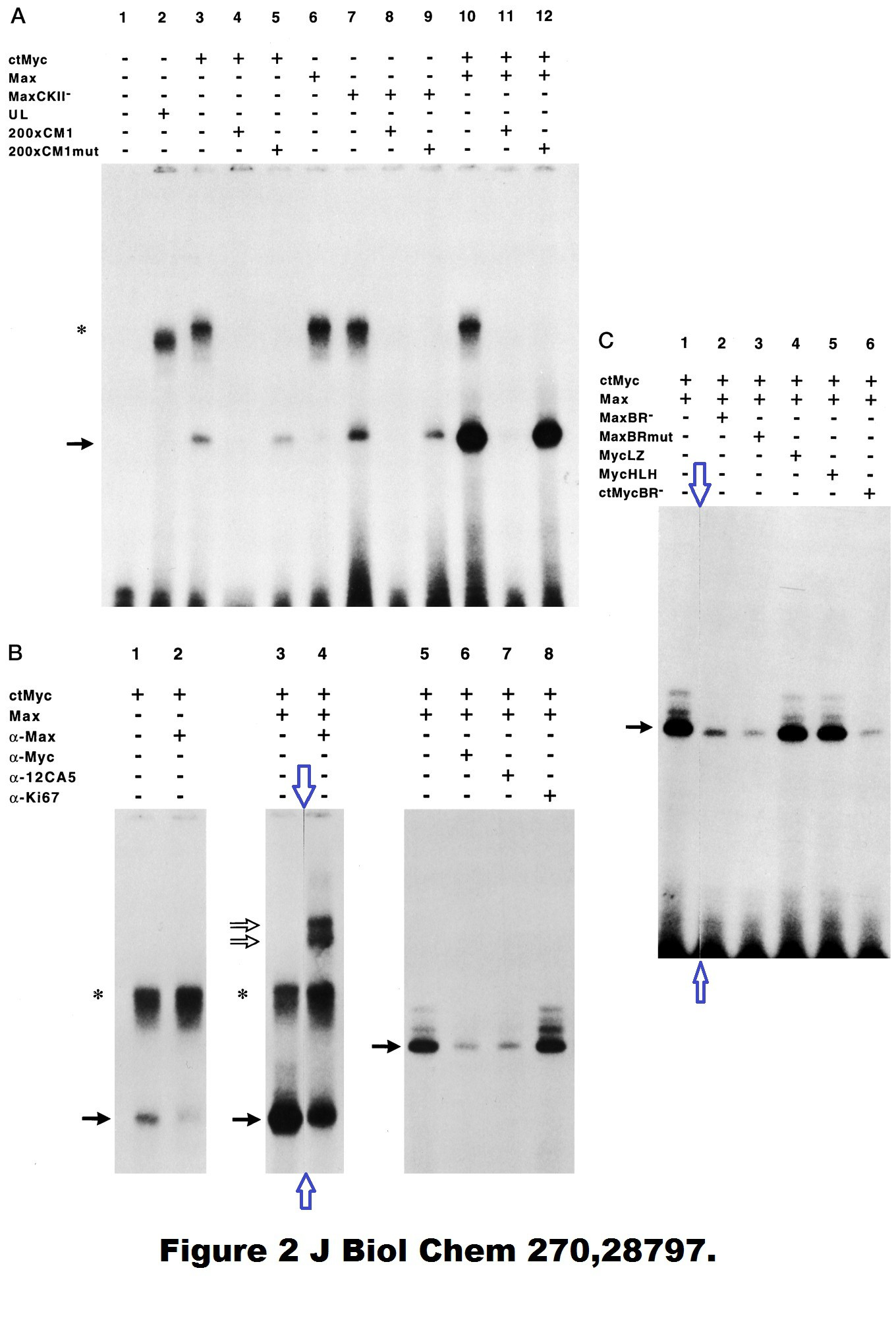
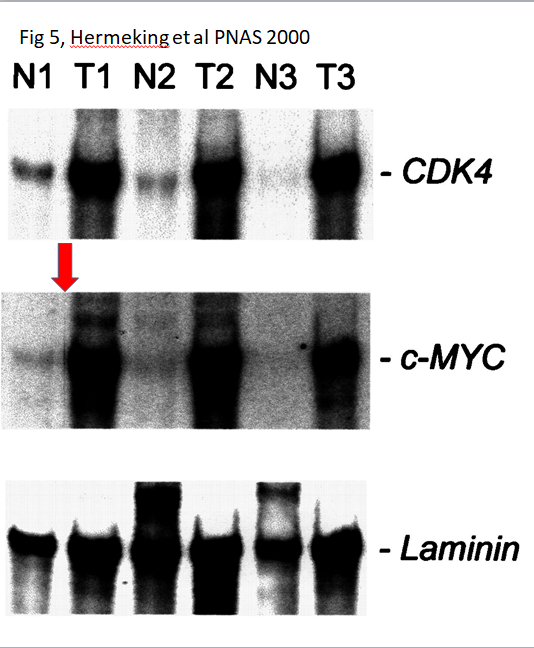
Regarding the 1995 JBC paper, it was the PhD student, as Hermking states:
“Work of a PhD student (Franz Kohlhuber) in the laboratory of my supervisor (Dirk Eick). The position where the membrane/gel/autoradiography was cut is clearly visible in my opinion. The data in itself is not “manipulated” by this, irrelevant results were removed from the figure.“
I presume the splicing in the PNAS 2000 paper was done by some PhD student in Vogelstein’s lab?
Most recently, this paper was flagged by the pseudonymous Clare Francis:
Helge Siemens , Jens Neumann , Rene Jackstadt , Ulrich Mansmann , David Horst , Thomas Kirchner , Heiko Hermeking Detection of miR-34a promoter methylation in combination with elevated expression of c-Met and β-catenin predicts distant metastasis of colon cancer Clinical cancer research (2013) doi: 10.1158/1078-0432.ccr-12-1703
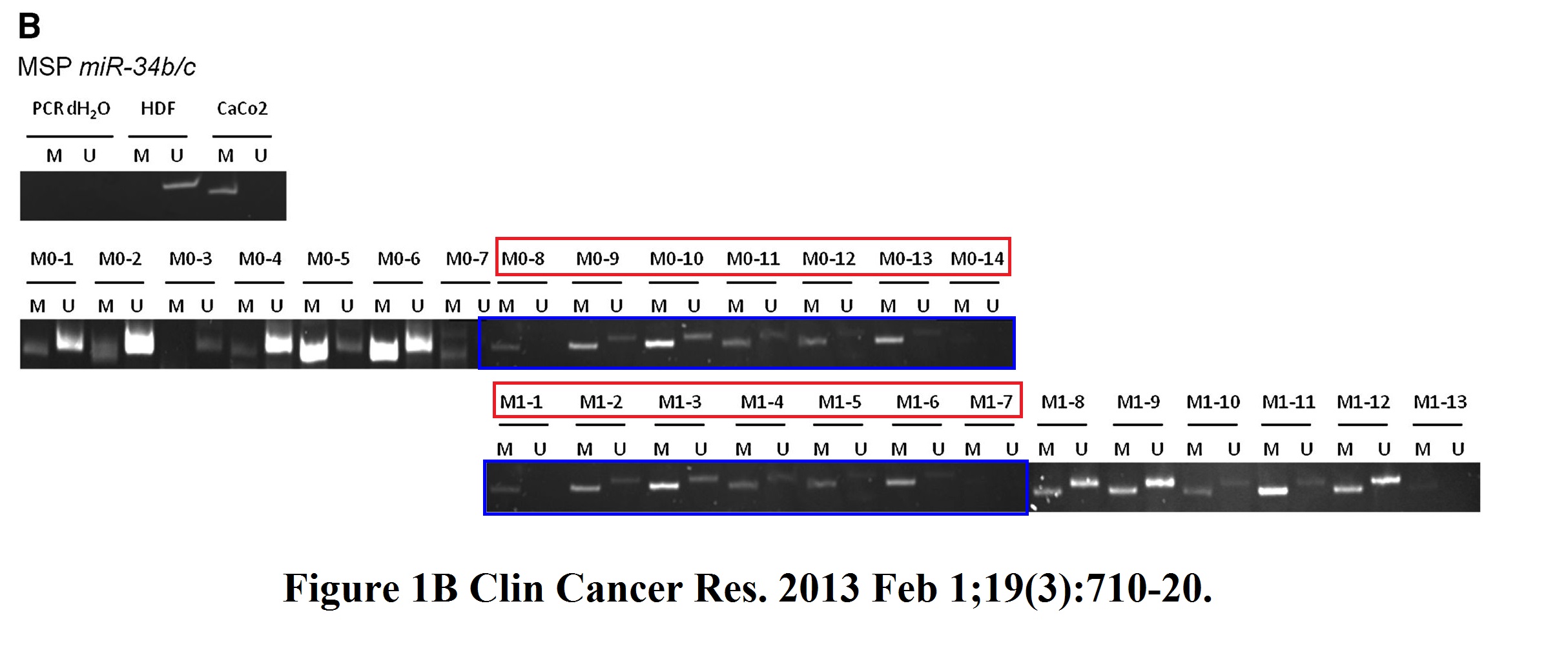

I asked Hermeking which PhD student was it this time, he replied:
“I will discuss this with the doctoral student at the time, Helge Siemens.
However, Fig. 1B (miR-34b/c methylation) is irrelevant for the statement of the publication. Like in result text, we have focused all further analyzes in this publication on miR-34a methylation, since miR-34b/c is associated in almost all tumors, whether with (M1) or without (M0).distant metastasis, is methylated. We had found this in other studies as well.
However, this should not be an excuse for this figure.
We will of course try to work on a corrigendum for this.”
I also think all these figures, and most likely the papers they are part of, are all irrelevant. Anyway, I don’t know about LMU Munich, but Johns Hopkins University has probably enough of these PubPeer antics.
Contact Form
I thank all my donors for supporting my journalism. You can be one of them!
Make a one-time donation:
I thank all my donors for supporting my journalism. You can be one of them!
Make a monthly donation:
Choose an amount
Or enter a custom amount
Your contribution is appreciated.
Your contribution is appreciated.




“Yardena Samuels , Luis A. Diaz , Oleg Schmidt-Kittler , Jordan M. Cummins , Laura Delong , Ian Cheong , Carlo Rago , David L. Huso , Christoph Lengauer , Kenneth W. Kinzler , Bert Vogelstein , Victor E. Velculescu Mutant PIK3CA promotes cell growth and invasion of human cancer cells Cancer Cell (2005) doi: 10.1016/j.ccr.2005.05.014”
https://www.weizmann.ac.il/mcb/Samuels/home
The first author benefits from these papers with problematic data:-
https://pubpeer.com/publications/728B8537D6CEB22560D9345F3F3D1E
https://pubpeer.com/publications/06080F7D25D1A1B598F4DE8DEF5238
https://pubpeer.com/publications/46CD919799986C18D15745C778C13C
https://pubpeer.com/publications/858FBC108F303612266C62378E71C5
See third lion here:
https://forbetterscience.com/2021/02/10/the-english-science-supremacy/
LikeLike
Same institute as Heiko Hermeking.
https://www.mhb.uni-muenchen.de/research/involved-professors/oncology/thomas-kirchner/index.html
Prof. Dr. med. Thomas Kirchner
Director, Institute of Pathology, Medical faculty – LMU
Hermekinghttps://www.pathologie.med.uni-muenchen.de/020wissenschaft/009ag_hermeking/index.html
Prof. Dr. med. Thomas Kirchner
More than 700 of publications, vast majority observational pathology.
https://pubmed.ncbi.nlm.nih.gov/?term=kirchner%20t&sort=date&ac=yes
Prof. Dr. med. Thomas Kirchner venturing into “science” with Florian Greten:-
https://pubpeer.com/publications/7DC774BD9A368A45CF48DD5305BBE9
https://pubpeer.com/publications/779511CD880489FA905B5DEF742AA8
https://pubpeer.com/publications/5DE4B1C5A4C6A62A92EF44732C444F
https://pubpeer.com/publications/8A70958AA4E4A680D4634FBFBBE3DC
FYI: https://fci.health/personen/florian-greten/
FLORIAN GRETEN,SPEAKER, DIRECTOR, FRANFURT CANCER INSTITUTE
Florian Greten’s Pubpeer record: https://pubpeer.com/search?q=florian+greten
Prof. Dr. med. Thomas Kirchner venturing into “science” with Heiko Hermeking:-
https://pubpeer.com/publications/FC8A5F7E0603C4CD1C3B726FAF92F8
LikeLike
Bert Vogelstein is most known for his work on p53, but what is that work, and how does it fit into the understanding of p53?
From EMBO Rep. 2010 Nov; 11(11): 822–826. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2966958/
we learn that p53 was discovered by at least 4 groups.
“The publication of these results, and other reports by Arnold Levine and Lionel Crawford, created the field of p53 research (Kress et al, 1979; Lane & Crawford, 1979; Linzer & Levine, 1979).
History mainly remembers the virological approach that led to the discovery of p53. However, Lloyd Old’s team at the Ludwig Institute for Cancer Research in New York, USA took an immunological approach and published their findings in the same year (DeLeo et al, 1979). Old showed that the humoral response of mice to some induced tumour cells was directed against a 53-kDa protein. His team found that in animals, SV40 tumours elicited an immune response specific to this protein. Crawford also described antibodies in human serum that target the 53-kDa protein in 9% of breast cancer patients. This immunological research was just as original, but the compartmentalization of research meant that scientists did not immediately recognize the link between the two sets of observations (DeLeo et al, 1979). Several years later, it was established that the protein was the same and it was imaginatively baptized p53 (Crawford, 1983).”
The 4 groups being those of:-
Lionel Crawford, Pierre May, Arnold Levine, Lloyd Old.
From Nat Rev Cancer. 2009 Oct; 9(10): 749–758.https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2771725/
“It was while studying these SV40-derived tumor antigens that several groups independently stumbled on p53. This happened in 1979, thirty years ago (Timeline). Working at the ICRF (now London Institute for Cancer Research), David Lane and Lionel Crawford realized that when sera from animals bearing SV40-induced tumors were employed to immunoprecipitate SV40 large T-antigen, a non-viral protein with an apparent molecular mass of about 53 kDa came along for the ride 3. Further analysis established that this cellular protein was physically complexed with SV40 large T-antigen. Thus the viral protein, previously shown to be largely responsible for the transforming and tumorigenic activity of the SV40 virus, had selected this hitherto unknown cellular protein as its partner for an intimate, specific interaction. At the same time, Daniel Linzer and Arnold Levine applied a similar immunological approach to SV40 transformed cells, and came up with essentially the same observations, namely that such cells harbored a complex between the SV40 large T-antigen and the cellular 53 kDa protein 4. Three other groups, those of Alan Smith in the UK, Robert Carroll in New York and Pierre May in France, simultaneously made very similar findings, all published in 1979 5-7.
Interestingly, Linzer and Levine also found that their antisera precipitated the same 53 kDa protein from teratocarcinoma (a germ cell tumor)-derived cells, despite the fact that the latter did not harbor any SV40 proteins; this indicated that a subset of the antibodies raised against the viral-induced tumor were capable of interacting directly with this cellular protein 4. In parallel, Lloyd Old and coworkers demonstrated that animals immunized with non-virally transformed cells produced antibodies to the same 53 kDa protein 8, rightfully qualifying it as a cellular tumor antigen. Moreover, Varda Rotter, working in the lab of David Baltimore, was able to identify the same protein being produced in excess in cells transformed by a retrovirus, the Abelson murine leukemia virus 9. Hence, very high levels of this new cellular protein were present not only in SV40-transformed cells but also in other types of cancer cells, but little or no p53 protein could be detected in non-transformed cells.”
This review credits the groups of:-
Lionel Crawford,Arnold Levine,Alan Smith,Robert Caroll,Pierre May,David Baltimore,Lloyd Old.
That is 7 groups discovering p53, with 4 of those 7 in common between the 2 reviews.
In the very beginning p53 was thought to be an oncogene, but there was growing evidence against that being the case.
From EMBO Rep. 2010 Nov; 11(11): 822–826. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2966958/
“studies of p53 began to encounter difficulties and the 131 articles published on the topic during this period did not integrate p53 into the family of oncogenes. Among these studies was the finding that both p53 alleles were inactivated in murine erythroleukaemia induced by the Friend virus (Mowat et al, 1985).”
EMBO Rep. 2010 Nov; 11(11): 822–826 also states that “high frequency of p53 gene rearrangements and deletions in human and murine osteosarcoma (Masuda et al, 1987).
Both ” both p53 alleles were inactivated in murine erythroleukaemia ” and “high frequency of p53 gene rearrangements and deletions in human and murine osteosarcoma” strongly suggest that p53 can act as a tumor suppressor gene. The principle was established in mice (1985) and extended to human tumours (1987).
“demonstration of inactivating mutations in human cancers eventually changed the status of p53 to a tumour suppressor gene (Baker et al, 1989; Takahashi et al, 1989)”.
Baker et al, 1989, is a highly cited paper by Vogelstein, where deletions of the region haboring p53 and and p53 mutations are common in human colorectal carcinomas, which comes the same year as another paper showing p53 as a frequent target for genetic abnormalites in human cancers (lung), yet both are predated by Mowat et al, 1985 and Masuda et al, 1987, which demonstrated that p53 acts as a tumor suppressor gene in the mouse and in human osteosarcomas.
Nature. 1985 Apr 18-24;314(6012):633-6. doi: 10.1038/314633a0.
Rearrangements of the cellular p53 gene in erythroleukaemic cells transformed by Friend virus
M Mowat, A Cheng, N Kimura, A Bernstein, S Benchimol
PMID: 3990796 DOI: 10.1038/314633a0
Proc Natl Acad Sci U S A. 1987 Nov;84(21):7716-9. doi: 10.1073/pnas.84.21.7716.
Rearrangement of the p53 gene in human osteogenic sarcomas
H Masuda 1, C Miller, H P Koeffler, H Battifora, M J Cline
Affiliation
1Department of Medicine, University of California, Los Angeles 90024.
PMID: 2823272 PMCID: PMC299371 DOI: 10.1073/pnas.84.21.7716
Science . 1989 Apr 14;244(4901):217-21. doi: 10.1126/science.2649981.
Chromosome 17 deletions and p53 gene mutations in colorectal carcinomas
S J Baker 1, E R Fearon, J M Nigro, S R Hamilton, A C Preisinger, J M Jessup, P vanTuinen, D H Ledbetter, D F Barker, Y Nakamura, R White, B Vogelstein
Affiliation
1Oncology Center, Johns Hopkins University School of Medicine, Baltimore, MD 21231.
PMID: 2649981 DOI: 10.1126/science.2649981
Science. 1989 Oct 27;246(4929):491-4. doi: 10.1126/science.2554494.
p53: a frequent target for genetic abnormalities in lung cancer
T Takahashi 1, M M Nau, I Chiba, M J Birrer, R K Rosenberg, M Vinocour, M Levitt, H Pass, A F Gazdar, J D Minna
Affiliation
1National Cancer Institute-Navy Medical Oncology Branch, Bethesda, MD 20814.
PMID: 2554494 DOI: 10.1126/science.2554494
LikeLike
Who discovered what in science is like in a war: history is written by the winners. If the winner is a white man, the world is happy to celebrate them. My guess is Rosalind Franklin only eventually got her partial credit because Watson spend decades after her cancer death publicly defaming and ridiculing her, so people wondered what really went on.
Behind every Nobel Prize winner are many subordinate scientists whose names were extinguished from history so the white dude gets the Nobel Prize.
LikeLike
p53 as a tumor suppressor gene.
EMBO Rep. 2010 Nov; 11(11): 822–826.
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2966958/
“…131 articles published on the topic during this period did not integrate p53 into the family of oncogenes. Among these studies was the finding that both p53 alleles were inactivated in murine erythroleukaemia induced by the Friend virus (Mowat et al, 1985).”
Mowat M, Cheng A, Kimura N, Bernstein A, Benchimol S (1985) Rearrangements of the cellular p53 gene in erythroleukaemic cells transformed by Friend virus. Nature 314: 633–636.
Nature Reviews Cancer volume 9, pages749–758 (2009)
Click to access nrc2723.pdf
p53 revisited: rise of a tumour suppressor
The realization that p53 is not actually an oncogene but rather the opposite, namely a
tumour suppressor, took several more years. In fact, clues that p53 might be a tumour suppressor had already been around. In retrospect, the first clue was provided by David Wolf and Rotter, who reported in 1984 that Trp53 (which encodes mouse p53) was inactivated by retroviral insertion in an Abelson murine leukaemia virus-transformed mouse cell line26. Similar observations were made by Sam Benchimol, Alan Bernstein andco-workers, studying leukaemias induced in mice by the Friend erythroleukaemia virus 27,28. Furthermore, Rotter and co-workers showed that TP53 was extensively rearranged and its coding sequences virtually deleted in the human leukaemia-derived cell line HL60, precluding production of p53 (Ref. 29). The simplest interpretation of these striking observations was that the loss of p53 promotes cancer, implying that sustained p53 function is necessary to prevent cancer.
Ben David, Y., Prideaux, V. R., Chow, V., Benchimol, S.
& Bernstein, A. Inactivation of the p53 oncogene by internal deletion or retroviral integration in erythroleukemic cell lines induced by Friend leukemia
virus. Oncogene 3, 179–185 (1988).
Mowat, M., Cheng, A., Kimura, N., Bernstein, A. &
Benchimol, S. Rearrangements of the cellular p53 gene in erythroleukaemic cells transformed by Friend virus. Nature 314, 633–636 (1985).
Wolf, D. & Rotter, V. Major deletions in the gene encoding the p53 tumor antigen cause lack of p53 expression in HL-60 cells.
Proc. Natl Acad. Sci. USA
82, 790–794 (1985).
It seems that Sam Benchimol and his colleagues in Toronto, and David Wolf and Varda Rotter at the Weizmann Institute, got there before Bert Vogelstein.
Credit where credit is due, Varda Rotter and David Baltimore have a claim to one of the first people to discover p53.
J Virol. 1980 Nov; 36(2): 547–555.PMCID: PMC353673
PMID: 6159484
Abelson murine leukemia virus-induced tumors elicit antibodies against a host cell protein, P50. V Rotter, O N Witte, R Coffman, and D Baltimore
LikeLike
“Regarding that 1999 Nature paper, Hermeking also declared to me:
In my opinion, it is well known that not all studies on the same topic come to the same conclusion. In this case, an author with a publication on the function of 14-3-3sigma that had a different/different result appears to be using the PubPeer forum to discredit authors and their co-authors who disagree with him.”
Examples of “not all studies on the same topic come to the same conclusion” have sprung to Hermeking’s mind and form the basis of his opinion. Could he enlighten the rest of us by sharing some of these examples?
There are problems with images in the original 1999 Nature paper.
There appear to be two instances of image duplication.
The major findings of the 1999 Nature paper cannot be reproduced by two other groups:-
Cancer Res. 2001 Oct 15;61(20):7660-8.
Neither p21WAF1 nor 14-3-3sigma prevents G2 progression to mitotic catastrophe in human colon carcinoma cells after DNA damage, but p21WAF1 induces stable G1 arrest in resulting tetraploid cells
P R Andreassen 1, F B Lacroix, O D Lohez, R L Margolis
Affiliation
1Institut de Biologie Structurale Jean-Pierre Ebel (CEA-CNRS), 38027 Grenoble Cedex 1, France.
PMID: 11606409
Radiat Res. 2004 Sep;162(3):270-86. doi: 10.1667/rr3221.
Computerized video time lapse study of cell cycle delay and arrest, mitotic catastrophe, apoptosis and clonogenic survival in irradiated 14-3-3sigma and CDKN1A (p21) knockout cell lines
Kenneth Chu 1, Noella Teele, Michael W Dewey, Norman Albright, William C Dewey
Affiliations collapse
Affiliation
1Radiation Oncology Research Laboratory, University of California San Francisco, San Francisco, California 94103, USA.
PMID: 15332997 DOI: 10.1667/rr3221
https://pubpeer.com/publications/876EA56A4549BCAA87455EA00E5A19#14
Materials and methods.
Cell lines. The human colorectal carcinoma HCT116 cell line and its derivatives with CDKN1A or 14-3-3 sigma or both these genes knocked out by homologous recombination disruption of both alleles were kindly provided by Dr. Bert Vogelstein of Johns Hopkins University.
The data in figures 2,3,6 and 7 for the HCT116 14-3-3 sigma -/- cells look more similar to the data for the HCT116 Wild Type cells than they do to the data for the HCT116 CDKN1A -/- or HCT116 double knockout cells.
Discussion, beginning paragraph 3.
“While knocking out CDKN1A increased the overall amount of cell death, as well as proliferation, knocking out 14-3-3 sigma increased only the amount of cell death, without allowing proliferation”.
Discussion, paragraph 4.
“when the CDKN1A protein was absent in either the single-knockout or double-knockout cells,
proliferation was essentially unchecked regardless of the 14-3-3sigma status”.
LikeLike
Correction years after publication.
Author Correction: 14-3-3σ is required to prevent mitotic catastrophe after DNA damage | Nature
There are more problematic data.
More problematic data Nature . 1999 Oct 7;401(6753):616-20.
Problematic data figure 1d.
4d, Western blot analysis. Cells with the indicated 14-3-3σ genotype were cultured in the absence (Con) or presence (Adr) of adriamycin and subsequently lysed in sample buffer. Extracts were fractionated by SDS–PAGE and probed with an antibody specific for 14-3-3σ.”
There does not seem to be any loading control.
There is a vertical, if slightly scalloped, change in signal between the +/+ Con (lane 1) and +/+ Adr lane (lane 2). The bands in lanes 2,3 and 4 are horizontal, yet the 3 bands in lane 1 are parallel to each other and are at about 15 degrees to the horizontal. Lanes 1 and 2 do not look like they come from adjacent parts of the same blot. It’s not obvious how the 3 bands in lane 1 relate to the bands in lanes 2, 3 and 4.
PubPeer – 14-3-3Sigma is required to prevent mitotic catastrophe after…
Nature’s reply.
“I am contacting you as you expressed concerns regarding the manuscript “14-3-3Sigma is required to prevent mitotic catastrophe after DNA damage” by Bert Vogelstein and colleagues.
We have investigated the matter thoroughly in line with the Committee on Publication Ethics guidelines.
We have found that the images in the article and the correction , meet our expectations for data from 1999, and we will not pursue the matter further at this time.
We will therefore consider the matter closed. “
“meet our expectations for data from 1999” sounds like it would not meet their expectations in 2024, and the extensive correction for 2 figures was in August 2023.
LikeLike
Proc Natl Acad Sci U S A. 2002 Jun 11; 99(12): 8265–8270.
doi: 10.1073/pnas.082240999
PMCID: PMC123056
PMID: 12060769
Targeted inactivation of CTNNB1 reveals unexpected effects of β-catenin mutation
Timothy A. Chan,* Zhenghe Wang,* Long H. Dang,* Bert Vogelstein,† and Kenneth W. Kinzler‡
Author information Article
*The Sidney Kimmel Comprehensive Cancer Center and †The Howard Hughes Medical Institute, Johns Hopkins Medical Institutions, 1650 Orleans Street, Baltimore, MD 21231
‡To whom reprint requests should be addressed. E-mail: kinzelke@jhmi.edu
Contributed by Bert Vogelstein
Figure 4C. Beta-Catenin panel. Band lane 2 has straight, vertical right edge.
LikeLike
Even without splicing, it is perfectly clear those Cadherins are on two physically separate gels, ie. without any loading control. And even then they had to splice the lanes from what may be a third gel.
https://pubpeer.com/publications/0EE8E1A5B0F83ABFA207C0EDB0F6D7
This is why scientists who diligently do loading controls on the same gel are unable to produce such groundbreaking science as Vogelstein.
LikeLike
” Apparently, Vogelstein and Kinzler are such big stars of cancer research, that Johns Hopkins University allows them to do to their mice what pedestrian scientists are not allowed to do, due to concerns about animal welfare and cruelty.”
Proc Natl Acad Sci U S A. 2004 Oct 19; 101(42): 15172–15177.
Published online 2004 Oct 7. doi: 10.1073/pnas.0406242101
PMCID: PMC523456
PMID: 15471990
Medical Sciences
Bacteriolytic therapy can generate a potent immune response against experimental tumors
Nishant Agrawal,†‡ Chetan Bettegowda,†§ Ian Cheong,†§ Jean-Francois Geschwind,¶ Charles G. Drake,† Edward L. Hipkiss,† Mitsuaki Tatsumi,¶ Long H. Dang,† Luis A. Diaz, Jr.,† Martin Pomper,¶ Mohammad Abusedera,¶ Richard L. Wahl,¶ Kenneth W. Kinzler,† Shibin Zhou,† David L. Huso,∥ and Bert Vogelstein†**
Author information Copyright and License information Disclaimer
** To whom correspondence should be addressed. E-mail: vodelbe@jhmi.edu
§C.B. and I.C. contributed equally to this work.
Contributed by Bert Vogelstein, August 25, 2004
Problematic data figure 4. Ulcerated, bleeding tumor.
LikeLike
” Apparently, Vogelstein and Kinzler are such big stars of cancer research, that Johns Hopkins University allows them to do to their mice what pedestrian scientists are not allowed to do, due to concerns about animal welfare and cruelty.”
Cancer Biol Ther. 2004 Mar;3(3):326-37. doi: 10.4161/cbt.3.3.704. Epub 2004 Mar 12.
Targeting vascular and avascular compartments of tumors with C. novyi-NT and anti-microtubule agents
Long H Dang 1, Chetan Bettegowda, Nishant Agrawal, Ian Cheong, David Huso, Philip Frost, Frank Loganzo, Lee Greenberger, Jozsef Barkoczy, George R Pettit, Amos B Smith 3rd, Hallur Gurulingappa, Saeed Khan, Giovanni Parmigiani, Kenneth W Kinzler, Shibin Zhou, Bert Vogelstein
Affiliation
1Howard Hughes Medical Institute and Sidney Kimmel Cancer Center, Johns Hopkins Medical Institutions, Baltimore, Maryland, USA.
PMID: 14739784 DOI: 10.4161/cbt.3.3.704
Figure 1B. Large (over half width of back), ulcerated tumor.
LikeLike
Sam W Lee once got in trouble for torturing mice with such tumour xenografts.
https://www.nature.com/articles/nature10167
https://pubpeer.com/publications/86A05A1191BCC54266244DB3839DAA
Lee later was exposed as research fraudster, kicked out of Harvard and slapped with retractions. Even his mentor, the psycho bully Stu Aaranson couldn’t save him.
Once you post it all on PubPeer, I will inform Johns Hopkins
LikeLike
HCT116 wild-type and and their isogeneic sublines HCT116-p21 -/- and HCT116-p53 -/- in Berlin.
Oncogene. 2005 Jun 9;24(25):4114-28. doi: 10.1038/sj.onc.1208579.
Loss of p21 disrupts p14 ARF-induced G1 cell cycle arrest but augments p14 ARF-induced apoptosis in human carcinoma cells
Philipp G Hemmati 1, Guillaume Normand, Berlinda Verdoodt, Clarissa von Haefen, Anne Hasenjäger, Dilek Güner, Jana Wendt, Bernd Dörken, Peter T Daniel
Affiliation
1Department of Hematology, Oncology and Tumor Immunology, University Medical Center Charité, Campus Berlin-Buch, Berlin-Buch, Germany.
PMID: 15750619 DOI: 10.1038/sj.onc.1208579
Click to access 1208579.pdf
“HCT116 wild-type cells and their isogeneic knockout
sublines HCT116-p21 -/- (Waldman et al., 1995) and HCT116-
p53-/- (Bunz et al., 1998) were kindly provided by Dr Bert
Vogelstein, Johns Hopkins Cancer Center, Baltimore, MD,
USA.”
Figure 1b much more similar and different than expected. How to explain?
J Biol Chem. 2005 Feb 25;280(8):7118-30. doi: 10.1074/jbc.M412330200. Epub 2004 Dec 6.
p14ARF induces G2 cell cycle arrest in p53- and p21-deficient cells by down-regulating p34cdc2 kinase activity
Guillaume Normand 1, Philipp G Hemmati, Berlinda Verdoodt, Clarissa von Haefen, Jana Wendt, Dilek Güner, Evelyne May, Bernd Dörken, Peter T Daniel
Affiliations collapse
Affiliation
1Department of Hematology, Oncology and Tumor Immunology, University Medical Center Charité, Campus Berlin-Buch, D-13125 Berlin-Buch, Germany.
PMID: 15582998 DOI: 10.1074/jbc.M412330200
LikeLike
J Biol Chem. 2005 Feb 25;280(8):7118-30. doi: 10.1074/jbc.M412330200. Epub 2004 Dec 6.
p14ARF induces G2 cell cycle arrest in p53- and p21-deficient cells by down-regulating p34cdc2 kinase activity
Guillaume Normand 1, Philipp G Hemmati, Berlinda Verdoodt, Clarissa von Haefen, Jana Wendt, Dilek Güner, Evelyne May, Bernd Dörken, Peter T Daniel
Affiliation
1Department of Hematology, Oncology and Tumor Immunology, University Medical Center Charité, Campus Berlin-Buch, D-13125 Berlin-Buch, Germany.
PMID: 15582998 DOI: 10.1074/jbc.M412330200
Problematic data figure 2A. much more similar than expected.
LikeLike
Additional problematic data Oncogene . 2005 Jun 9;24(25):4114-28. doi: 10.1038/sj.onc.1208579
Problematic data figure 6a. Much more similar than expected.
LikeLike
HCT116 cells in Berlin.
J Cell Biol . 2007 Nov 19;179(4):701-15. doi: 10.1083/jcb.200703040.
Mcl-1 determines the Bax dependency of Nbk/Bik-induced apoptosis
Bernhard Gillissen 1, Frank Essmann, Philipp G Hemmati, Antje Richter, Anja Richter, Ilker Oztop, Govindaswamy Chinnadurai, Bernd Dörken, Peter T Daniel
Affiliation
1
Department of Hematology, Oncology, and Tumor Immunology, University Medical Center Charité, 13125 Berlin, Germany.
PMID: 18025305
PMCID: PMC2080900
DOI: 10.1083/jcb.200703040
Problematic data figure 2B. Much more similar than expected.
LikeLike
I informed Peter Daniel of these two papers.
LikeLike
To me, it does seem kind of weird that Vogelstein got do famous and became such a big money grabber when he didn’t seem to do anything that earth shattering, like Bob Weinberg and his idea to transfect oncogenes into cell lines to ID them—-that really opened up the genetic basis of cancer research. All I an tell is the Vogelstein seem to ride a hot wave of research—tumor surpressors, which he never really seem to lead in a major fashion like Weinberg.
I had a good friend that was a grad student for Eric Lander. Didnt mention if he was abusive or not.
LikeLike
Bert Vogelstein is well-known for his “adenoma to carcinoma of the colon” model with its accumulation of genetic changes during development of carcinomas. It’s a model, but no new principles. There are many lines of evidence for a multi-step model of tumor development using different methods, and there are other multi-step models. The “two-hit hypothesis” for the development retinoblastoma (inherited and non-inherited), associated with Alfred Knudsen relied on mathematics. 2 steps are more than 1. The “adenoma to carcinoma of the colon” extends this to 3,4, or more steps, but that is what many people believed before.
The mouse leukemia models of malignancy are often overlooked, but powerful as you have a biological output, the mice get leukemia, and you find genes, but not too many, in a relatively short time. The mouse erythroleukemia model showed that p53 is a tumor suppressor gene as often the p53 gene is either so rearranged, sometimes to the degree of being absent, that it cannot be working as an oncogene. Such work is most commonly associated with the work of Sam Benchimol and Yaakov BenDavid in Toronto, and Varda Rotter, first in David Baltimore’s lab, and since then at the Weizmann Institute. The papers which showed these things are not in obscure journals. To his credit, in early publications, Bert Vogelstein does correctly credit them with those discoveries.
Ben David, Y., Prideaux, V. R., Chow, V., Benchimol, S.
& Bernstein, A. Inactivation of the p53 oncogene by internal deletion or retroviral integration in erythroleukemic cell lines induced by Friend leukemia
virus. Oncogene 3, 179–185 (1988).
Mowat, M., Cheng, A., Kimura, N., Bernstein, A. &
Benchimol, S. Rearrangements of the cellular p53 gene in erythroleukaemic cells transformed by Friend virus. Nature 314, 633–636 (1985).
Wolf, D. & Rotter, V. Major deletions in the gene encoding the p53 tumor antigen cause lack of p53 expression in HL-60 cells.
Proc. Natl Acad. Sci. USA 82, 790–794 (1985).
LikeLike
Interesting. Thank you, Zeb. BTW, I have Weinberg’s book on cancer and IMO its better than Vogelstein’s.
LikeLike
“To his credit, in early publications, Bert Vogelstein does correctly credit them with those discoveries.”
Science . 1989 Apr 14;244(4901):217-21. doi: 10.1126/science.2649981.
Chromosome 17 deletions and p53 gene mutations in colorectal carcinomas
S J Baker 1, E R Fearon, J M Nigro, S R Hamilton, A C Preisinger, J M Jessup, P vanTuinen, D H Ledbetter, D F Barker, Y Nakamura, R White, B Vogelstein
Affiliation
1Oncology Center, Johns Hopkins University School of Medicine, Baltimore, MD 21231.
PMID: 2649981 DOI: 10.1126/science.2649981
Google scholar
Chromosome 17 deletions and p53 gene mutations in colorectal carcinomas
…, DF Barker, Y Nakamura, R White, B Vogelstein – Science, 1989 – science.org
Previous studies have demonstrated that allelic deletions of the short arm of chromosome 17
occur in over 75% of colorectal carcinomas. Twenty chromosome 17p markers were used …
Cited by 2729
“Third, p53 genes are often inactivated through proviral integration in
Friend virus-induced mouse leukemias (25,26). Fourth, rearrangements of the p53
gene occur in the human leukemia cell line HL60 and in some osteosarcomas, and no
p53 gene product is detectable in HL60 cells (27).”
References.
25. G. G. Hicks and M. Mowat, J. Virol. 62, 4752(1988).
26. M. Mowat et al., Nature 314, 633 (1985); D. G.Munroe, B. Rovinski, A. Bemstein, S. Benchimol,Oncogene 2, 621 (1988).
27. D. Wolf and V. Rotter, Proc. Natl. Acad. Sci. U.S.A.82, 790 (1985); H. Masuda, C. Miller, H. P. Koeffler, H. Battifora, M. J. Cline, Proc. NatI. Acad.
Sci. U.S.A. 84, 7716 (1987)
LikeLike
“To his credit, in early publications, Bert Vogelstein does correctly credit them with those discoveries.”
Click to access 342705a0.pdf
Nature. 1989 Dec 7;342(6250):705-8. doi: 10.1038/342705a0.
Mutations in the p53 gene occur in diverse human tumour types
Janice M. Nigro*, Suzanne J. Baker*, Antonette C. Preisinger*, J. Milburn Jessupt,
Richard Hostettert, Karen Clearyt, Sandra H. Bigner*, Nancy Davidson*, Stephen Baylin*, Peter Devilee §, Thomas Gloverll, Francis S. Collinsll, Ainsley Weston~,
Rama Modali~, Curtis C. Harris~ & Bert Vogelstein*#
* The Johns Hopkins Oncology Center, 424 North Bond Street, Baltimore.
Maryland 21231. USA
t M. D. Anderson Hospital. Houston, Texas 77030, USA
:j: Department of Pathology, Duke University, Durham,
North Carolina 27710, USA
§ Department of Human Genetics, University of Leiden, Holland
II Department of Human Genetics, University of Michigan, Ann Arbor,
Michigan 48109, USA
~ Laboratory of Human Carcinogenesis, National Cancer Institute,
Bethesda, Maryland 20892, USA
Mutations in the p53 gene occur in diverse human tumour types
…, A Weslon, R Modali, CC Harris, B Vogelstein – Nature, 1989 – nature.com
THE p53 gene has been a constant source of fascination since its discovery nearly a decade
ago 1, 2. Originally considered to be an oncogene, several convergent lines of research …
Cited by 3443 times according to google scholar
In the abstract.
“Originally considered to be an oncogene, several convergent lines of research have indicated that the wild-type gene product actually functions as a tumour suppressor gene 3- 9.”
Foundations well and truly laid.
References.
3. Finlay. C. A .. Hinds. P. W. & Levine, A. J. Gell 57, 1083-1093 (1989).
4. Eliyahu. D. et al. Proc. natn. Acad. Sci. USA. (in the press).
5. Sturzbecher. H.-W .. Addison, C. & Jenkins, J. R. Malec. cell. Bioi. 8, 3740-3747 (1988).
6. Munroe. D. G .. Rovinski, B .. Bernstein, A. & Benchimol. S. Oncogene 2, 621-624 (1988).
7. Wolf, D. & Rotter. V. Proc. natn. Acad. Sci. US.A. 82, 790-794 (1985).
8. Masuda. H .. Miller. C., Koeffler, H. P., Battifora, H. & Cline, M. J. Proc. Natn. Acad. Sci. US.A. 84, 7716-7719 (1987).
LikeLike
J Cell Biol . 2007 Nov 19;179(4):701-15. doi: 10.1083/jcb.200703040.
Mcl-1 determines the Bax dependency of Nbk/Bik-induced apoptosis
Bernhard Gillissen 1, Frank Essmann, Philipp G Hemmati, Antje Richter, Anja Richter, Ilker Oztop, Govindaswamy Chinnadurai, Bernd Dörken, Peter T Daniel
Affiliation
1
Department of Hematology, Oncology, and Tumor Immunology, University Medical Center Charité, 13125 Berlin, Germany.
PMID: 18025305
PMCID: PMC2080900
DOI: 10.1083/jcb.200703040
Problematic data figure 2B. Much more similar than expected.
LikeLike
“Bad luck in Cancer Research”.
Blood. 2005 Sep 1;106(5):1801-7. doi: 10.1182/blood-2004-11-4513. Epub 2005 May 10.
A rapamycin derivative (everolimus) controls proliferation through down-regulation of truncated CCAAT enhancer binding protein {beta} and NF-{kappa}B activity in Hodgkin and anaplastic large cell lymphomas
Franziska Jundt 1, Nina Raetzel, Christine Müller, Cornelis F Calkhoven, Katharina Kley, Stephan Mathas, Andreas Lietz, Achim Leutz, Bernd Dörken
Affiliation
1Charité, Campus Virchow-Klinikum, University Medicine Berlin, Augustenburger Platz 1, D-13 353 Berlin, Germany. fjundt@mdc-berlin.de
PMID: 15886325 DOI: 10.1182/blood-2004-11-4513
Problematic data figure 4B. Vertical, straight, truncated ends FL bands C/BPbeta panel, yet no vertical changes in signal in pS6 panel.
LikeLike
Reply to:
“Leonid Schneider
February 13, 2022
I informed Peter Daniel of these two papers.”
Bit more information for Peter Daniel.
“Bad luck in Cancer Research”.
PLoS One. 2012; 7(4): e34549.
Published online 2012 Apr 10. doi: 10.1371/journal.pone.0034549
PMCID: PMC3323539
PMID: 22506026
Mutual Regulation of Bcl-2 Proteins Independent of the BH3 Domain as Shown by the BH3-Lacking Protein Bcl-xAK
Michael Plötz, 1 Amir M. Hossini, 1 Bernhard Gillissen, 2 Peter T. Daniel, 2 Eggert Stockfleth, 1 and Jürgen Eberle 1 , *
Dhyan Chandra, Editor
Author information
1 Department of Dermatology and Allergy, Skin Cancer Center, University Medical Center Charité, Berlin, Germany,
2 Department of Hematology, Oncology and Tumor Immunology, University Medical Center Charité, Berlin, Germany,
Roswell Park Cancer Institute, United States of America
E-mail: juergen.eberle@charite.de
Conceived and designed the experiments: JE MP ES PD. Performed the experiments: MP AH. Analyzed the data: JE MP BG. Contributed reagents/materials/analysis tools: BG PD. Wrote the paper: JE MPE.
Problematic data figure 6.
Much more similar and different than expected after vertical stretching,
and vertical flipping and minor re-sizing.
LikeLike
Reply to:
“Leonid Schneider
February 13, 2022
I informed Peter Daniel of these two papers.”
Bit more information for Peter Daniel.
“Bad luck in Cancer Research”.
Figure 1 Int J Cancer. 1996 Jul 3;67(1):138-41 contains data very similar to data in J Clin Invest. 1996 Jun 1;97(11):2651-9 representing different proteins and cells.
FYI:-
Int J Cancer. 1996 Jul 3;67(1):138-41. doi: 10.1002/(SICI)1097-0215(19960703)67:13.0.CO;2-9.
Induction of the death-promoting gene bax-alpha sensitizes cultured breast-cancer cells to drug-induced apoptosis
C Wagener 1, R C Bargou, P T Daniel, K Bommert, M Y Mapara, H D Royer, B Dörken
Affiliation
1Max Delbrück Center for Molecular Medicine, Berlin-Buch, Germany.
PMID: 8690514 DOI: 10.1002/(SICI)1097-0215(19960703)67:13.0.CO;2-9
J Clin Invest. 1996 Jun 1;97(11):2651-9. doi: 10.1172/JCI118715.
Overexpression of the death-promoting gene bax-alpha which is downregulated in breast cancer restores sensitivity to different apoptotic stimuli and reduces tumor growth in SCID mice
R C Bargou 1, C Wagener, K Bommert, M Y Mapara, P T Daniel, W Arnold, M Dietel, H Guski, A Feller, H D Royer, B Dörken
Affiliation
1Max Delbrück Center for Molecular Medicine, Berlin-Buch, Germany.
PMID: 8647960 PMCID: PMC507353 DOI: 10.1172/JCI118715
LikeLike
Another two scientist group from Harvard with papers containing lots of problematic images.
https://hsdm.harvard.edu/baron-laboratory
https://www.pubpeer.com/publications/524BB49AB2CCF9738F8BF8516A7AC3#1
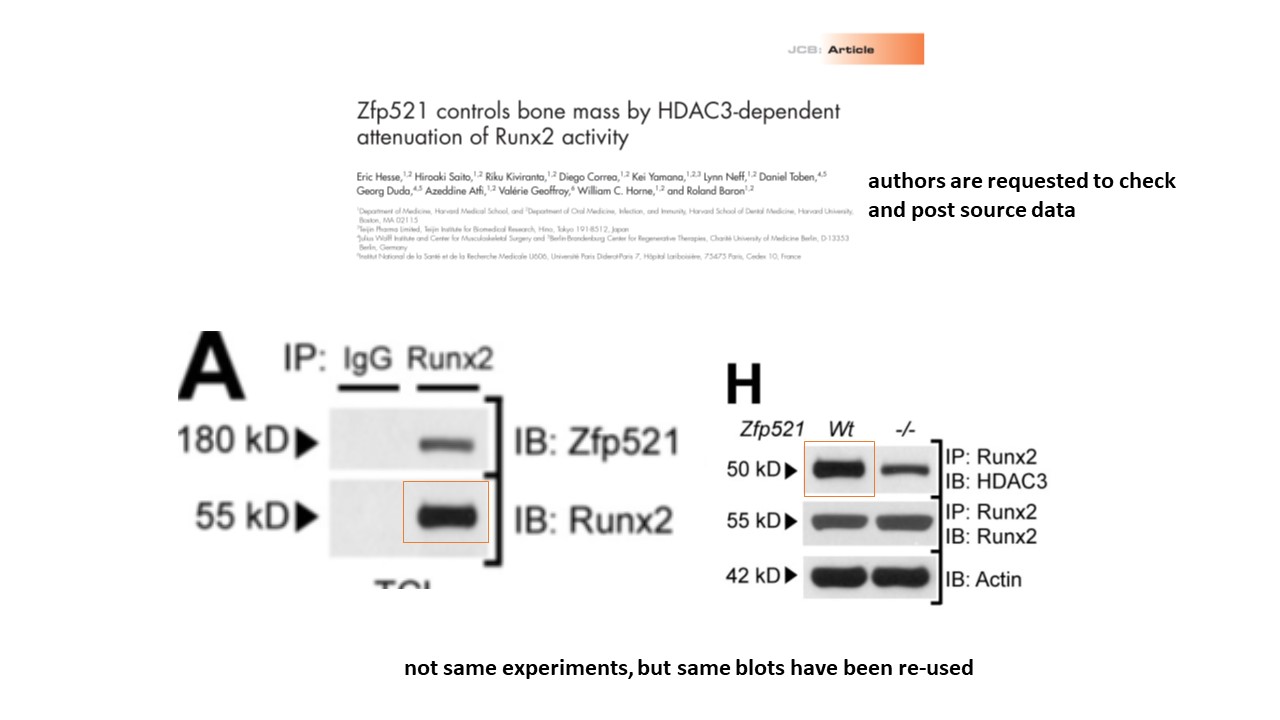



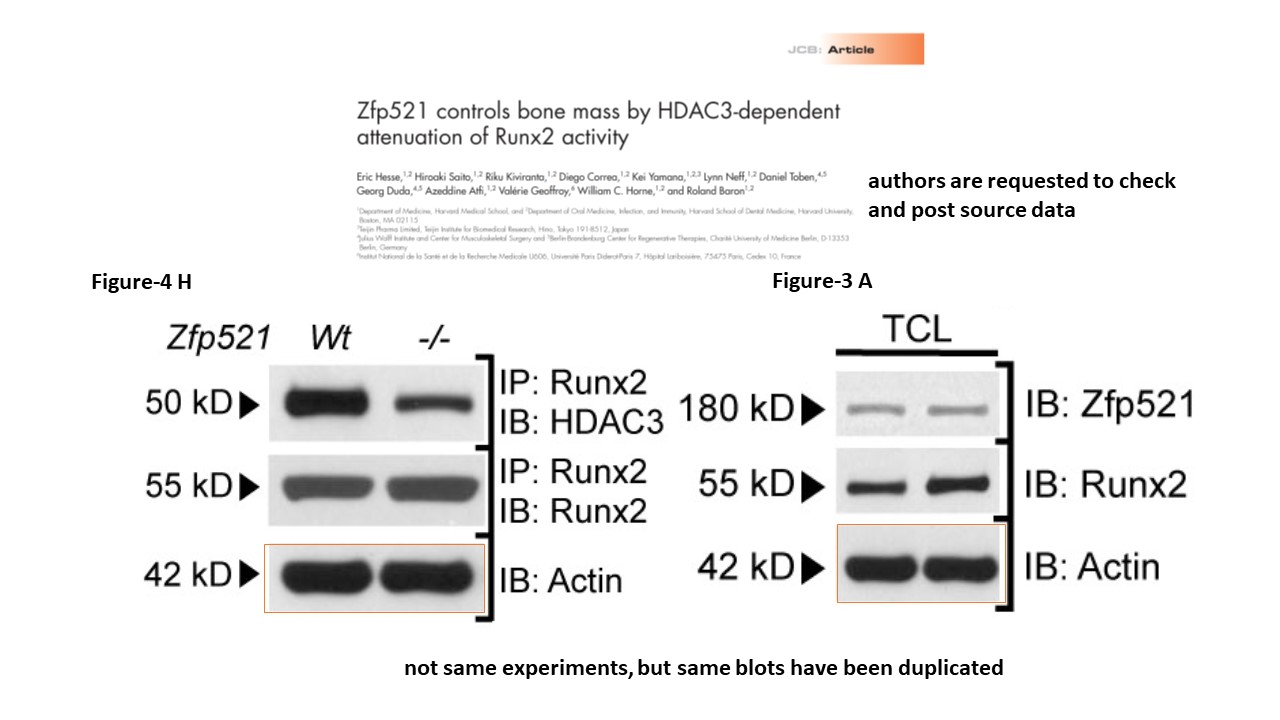
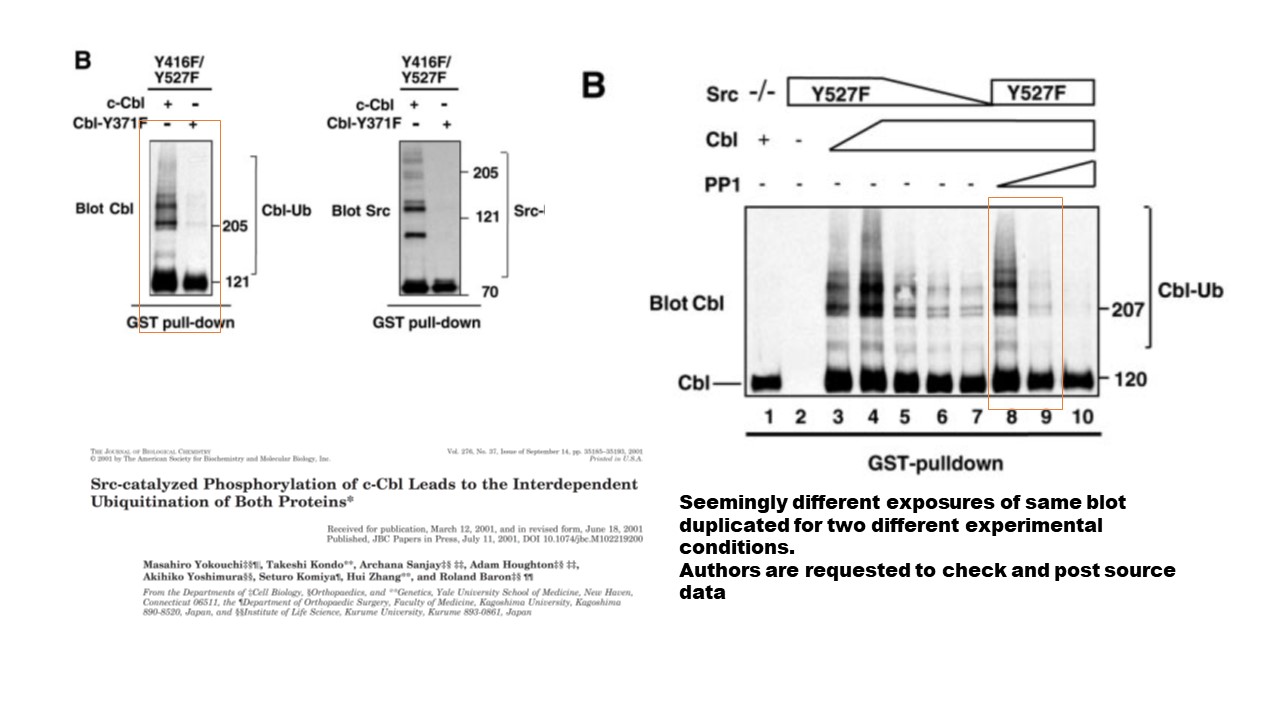
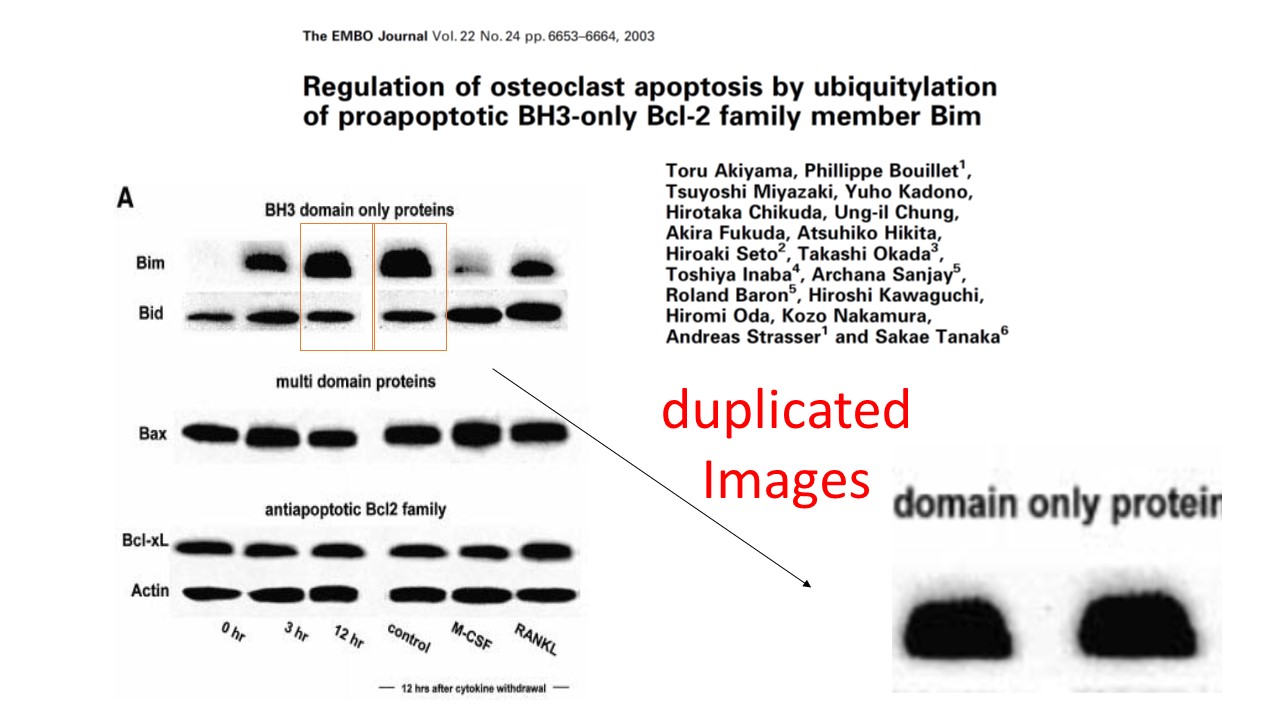
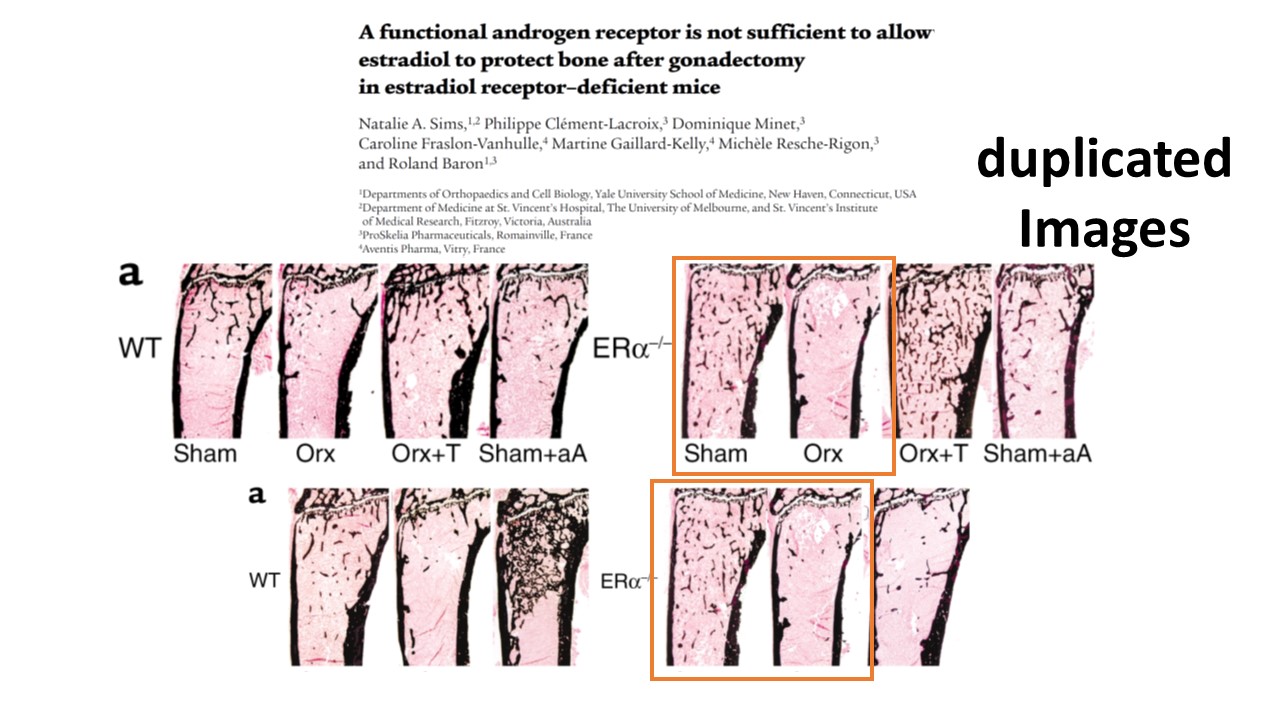
https://www.pubpeer.com/publications/5B0004BB5E0FEA751B5831C093FB4F
https://www.pubpeer.com/publications/A6FBADAF7347D5831049F387C5C12E
https://www.pubpeer.com/publications/F707EACA09BAD155743918BD98303A
LikeLike
More troubling images from the same author,
https://connects.catalyst.harvard.edu/Profiles/profile/1241265
LikeLike
“In any case, Hermeking states that these experiments were done by a “postdoc from the lab of Tal Mak in Canada“, i.e. the University of Toronto professor Tak Mak who also contributed this paper to PNAS as Academy member.
Update 10.02.2022. Now Tak Mak sent this postdoc to threaten me with a lawsuit also. This message just arrived, with Mak in cc, from Yuwen Su, now PI at National Health Research Institutes in Taiwan”
Tak Mak’s Pubpeer record. Perhaps he is not a great as he is cracked up to be.
https://pubpeer.com/publications/D29AA5A9F536B1498E6F2596D56D7A
https://pubpeer.com/publications/DA7A7647F2B8B39C0C0975D60E9DE3
https://pubpeer.com/publications/E1FBDE549A94C9F5A31B6C02B9E13D
https://pubpeer.com/publications/C8ECF3BABB1BCA31DBFC39F83D3401#1
https://pubpeer.com/publications/E9F7A7D43DE23462DC19E9235D764F
https://pubpeer.com/publications/B97147DA78A9982EF04EEA8DBFCA20#1
https://pubpeer.com/publications/06E69F4C563450DB5695D00715689A#1
https://pubpeer.com/publications/772D31E7E0538FB8AF5A8D7DC045FB#6
https://pubpeer.com/publications/E32A6C2019897001205ED25962E48C#1
https://pubpeer.com/publications/2B4278534852139EB5E4CD84A185A7#4
https://pubpeer.com/publications/A943FBC8525AA61C84DA5D4A646E34#1
https://pubpeer.com/publications/85275786B91CF866F1D03AF399E340
https://pubpeer.com/publications/4729C281F742107D09CDCA4D6BCA32
https://pubpeer.com/publications/D2187143725494B9CA948B36548D46
https://pubpeer.com/publications/B527488DDC43627DF00644AD9109B0
https://pubpeer.com/publications/AD9CBDD32ED22BD0D57F325F8CB27A
https://pubpeer.com/publications/788DFFA736DFE3931677ED20822077
https://pubpeer.com/publications/EA009AE50CC030324D24D752519816#1
https://pubpeer.com/publications/AF1E3082EE8E45E523B94153161A9F#1
LikeLike
14 April 2022 corection.
https://aacrjournals.org/clincancerres/article/28/8/1739/694175/Correction-Detection-of-miR-34a-Promoter
In the original version of this article (1), three errors exist in Fig. 1: splice lines were missing in both Fig. 1A and B, the same gel set was used to represent lanes M0-8 to M0-14 and M1-1 to M1-7 in Fig. 1B (lanes M1-1 to M1-7 are correct), and the figure legend referred to H2O instead of dH2O. The errors have been corrected in the latest online HTML and PDF versions of the article. The authors regret the errors.
Reference
1. Siemens H , Neumann J , Jackstadt R , Mansmann U , Horst D , Kirchner T , et al . Detection of miR-34a promoter methylation in combination with elevated expression of c-Met and β-catenin predicts distant metastasis of colon cancer. Clin Cancer Res 2013;19:710–20.
Only one more name on the original article, funny that is left off the correction: Heiko Hermeking
https://aacrjournals.org/clincancerres/article/19/3/710/208961/Detection-of-miR-34a-Promoter-Methylation-in
LikeLike
Pingback: Croce begat Calin, and Calin begat Girnita… – For Better Science
Another part of Bert Vogelstein’s legacy,
Lin “stock images” Zhang, University of Pittsburgh
Lin Zhang, Ph.D. | Integrative Systems Biology | University of Pittsburgh
Under Representative publications.
Zhang, L., Yu, J., Park, B.H., Kinzler, K.W. and Vogelstein, B. (2000) Role of BAX in the apoptotic response to anti-cancer agents. Science 290: 989-992.
Zhang, L. *, Zhou, W. *, Velculescu, V.E., Kern, S.E., Hruban, R.H., Hamilton, S.R., Vogelstein, B. and Kinzler, K.W. (1997) Gene expression profiles in normal and cancer cells. Science 276: 1268-1272.
Lin Zhang will easily get lost in the sea of people with Lin or Zhang as part of their name.
The legacy of image duplication.
https://pubpeer.com/publications/E6F4219D99316863E02F121FAD89A3#
https://pubpeer.com/publications/E156B6A5E7C78EB869611C8A53360E
https://pubpeer.com/publications/8A113C00960A3A2B45191C8422C140
https://pubpeer.com/publications/1AB711DC681C70BC5C9C464EFB99BF
https://pubpeer.com/publications/9C1EE92E042324FD770AF94FC0A5FC
https://pubpeer.com/publications/D9CE35EE9790C00FE904DCCFC792E2
https://pubpeer.com/publications/9F7EADFA23893108CA42E09EA536C0
LikeLike
Bert Vogelstein has 607 publications according to Pubmed
https://pubmed.ncbi.nlm.nih.gov/?term=vogelstein%20b&sort=date&ac=yes
Joseph Stalin is often credited with the aphorism that ‘quantity has a quality all its own’.
Presumably Stalin was talking about tanks or steel, or the people murdered during the political purges, about 700,000 (the party killing its own members), or the much larger number of people who died during the forced famines (in the Ukraine, Northern Caucasus, Volga Region, Kazakhstan, the South Urals and Western Siberia, it is estimated that 5.7 to 8.7 million people died). Vogelstein having so many publications has denied other people funding opportunities (similar to the forced famines) as funding is decided in the real world, where people are lazy, and only the count the number of publications, not their scientific contribution.
Quantity may be good for Bert Vogelstein, but is not good for others.
Vogelstein has become King of Cancer, when he has been made famous by the “adenoma to carcinoma model” of carcinogenesis , a derivative of the “two-hit hypothesis” proposed
(with evidence) by Alfred Knudsen, neither did Vogelstein discover oncogenes, tumour suppressor genes, or even the most commonly mutated gene in cancer p53, yet despite this, Vogelstein is King of Cancer.
607 publications make Vogelstein’s name difficult to avoid,
like Carol Croce with 1212 publications.
https://www.ncbi.nlm.nih.gov/search/all/?term=croce%20cm
LikeLike
At 754 publications Peter Carmeliet will never have to worry about having enough to eat..
https://www.ncbi.nlm.nih.gov/search/all/?term=Carmeliet%20p
He doesn’t keep kosher though.https://pubpeer.com/search?q=Carmeliet
LikeLike
Nature. 1999 Oct 7;401(6753):616-20. doi: 10.1038/44188.
14-3-3Sigma is required to prevent mitotic catastrophe after DNA damage
T A Chan 1, H Hermeking, C Lengauer, K W Kinzler, B Vogelstein
Affiliations collapse
Affiliation
1The Johns Hopkins Oncology Center, Program in Human Genetics, The Johns Hopkins University School of Medicine, Baltimore, Maryland 21231, USA.
PMID: 10524633 DOI: 10.1038/44188
Never too late for Nature for the right people!
https://pubpeer.com/publications/876EA56A4549BCAA87455EA00E5A19#77
Elisabeth M Bik commented August 2023
Correction 24 years after publication.
Correction, 22 August 2023: https://www.nature.com/articles/s41586-023-06446-1
“A possible duplication of the top panel in Fig. 2a has been brought to our attention. As the source data for the images are no longer available, we have repeated the control with the cells frozen in 1999 and thawed 23 years later. This new experiment showed that the nuclear morphologies of cells with and without 14-3-3σ in the absence of DNA damage induced by adriamycin are indistinguishable, as shown in Fig. 1 of this correction. Figure 3d of the original paper also shows independent confirmation of the indistinguishable nuclear morphologies.
We also wish to clarify that the legend of Fig. 4a should have stated that the sequential anti-cdc2 and the anti-cyclin B1 stainings in the right panel represent different orientations of overlapping fields of the same cells. Figure 2 of this correction shows the staining when the fields are aligned, and at higher resolution than in the original. Figure 4a was not intended to show co-localization; co-localization between cdc2 and 14-3-3σ was shown at high magnification in the original Fig. 4c.”
LikeLike
That’s reproducibility! Experiments from a quarter a decade ago, easily redone and reproduced!
What happened to Bert’s and Heiko’s plans to sue people?
LikeLike
“Experiments from a quarter a decade ago, easily redone and reproduced!”
Don’t you mean a quarter of a century ago? 1999-2023 is 24 years, nearly a quarter of a century.
A decade is 10 years.
LikeLike
Correct, wrote too fast.
LikeLike
Possibly more problematic data Nature. 1999 Oct 7;401(6753):616-20. doi: 10.1038/44188.
Not all of a piece, and no loading control.
https://pubpeer.com/publications/876EA56A4549BCAA87455EA00E5A19#80
LikeLike
Fresh problematic data for Heiko Hermeking:
PubPeer – Paracrine Induction of Epithelial-Mesenchymal Transition Bet…
LikeLike