In 2014, Institute of Plant Genetics of the Polish Academy of Sciences in Poznan received a Horizon 2020 grant from the European Union, in value of €2 million. The project Bio-Talent is designed to set-up a new Department of Integrative Plant Biology, one of the three key scientists involved is a certain Gregory Franklin, and his highly productive publication output will make a large share of Bio-Talent’s final report to the EU, when the grant ends in June 2019. There is just one snag: the Franklin nanotechnology lab produces fabricated data, made up in Photoshop or from creative reuse of unrelated published material. Another recurrent name is Franklin’s newly arrived PhD student and member of EU-funded Bio-Talent team, Qaisar Maqbool. He even has his own stand-alone papers consisting of fabricated data. Some Bio-Talent it sure is, but is it really what EU wanted to be funding in Poland?All this was analysed and posted on PubPeer by a reader of my site, and the case started with a recent Frontiers paper from Pakistan, where the topic editor Franklin somehow apparently appointed himself as last and corresponding author: Maqbool et al, Frontiers in Pharmacology 2018. The paper is a classic green nanotechnology compost heap slurry: you make some boring metal nanoparticles, but with herbal favour, usually using a plant of complementary medicine “value”, and declare those “green” nanoparticle to efficient killers of cancer cells or multi-drug-resistant bacteria. What Maqbool and his colleagues back in Lahore, Pakistan, claim to have discovered was that CuO and CeO2 nanoparticles flavoured with leaves from olive tree (I guess they tried to peddle to the Mediterranean market) kill bacteria and are absolutely non-toxic to eukaryotes.
The road to get the desired results with such green nanotechnology is usually sloppy science, like accidentally not using same dosages of toxic metal nanoparticles in controls versus herbal setups in vitro or even in vivo (most journals never ask for animal research ethics permits), and if everything else fails: data manipulation in silico. This is apparently what the Poznan authors had to resort to. Electron microscopy images of nanoparticles were generated in Photoshop, with bits and pieces cloned and reused. Same groups of nanoparticles appear several times in the same image, as the PubPeer commenter said:
“A1 is festooned with little repeating elements akin to a dog’s bollocks. It appears to be a digital composition”.
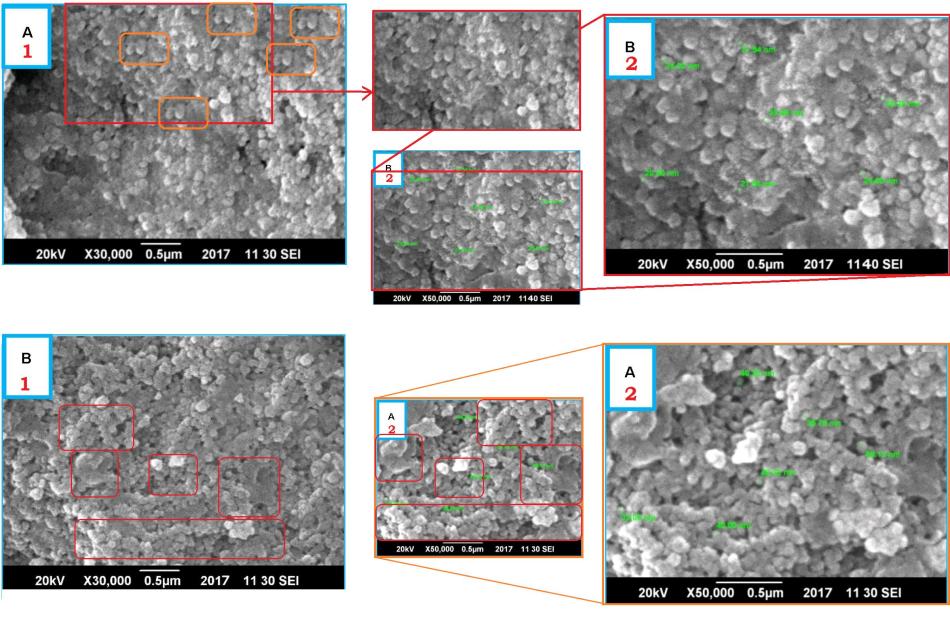

Franklin’s contribution to this masterpiece was two-fold: he “provided his expert opinion and technical expertise related to the accomplishment of biomedical application part“, being Maqbool’s new boss and Frontiers topic editor who invited that masterpiece. Which brought Franklin the practical last authorship, to complement his Bio-Talent report to EU Commission, who then paid the article processing charge of $2950 for a fraudulent paper from Pakistan. Everyone is happy: young Mr Maqbool, his new master Dr Gregory, Frontiers, their academic employer in Poznan, the EU Commission and so should you be. This is namely the top peer-reviewed science you get:
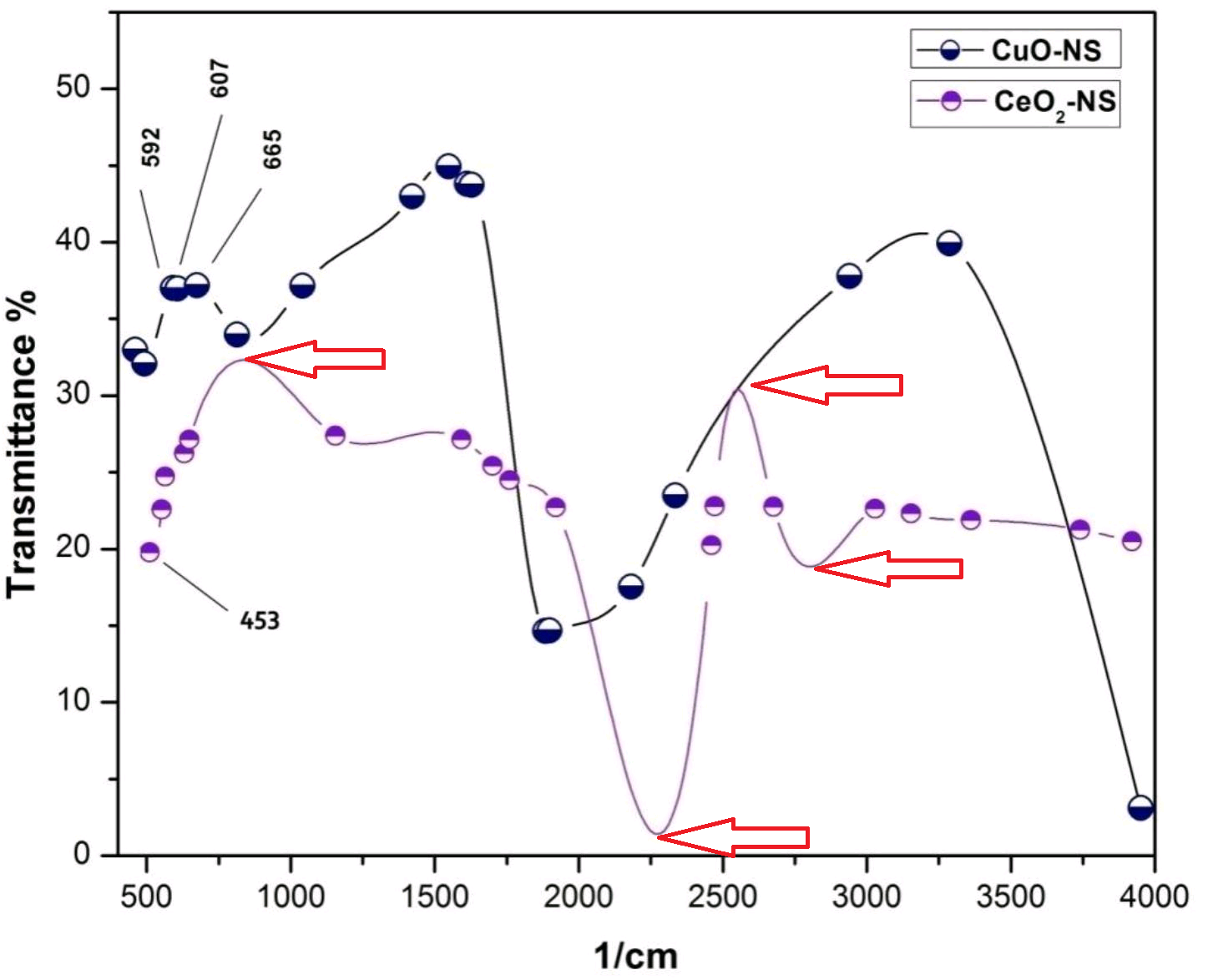
Apparently Frontiers “rigorous peer review” decided that yes, one can invent peaks and troughs of a spectrum without any support from actual data points. This is what the authors did, for the spectrum of CeO2-olive-leaf-flavoured nanoparticles. This kind of data fabrication is in plain sight for editors and peer reviewers, there are absolutely no excuses to have missed that.
If you think at least the CuO spectrum might be trustworthy: it is not. That one, just like the HPLC chromatogram from Figure 2 or the X-Ray diffraction (XRD) analysis from Figure 5, were dug up from older Maqbool papers, Maqbool RSC Advances 2017, and Maqbool et al IET 2017.
Not just the spectra, diffraction and chromatography analyses, also electron microscopy images in that Maqbool-Franklin Frontiers co-production were reused from older material: Anwaar et al Frontiers in Plant Science 2016 (that one might lack Franklin as author, yet it was edited by his colleague in Poznan), and the solo masterpiece, to qualify as Franklin’s PhD student, Maqbool RSC Advances 2017. In the Anwaar paper in Frontiers, the images used to stand in for nanoparticles designed to help rice grow, for which they had to be prepared using neem leaf extract (Azadirachta indica), because neem in Bengal area is traditionally eaten with rice, so it makes perfect sense.
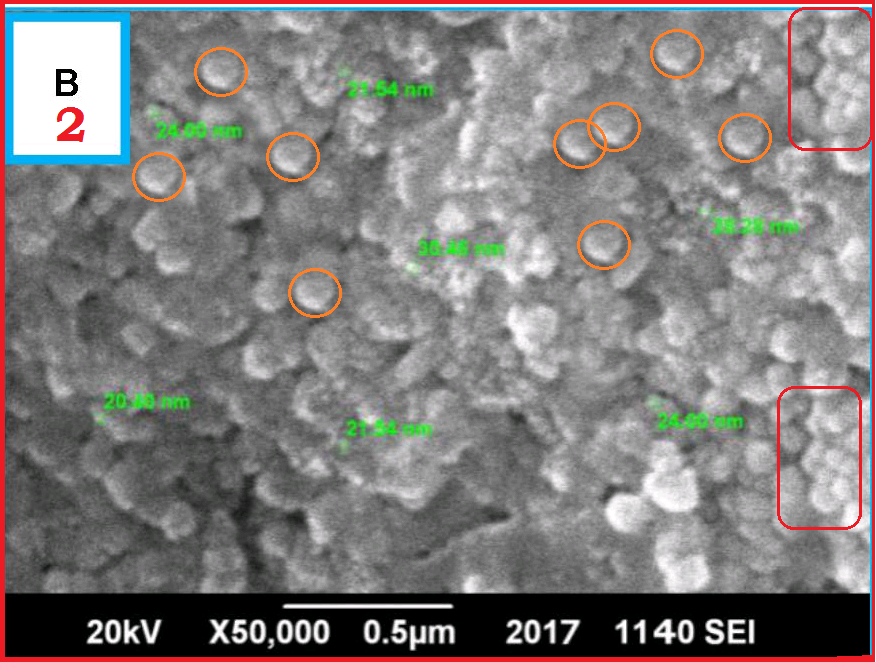

The strange Maqbool RSC Advances 2017, authored solely by the young student Mr Maqbool, about to begin his PhD under the great Franklin, is full of surprises itself. Look at this thermogravimetry analysis (TGA) in Figure 4:
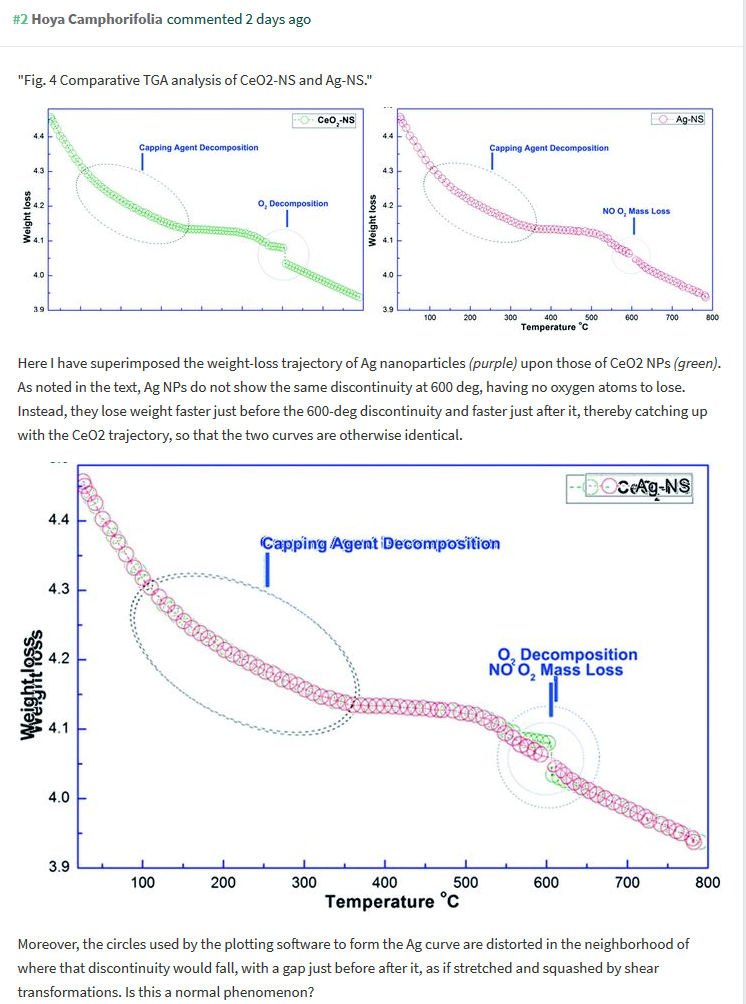
It is peculiar how similar those curves are. If one overlays them, only a tiny wiggly bit at the “O2 decomposition” point is different, which must be a clever optical psychological trick to confuse the reader. And it gets better. One can also overlay these two with two more curves, for organometallic (OM)-Ag nanoparticles from Fig 7 of Maqbool et al RSC Advances 2018 (last author is Franklin), and for CeO2 nanoparticles from Maqbool et al IJN 2016, Fig 9. They all fit neatly on top of each other, because they are obviously the same measurement.
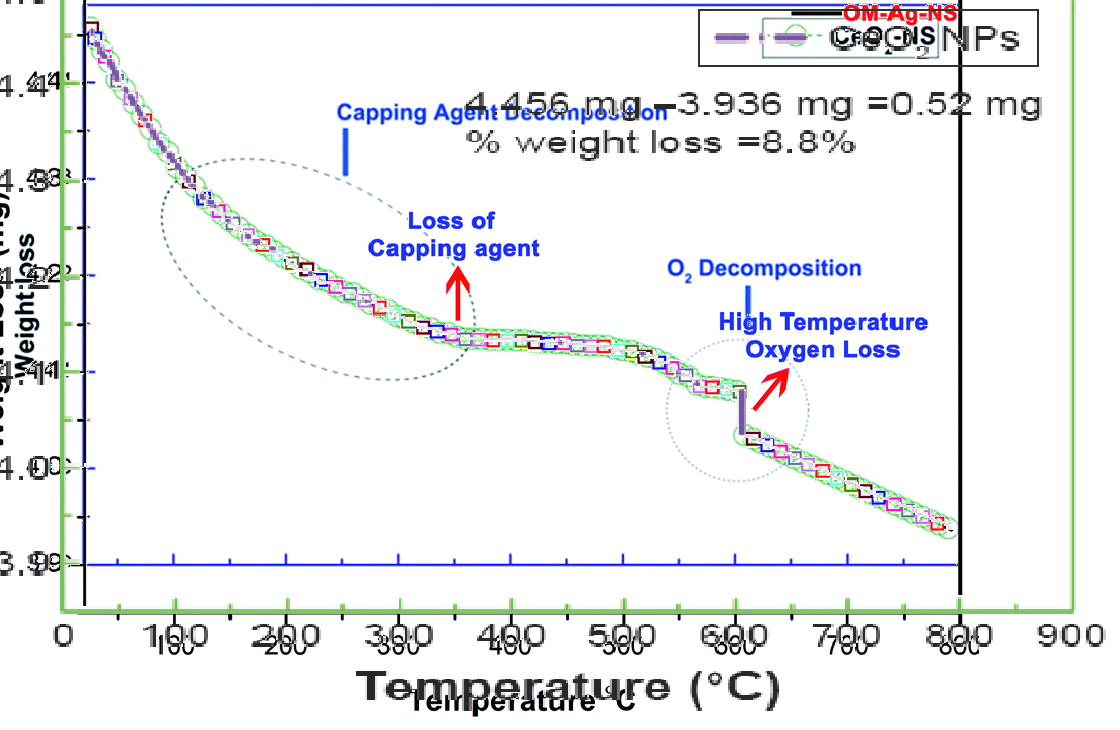
One can only guess where the original curve comes from, and what it originally showed, and who might its original author be. For sure we now understand the secret to Mr Maqbool’s and Dr Franklin’s amazing productivity. The Maqbool et al RSC Advances 2018 paper has another recycled graph inside Figure 7, where the spectrum for St John’s-wort extract (mind your medicinal herbarium!) miraculously proves exactly identical to that of organometallic silver nanoparticles, just a bit shifted and squeezed.
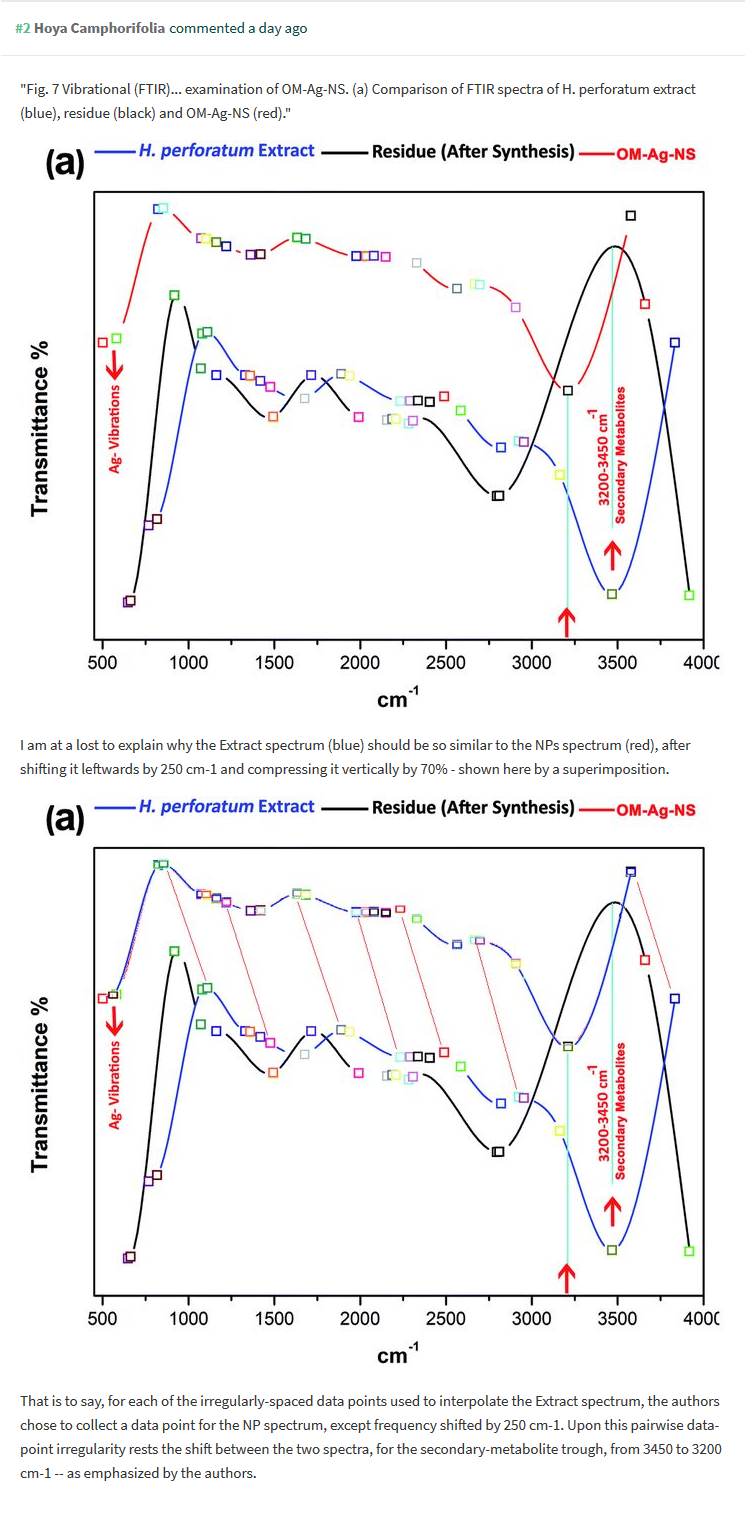
Of course also that fake compost dump of a research paper claims to kill drug-resistant bacteria with nanoparticles, this time St John’s-wort flavoured (to cater to northern Europeans, presumably). There is hope: the Editor in Chief of RSC Advances, Russell Cox, promised action on those two Maqbool papers:
“I will forward these points to our editorial office with a request that the papers in question are examined as quickly as possible. For your information I am an expert in the chemistry of fungi, and have little expertise in materials chemistry. RSC advances publishes thousands (literally) of papers each year and it is almost impossible for us to screen every image during the review process. We thus rely on the broader chemistry community to point out potential failure of ethical behaviour in authors, and I thank you for your vigilance in this matter. We have an established procedure for dealing with cases of possible fabrication and duplication which we will follow in this case”.
One should not assume Franklin always needed Maqbool to produce such original pseudoscience. In this regard, see the works of Franklin as last author from his earlier stint in Portugal, like Marslin et al Colloids & Surfaces B, 2015. There is a very interesting image duplication in it:

The paper Marslin et al Planta Medica 2016 also has an image duplication which is unlikely to have happened by oversight. On the other hand, the paper Maqbool et al Int J Nanomed 2016 misses Franklin, but has same kind of duplicated and re-sized image, yet size bar remained same. What a nightmare team those two make.
Almost all those analyses were made and posted on PubPeer by a certain reader of my site, who pretends to be a waxy plant, Hoya camphorifolia. Let us hope Franklin and Maqbool won’t turn poor Hoya into some medicinal nanoparticles next, for revenge, and publish it in Frontiers, paid with EU grant money.
Another reader of my site wrote to me to point out that a recent preprint by Maqbool and Franklin also contains fake data, Sidorowicz et al MDPI Preprints 2018. It is also about St John’s-wort. The electron microscopy image was made by copy-paste in Photoshop. For all we know, those St John’s-wort flavoured silver nanoparticles might be a cloned picture of a hamburger.
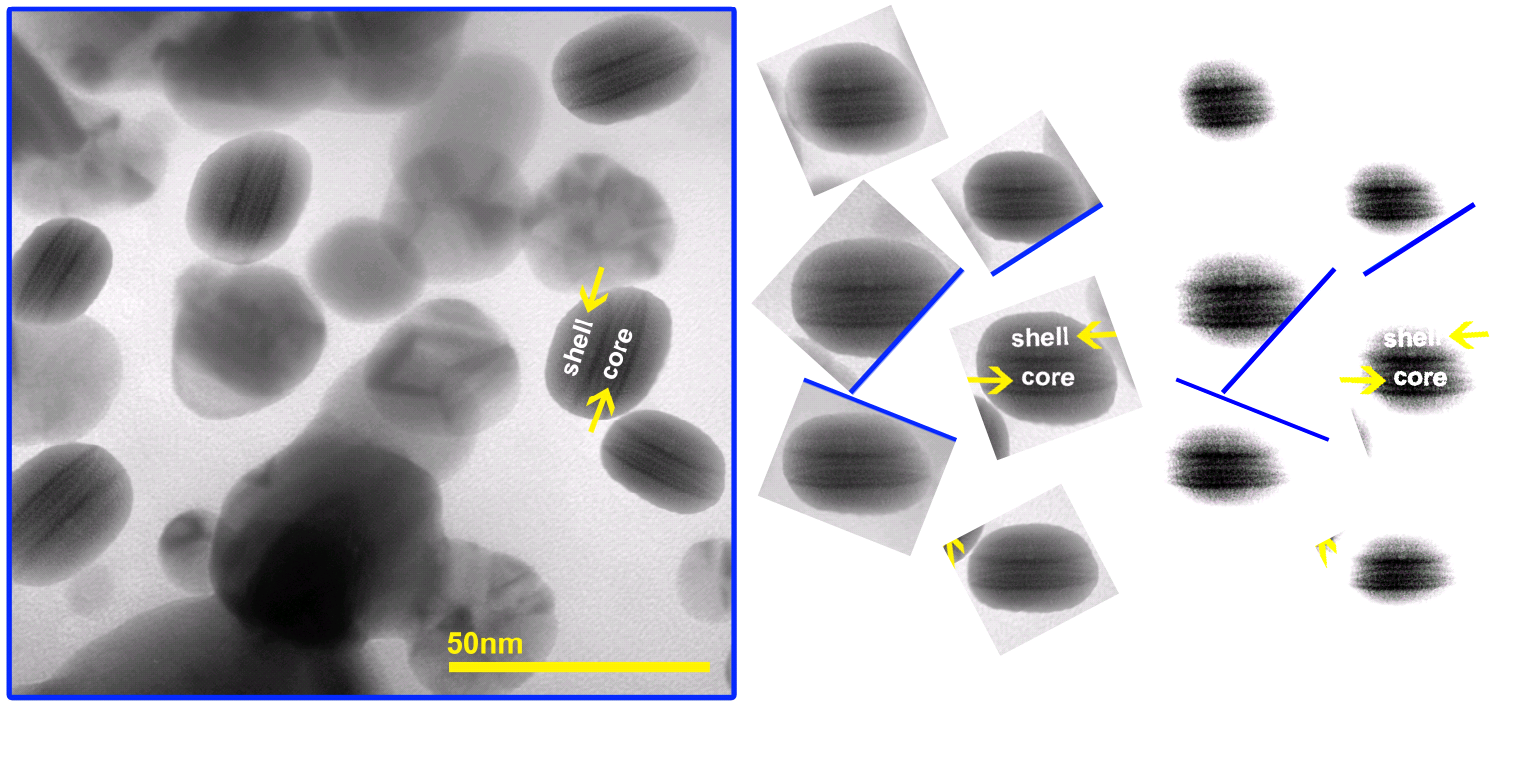

Once again, the authors invented some peaks. This time at least, one cannot blame any peer reviewers for approving that, this is still a preprint.
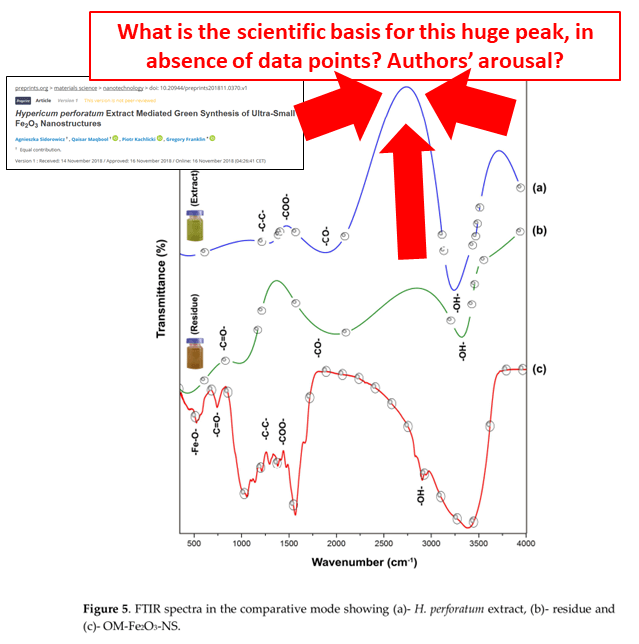
Carnivora!
The initially discussed Frontiers in Pharmacology paper Maqbool et al 2018 was part of a research topic “Nanoformulations containing Phytochemicals/ Herbal extracts“, organised in the section “Ethnopharmacology” by Franklin and two friends. What other amazing works did it provide the scientific community with, before the submission closed? A mini-review, and a paper from Gdansk, Poland: Krychowiak et al 2018, that’s it. This “original research” paper, where silver nanoparticles were flavoured with extracts from carnivorous plants of sundew family, has an interesting backstory. For one, the co-author Anna Kawiak from the University of Gdansk had to retract her study Kawiak & Domachowaska PLOS One 2016 , because it was too fraudulently photoshopped. That paper claimed to have established anti-cancer properties of carnivorous plant extracts, there is another one on same topic in same journal, Kawiak & Lojkowska 2016 which PLOS chose to do nothing about, despite horrendous data manipulations found there. Here is just one example, the duplicated background of western blot image suggests that a gel band was digitally superimposed:
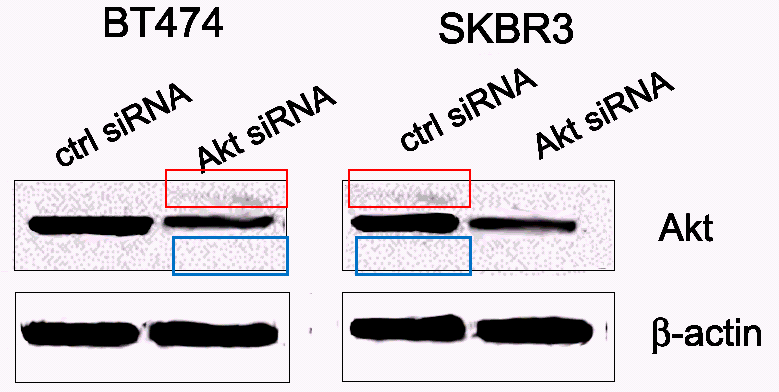
You might wonder why would someone so desperately want to prove that sundew can cure cancer as to resort to outright data fakery. Sure, nanoparticles flavoured with medicinal plants, nanotechnology meets Ayurveda, that is cool and fashionable, but extracts from carnivorous plants to cure cancer? How does one arrive to that conclusion? My regular contributor Smut Clyde blogged about that topic some years ago:
“Despite a name that is redolent of the homeopathic pharmacopeia, it turns out that “Carnivora” is not a highly-diluted preparation of big cats, rabid mustelids, pinnipeds and hyaenas,* to be taken as a counter-agent to the effects of partial consumption by tigers”
We learn that this school of silly thought is called “Carnivora”, first postulated by some German nutcases in the 1980ies. Soon, more nutcases in Germany and other countries including Poland joined the craze, because if a carnivorous plant like sundew or Venus flytrap can digest a fly, it perfectly stands to reason that it would also dissolve cancer. Please don’t shake your head incredulously now, for this theory has been approved and peer reviewed by none other that Frontiers! Marc Diederich and his colleagues in Luxembourg proved in the Gaascht et al Frontiers in Oncology 2013 that yes, Venus flytrap extract is bound to cure cancer. It is just an opinion piece moonlighting as a literature review, but who needs reliable research data to postulate fringe discoveries, that being Frontiers?
It makes perfect sense that that Frontiers topic contains just these two complimentary “research” papers on green nanotechnology: Franklin & Maqbool copy-paste orgy of fake data, and Kawiak’s potentiation of Carnivora quackery with nanoparticles. At least the chief editor of the Ethnopharmacology section at Frontiers in Pharmacology, Michael Heinrich, promised to take care of the Franklin case. He wrote to me:
“Thanks for bringing this to our attention. I have contacted the Research Integrity Department of Frontiers and they will investigate immediately. This will take a few weeks and we will get back to you afterwards. I cannot comment without further data, but want to draw your attention to our ‘Four Pillars of Best Practice’ which we also use to evaluate MSs (and as a key guide to authors) specifically in the area of ethnopharmacology: https://www.frontiersin.org/journals/pharmacology/sections/ethnopharmacology#about). Here we define in detail what constitutes best practice for manuscripts submitted to Frontiers in Pharmacology; Section Ethnopharmacology. They build on the general requirements of Frontiers in Pharmacology”.
Under the provided link we learn:
“Research must be certified by peers before entering a stream of knowledge that may eventually reach the public – and shape society. Therefore, Frontiers only applies the most rigorous and unbiased reviews, established in the high standards of the Frontiers Review System. Furthermore, only the top certified research, evaluated objectively through quantitative online article level metrics, is disseminated…”
That is reassuring. Kawiak’s Carnivora and Franklin’s fake data, even his blatantly made-up data-free peaks all passed the highest quality control in the world of science, the Frontiers peer review.

Donate!
If you are interested to support my work, you can leave here a small tip of $5. Or several of small tips, just increase the amount as you like (2x=€10; 5x=€25). Your generous patronage of my journalism will be most appreciated!
€5.00


I also wonder what is the opinion of the scholarship program which supports the studies of the PhD student. According to the BioTalent website, he “is a Gold medalist in MSc-Biology and won 2018 outstanding research paper of the year award by HEC (Higher Education Commission, Pakistan)”. Does HEC check the quality of papers before giving an award? Do they suspend a scholarship in such a case?
LikeLike
Some high level irony at Bio-Talent:they invited various plant scientists as seminar speakers. Among them, Vicki Vance, the original whistleblower in the Olivier Voinnet fraud affair, Edward Farmer who used to be one of the two external investigators of Voinnet at ETH, and Cyril Zipfel, former director of The Sainsbury Lab, whose task was to deal with the aftermath of Voinnet’s fraud there.
How did Franklin feel meeting these people? Or maybe everyone knew and they were invited “ironically”?
http://www.biotalent.eu/index.php/newsletter
LikeLike
Thank you for sharing. I am deeply concerned about the data manipulation issues in Anna’s papers. We collaborated and I sincerely hope that this matter will be addressed promptly. In joined paper in CDDis we overlooked the dublication made by a mistake and issued correction asap
LikeLike
Thanks for the link to Riddled!
I was wrong there to date ‘Carnivora’ to 1987. That was just when Keller applied for a patent, and published his made-up cure result. He had been selling the stuff from the late 70s on, and the Psiram article on Carnivora links to articles in Der Spiegel and Die Zeit from 1985, about the consumer base Keller had constructed by then.
https://www.psiram.com/de/index.php/Carnivora
So it has a 40-year history rather than 30 years, but still quite recent. Even so, it has been adopted in the ‘Green Mysticism’ school of thought, who treat it on a par with the Time-honoured Traditional Wisdom of Medieval herbalism from different cultures.
This makes it difficult to be sure which promoters flytrap-juice are simply on the payroll of “Big Carnie”, and which are sympathetic to cancer scams because their brains are wired up funny.
Among the former group, we find Todorov and Iliaranova, who wrote a great profusion of papers about the active ingredients in Flytrap Juice and how well they killed cancer in test-tubes… most of them published as in-house reports for Carnivora-Forschungs GmbH.
In the latter group, I keep coming across the names of Kreher, Wagner, Jurcic and / or Neszmelyi, who were publishing from the late 80s onwards. Perhaps they read Keller’s completely fictitious cure rates, believed them, and have been trying ever since to work out which constituents of flytrap produce the non-existent cures. You quote them as “more nutcases in Germany”. There is a strong Mitteleuropea / Eastern-Europe trend to the Green Mysticism tradition (Todorov and Iliaranova were Bulgarian).
Anyway, Kreher & Wagner cite Keller (1987) in reverential terms. Diederich cites Kreher & Wagner, so he must be aware of Keller’s position as the instigator of the whole Carnivorous-plant / Cancer Cure scam. He is careful never to cite Keller directly, though, so he must also be aware that Keller’s name brings undesirable associations of “unscrupulous lying charlatan”.
With all that in mind, it would be interesting to know more about the unspecified Third Party who provided Kawiak with fabricated data supporting sundew juice as a cancer cure. The initial suspicion is that she outsourced experimental work to an outside laboratory who cheated her, but it could also be that they were employing her.
LikeLike
I forgot to say… it was a little disingenuous of Diederich to publish his Flytrap paean in “Frontiers in Oncology”, without noting that it was effectively an advertisement for a commercial product (the subject of extravagant curative claims). To write a history of traditional uses for flytraps without mentioning the central role of Keller and his cancer scam, is a real Hamlet-without-the-Prince-of-Denmark situation.
LikeLike
On the peaks: I would say those are results of polynomial interpolation. Note how there is a pair of points very close to each other on the horizontal axis, but quite different on the vertical axis, next to the bigger peaks/troughs. A polynomial passing through both points must have a very steep tangent line (high derivative), and, since polynomials are smooth, it will cause a big peak or trough next to such a pair of points.
Such naive polynomial interpolation of noisy data is not a good practice. Precise polynomial interpolation of noisy data will always cause such problems; there are problems even with non-noisy data. Wikipedia is helpful here: https://en.wikipedia.org/wiki/Polynomial_interpolation#Convergence_properties
Possible solutions would be smoothing out the data before interpolation (to remove noise), using splines or other more local interpolants, or using some kind of regression or regularization.
If the peaks or the troughs are used in arguments in the paper, than there is a definite problem.
LikeLike
The red curve here (c) is the weirdest polynomial interpolation I have seen.
Also compare curves (a) and (b) over at the extreme right, where they have the same data points; a peak has been interpolated between 3500 and 4000 cm-1 for (a), but not for (b).
LikeLike
The curve (c) is doing something else. It seems that the curve is perturbed towards the average of the curve between some of the points, but is not quite that, either. I have no idea what is happening there. It might be a polynomial (or Fourier approximation) of very high degree, but looks quite strange for that, too. I do not know.
The rightmost interpolation in (b) does look suspicious; I would have expected it to be more rounded, but only a little. The three points to the left of it are not on a line, but rather in a position which should make it fairly easy to draw a smooth polynomial without a peak so that it reaches the rightmost point. In (a), the configuration of the four points does not seem as conductive for drawing a gentle curve towards the rightmost point there.
The method to test this hypotheses of polynomial interpolation would be to try to repeat it. I am not quite interested enough to do that, but would be interested in the results if anyone does so.
LikeLike
Just for clarification (c) is actually the only graph in that plot that actually looks like an ir transmittance spectrum if you ignore the points. Maybe some protein or tissue powder, because it shows amide I and II bands between 1500 and 1700 and C-C vibrations all around 1000 wavenumbers. It might even be an actual spectrum from hypericum. Just look up some ir spectra in the nist database and you can see the typical shapes. I guess the points placed on the line are just randomly added. The green and blue graph look like somebody tried to connect the points by a spline, except for the two green points above 3700 1/cm, just like you can do with plotting graphs in excel. And splines are a no go in data presentation. So it’s a really bad graph even if it actually was real data resembled by those points.
LikeLike
… as an organic chemist, I have seen a plethora of IR spectra, and I do not understand at all, how any sane reviewer could have passed the papers with these IR spectra of Maqbool. He claims to have used FTIR, which stands for Fourier-transform IR – the signal is generated by using fourier transformation on the raw data, and there is no way how you can get several isolated points with random displacement by this method (you get a continuous-looking curve depending on your sampling, but definitelly not 10-20 randomly placed points). And mind – this is not done by user, but by the machine, you only get the data AFTER the FT, so I simply cannot see a way how this “data” was not simply faked by inventing some numbers. If Mr. Maqbool can show me how – he may as well cut off my legs and call me a shorty.
Btw. the huge “erect” peak in the spectrum is actually a lack of signal (it shows transmittance), not that it makes any difference…
Who cares about the frontiers, but RSC? Honestly, I’m dissapointed.
LikeLike
You may enjoy this version from Maqbool in RSC Advances, where transmittance ranged from 10% down to -20%.
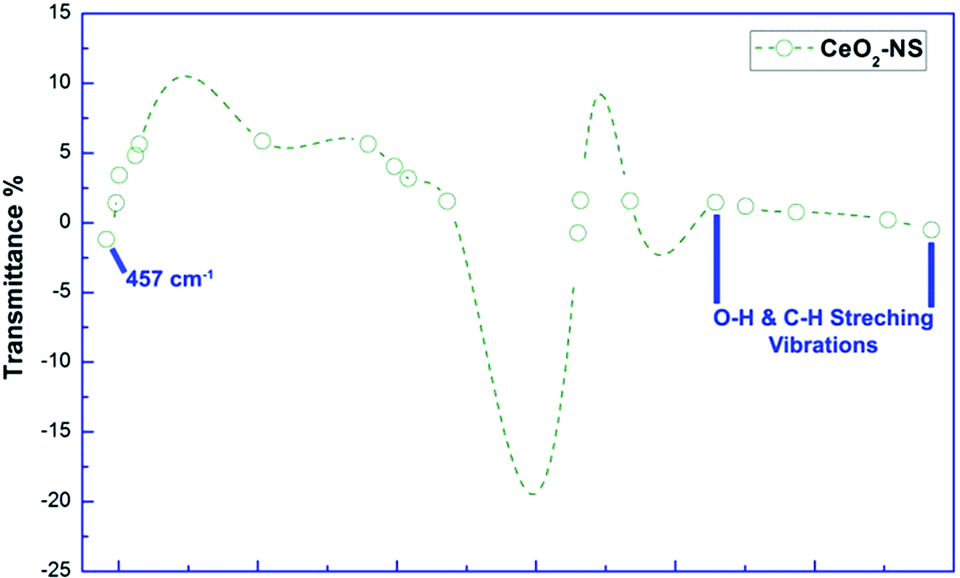
LikeLike
… bloody hell, I missed that one. So it’s a CeO2 laser, if I understand this correctly?! But anyhow – the crew at the RSC adv. are chemists, the editors were presumably chemists… so how did this happen??
LikeLike
So I’m going out on a limb here, but wouldn’t negative transmittance be luminance? Cold fusion, perhaps?
LikeLike
These clowns can’t even photoshop convincingly, but we have to give them credit for gaming a system that clearly has no capacity for due diligence. And winning awards, too. One might wonder what the genuinely half-bright could accomplish in such an ecosystem, but I think we have ample evidence of that as well in these pages. But eventually they get greedy, sloppy and exposed: politically and scientifically. Ironic that the same company that created Photoshop, the chosen tool of dumb fakers, also came up with Acrobat, the Achilles’ heel whereby they are eventually discovered and discredited. I salute the efforts of the nanotech sleuths, and I marvel at how they do not just become depressed like I do when wading through this dreck. “Royal” chemistry society indeed – who’s their head, Prince Phillip?
LikeLike
I salute the efforts of the nanotech sleuths, and I marvel at how they do not just become depressed like I do
F X Coudert is indefatigable.
LikeLike
Again, I salute the indefatigability of those who persist in fighting the good fight for science, against the opposition of bad authors, the lousy reviewers and editors who enable them, and the crap journals that publish them (at least the latter have a monetary incentive). I love the erratum for the paper flagged in the tweet: we used the wrong figure – as if there could ever be any excuse for generating a Frankenstein’s photoshop monster like the original (which was apparently used several times). Luckily most of the real garbage is paywalled. If we all only cited open access papers this stuff would evaporate in the dark where it belongs.
LikeLike
Open access like Frontiers and PLoS One I assume?
LikeLike
I have some good experience with sending expression of concerns to the research institutes and journals together with an easy-to-understand presentation with figures of the problematic issues, and I will encourage you to do the same. I am talking about cases where you have a longer list of concerns. Presentation can be made by simply collecting figures from Pubpeer (including a link), but often they need to be improved to make it easier to understand the problem. This is of course time consuming and if that’s a problem, an alternative way could be to present links to pubpeer posts/blogs only.
Unfortunately the information from Pubpeer and Leonids’s blog is not always reaching the research institutions and journals involved, and it’s easier to convince the journal editors and research institution leadership that something is terribly wrong by providing a full presenation of the case. If they receive an expression of concern, public institutes usually have to address the problem.
LikeLike
In response to the possible serious scientific misconduct made by Dr Franklin Gregory and his PhD student MSc Qaisar Maqbool we would like to express our deep concern and assure that we are investigating this issue closely and the outcome of our investigation will be provided soon. We would like to make the statement that whereas it is our responsibility to maintain the highest ethical standards of scientific work it is difficult to track the possibility of data manipulation in publications. The research projects of Dr Franklin are not funded from the BIO-TALENT project and Mr Maqbool is not employed from this source of funds. Dr Franklin leads an independent research team, therefore we appeal to not link the activity of all other scientists involved in other research areas in our Institute as well as the BIO-TALENT – funded department with possible misconduct. The case of Dr Franklin and his PhD student will be studied individually and the findings of this investigation will be released promptly and appropriate actions taken.
Sincerely yours
Prof. Bogdan Wolko
Director of the IPG PAS
LikeLike
Dear Prof Bolko,
many thanks for your reply, as you know I tried to reach you by email before. Usually, when a team is listed like this, one assumes they all receive funding from the relevant grant:
http://www.biotalent.eu/index.php/biotalent-team
Indeed Mr Maqbool is not listed, but then again he arrived to the group only recently.
http://www.biotalent.eu/index.php/newsletter
In other news, I saw that the authors withdrew their fake preprint:
LikeLike
We would like to inform all interested parties that, as a consequence of our internal investigation Mr. Qaisar Maqbool has claimed his responsibility for the manipulation of graphical documentation and using data from untrusted sources in the questioned works. Dr. Franklin Gregory claims, that he was not aware of the fact that data was manipulated. As a consequence, Mr. Maqbool’s appointment with the Institute will be terminated. In order to avoid similar problems in the future, the further work of Dr. Franklin Gregory will be supervised by the Institute authorities. At present we are contacting editors of the appropriate journals to provide corrections or retract the following articles:
Maqbool Q., Kruszka D., Kachlicki P., Franklin G. (2018) Organometallic Ag nanostructures prepared using Hypericum perforatum extract are highly effective against multidrug-resistant bacteria. RSC Advances, 8: 30562-30572.
Maqbool Q., Nazar M., Maqbool A., Pervez M.T., Jabeen N., Hussain T., Franklin G. (2018). CuO and CeO2 Nanostructures Green Synthesized Using Olive Leaf Extract Inhibits the Growth of Highly Virulent Multidrug Resistant Bacteria. Frontiers in pharmacology, 9, 987.
The MDPI preprint has been retracted.
As a research institution we would like to say, that whatever happened is devastating to us. We would also like to declare that none of us inspired or motivated this misbehavior.
Prof. Malgorzata Jedryczka
Deputy Director for Research
Institute of Plant Genetics of the Polish Academy of Sciences
LikeLike
I am puzzling how Mr Maqbool was ever recruited to the group. It does not require deep scrutiny of this papers to see the microscopy duplications and manipulations, or to recognise that his squiggles presented as FTIR spectra are some kind of weird cargo-cult imitations, Whose idea was it to take him on?
There are unanswered questions about the Frontiers paper (part of a Research Topic edited by Gregory Franklin). How much did Dr Franklin contribute to it, earning him the lead authorship?
Is it possible to comment on the two publications from Dr Franklin without Maqbool’s involvement, that have been flagged at PubPeer for overlapping-image problems?
LikeLike
Response to Małgorzata Jędryczka. Oh Yes its is devastating, In fact it was already devastating from the Dec. 2014 when you did corrupt Bio-Talent requirement procedures for the leader and the ERA-Chair holder. Dr R. Lewandowski should never be recruited as Bio-Talent leader and ERA-Chair professor. His experience in 2014 was worth nothing. If you would have employed me, instead of Lewandowski you did never would get such problems, you did not never would have to cooperate with fake scientists..Since 2014 I have published over 20 primary experimental articles and got 2 patents. My index H increased from 26 to 37. Only true experiments are published. You would never got problems with the final report to the EU. Shame on you!
LikeLike
It was the right decision to retract this manuscript. I look forward the outcome of the institutional investigation.
LikeLike
A rather touching confession by one of the authors involved on PubPeer.
https://pubpeer.com/publications/87AA384477944F61DC1B0967C60303#6
He makes a telling point that behind many of these crap papers is a ruined career by a trainee caught in the web of corrupt supervisors and colleagues on the make.
I hope SWT Maqbool enjoys his brief moment in the sun.
LikeLike
I hope that the first author of the preprint can bounce back without damage to her reputation.
LikeLike
An institutional investigation should determine who was engaged in scientific misconduct, and the other papers should also be investigated of the two authors. Although other authors may have been not involved in data fabrication, they also have a certain degree of responsibility by lacking proper overview of the work/manuscript.
LikeLike
Pingback: Preprinters of the World Unite – For Better Science
He writes his name sometimes “Franklin Gregory”, sometimes “Gregory Franklin”, maybe he has to get used to the name, or is it really his true name?
It is saddening that if the EU supports eastern European countries, they spend the money on attracting third world researchers with doubtful expertise.
At least, “Franklin holds a PhD degree in biotechnology and an LLM in European and Transglobal Business Law with specialisation in IP Rights.” (research-gate self-description), so his focus is not on science anyway.
LikeLike
“Franklin Gregory” versions of the name could easily be someone else’s screw-up.
LikeLike
Consistent with the general perception that “nanoscience” is largely a scam aimed at hoodwinking unwary investors, governments and otherwise. All that BS about green synthesis and cancer drugs. According to his online profile, before he was outed the great P.K. Sharma was expecting to be snapped up by a high-flying nanotech enterprise. Or maybe he was.
LikeLike
Dr R. Malinowski who was selected as the Leader of the Bio-Talent project, won only a few points in the Bio-Talent recruitment in Dec. 2014. At that time he was a fresh postdock, without considerable academic or cooperation with the industry and with no experience of being a group leader, with several publications as a leading author. Yet he was selected by Bio-Talent Recruitment Committee as the best candidate. In the same recruitment my scientific,academic and innovative achievements measured by the rules and scores of the announced Bio-Talent competition were more than 50 times higher than those of Dr. Malinowski (I did scored 260 points almost maximum). Between 2015-2019 I published over 22 experimental papers and made 2 patents. It is this corrupt selection that causes the current problems of IGR-PAN.
LikeLike
Email from. Gregory Franklin
Dear Dr Schneider
As per the request of Prof. Małgorzata Jędryczka, I am replying to your email.
First of all, I sincerely apologise to my institute, Directors, Head of the department and colleagues and all the people to whom this email is copied for this unfortunate situation and inconvenience.
As you could see from the vast amount of fabricated data published by Mr Maqbool https://pubpeer.com/search?q=Qaisar+Maqbool+ most of them published before coming to Poland, I am only a victim here, as I am his PhD supervisor for the past 7 months. I was really impressed with his 2017 single author RSC Advances article and believed all the data produced by him at IPG-PAS. I was not even aware that this kind of image manipulations could be even possible. I apologise for my ignorance.
Regarding the two articles you mentioned:
Article 1
Marslin G, Sarmento B, Franklin G, Martins JAR, Silva CJR, Gomes AFC, Sárria MP, Coutinho O, Dias ACP (Corresponding author)(2017). Curcumin Encapsulated into Methoxy Poly(Ethylene Glycol) Poly(ε-Caprolactone)
Nanoparticles Increases Cellular Uptake and Neuroprotective Effect in Glioma Cells. Planta Medica 83(5): 434-444
DOI: http://dx.doi.org/10.1055/s-0042-112030
I am neither first nor corresponding author in this article, although my affiliation is IPG-PAS (Article 1, attached). I consulted with the corresponding author of this article regarding the question raised by pubpeer on panel C of the Figure 12 showing the cellular uptake of fluorescent nanoparticles after 4, 6 and 24 hours of co-incubation in 24 well animal cell culture plates. I was given to understand that, as the images were taken from the same cell population repeatedly over different period, there might be a possibility that the same group of cells could appear in the images between time points. The authors responsible for this articles will explain to the journal if needed. I have posted my response in your pubpeer blog: https://pubpeer.com/publications/AEB725E285C746261AA27D690A4D9B#2
Article 2
PEGylated ofloxacin nanoparticles render strong antibacterial activity against many clinically important human pathogens” [Colloids and Surfaces B: Biointerfaces 132 (2015) 62–70] Gregory Marslin, Ann Mary Revina, Vinoth Kumar Megraj Khandelwal,Krishnamoorthy Balakumar, Caroline J. Sheeba, Gregory Franklin (Corresponding author)
This work was performed and published from my previous institution in Portugal, /Centre for the Research and Technology of Agro-Environment and Biological Sciences (CITAB). /As we relied on an outside facility (about 50 km away from my work place) for the TEM images and that we overlooked an error in the file names containing the type of nanoparticle and magnification of images, an error occurred with respect to Fig 1B. Although this misrepresentation of image (1B) does not affect the results or conclusions, we have already submitted an erratum. Please understand that we did not have any intention to manipulate anything. As you aware, TEM images of nanoparticles PEGylated or not generally look similar, I could not spot the error before submission. I am also attaching the full article (Article 2). I thank you for spotting ing the error, which provided us an opportunity to correct it. I have also responded to the comment in your pubpeer portal https://pubpeer.com/publications/46148AD4E0BFC61B68D6267E2AB7AD#2
If you need any further information, please feel free to contact me.
Yours sincerely
Franklin
LikeLike
“Dear Franklin,
thank you very much for the explanations concerning your two articles with questioned images. Great care about the quality of documentation of scientific results is of great importance, so I am pleased you have informed the editors of these papers. It is important to know the images have not changed the conclusions from your studies.
Best regards,
Gosia(Malgorzata Jedryczka”
LikeLike
Regarding article 2 (Colloids and Surfaces B: Biointerfaces 132 (2015) 62–70] Gregory Marslin, Ann Mary Revina, Vinoth Kumar Megraj Khandelwal,Krishnamoorthy Balakumar, Caroline J. Sheeba, Gregory Franklin), a reader provided this important information about co-authors:
“Gregory Franklin and Gregory Marslin are brothers. Ann Mary Revina is wife of Gregory Marslin and she never did any research as i know. Caroline J. Sheeba is wife of Gregory Franklin who is from completely different area of research. But in many articles, i can see all of them together!”
LikeLike
It is not the first instance when I see Pakistani authors to use their given names as their family names/surnames. It hides family relationships, such as in this case. It is unlikely that Gregory is a Pakistani family name. I guess they just use two of their given names.
LikeLike
If the case is over, than it is all fine. However, did the Higher Education Pakistan fellowship program agree to terminate the contract and stipend? Or will Mr. Maqbool be moved to another lab, where he can continue his fellowship?
LikeLike
The case is far from over. Looks like Maqbool landed up in exactly the right place for is brand of “science.”
If you want to know what Gregory is up to, consult his Research Gate profile:
“In addition to a PhD, he also holds an LLM degree in European and Transglobal Business Law with specialization in Intellectual Property Rights from the University of Minho. His current research focuses on the application of biotechnology for plant improvement, metabolic engineering in plants to produce pharmaceutically important secondary metabolites, exploration of new drug leads and Nano drug delivery. He has co-ordinated many industrial as well as research projects. He is a scientific entrepreneur strongly motivated towards applying research findings for business development, job creation, and human welfare.”
He is clearly trying to generate as much likely-looking “research” as possible in hopes of roping in potential investors. Let us hope the scam does not go beyond a bit of hustling within what appears to be a pathetic academic system and a highly flawed publication matrix.
LikeLike