France is the place where research integrity truly happens, as a circus clown show. In other countries, fraud is a dirty business dealt in the dark alleys of academia, but in France, it is the prime time entertainment, with all the elites participating, playing victim of persecution, denouncing traitors of the Republic, invoking Nazis and Vichy regime, and issuing legal threats, all on the internet and in national newspapers. Let me sum up so far:
- Olivier Voinnet, star of plant sciences, went from being a fraudulent pariah to a Dreyfus figure, a champion of research integrity who single-handedly uncovered a huge fraud conspiracy committed by his right-hand man Patrice Dunoyer, but the main research findings remain unaffected.
- Anne Peyroche, former CNRS president, was found guilty of having published research fraud and faces a sack by Commissariat à l’énergie atomique.
- Catherine Jessus, until recently chief biologist at CNRS, was defended by all the French elites of Academie de Sciences and CNRS against malicious slander committed by anonymous critics, myself and Le Monde. Turned out, exactly same kind of data manipulations can be fraudulent in Voinnet papers but perfectly scientific in papers by Jessus.
- Francis-Andre Wollmann, the dialectic investigator of both Voinnet and Jessus, Academie de Sciences member and Knight of the Honour Legion, himself coauthored papers with manipulated data.
- Guido Kroemer, another French star scientist and Photoshop artist, used to host in his Paris lab his Spanish cancer research colleague Carlos-Lopez-Otin, a fugitive persecuted by retractions, ridicule and mysterious disappearance of mice. It is not known if Carlos’ friends from Opus Dei came to visit or not, but they sure helped him return back to Oviedo in early March 2019.
- Frédérique Vidal, Minister for Research in the current Macron government, apparently interfered with the misconduct investigations in both affairs of Peyroche and Jessus. It becomes progressively clearer why Vidal tends to side with dishonest scientists: she herself published some manipulated data as university professor. And in this article, I present even more material on this topic, in two new papers authored by Vidal.
The Minister for Research Frederique Vidal is a specialist in molecular genetics of germ cell development. She received her PhD from University of Nice and was appointed assistant professor there in 1995, and full professor in 2004. In 2009, Vidal rose to the rank of Head of Department of Life Sciences, and in 2012, President of the University of Nice. Vidal remained in this position until her appointment as Minister of Higher Education, Research and Innovation in 2017. Like Wollman, she is also Knight of the Honour Legion.
I previously reported a duplicated gel in this Vidal paper:
F. Vidal, P. Lopez, L.A. Lopez-Fernandez, F. Ranc, J.C. Scimeca, F. Cuzin, M. Rassoulzadegan
Gene trap analysis of germ cell signaling to Sertoli cells: NGF-TrkA mediated induction of Fra1 and Fos by post-meiotic germ cells
Journal of Cell Science 2001 Jan;114(Pt 2):435-43.
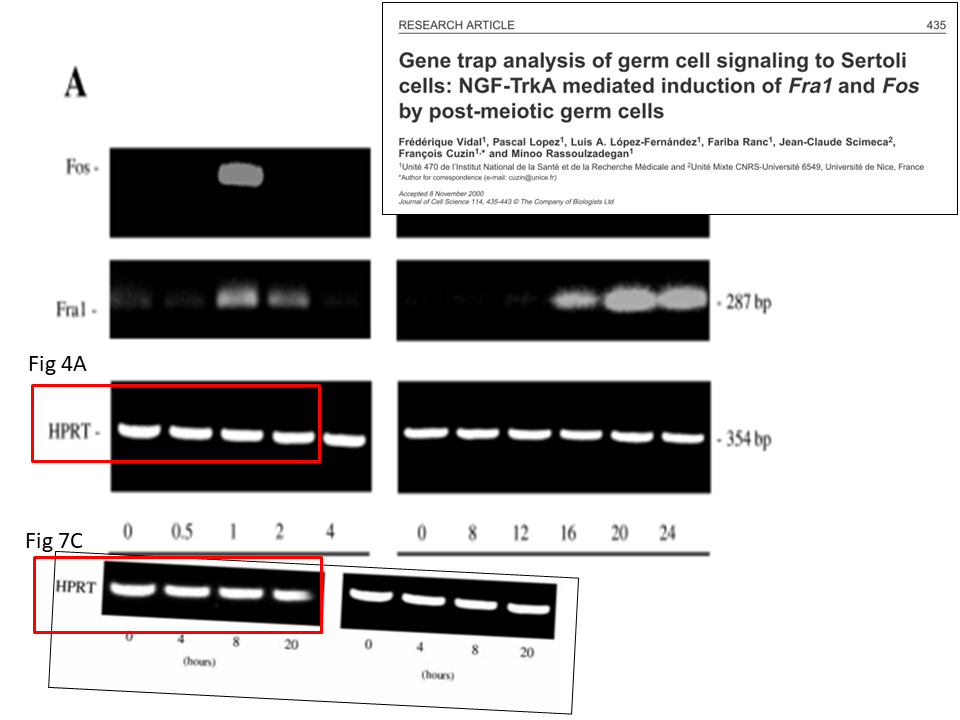
A certain expert then provided additional evidence on PubPeer, it turned out the gel featured in the same paper not twice, but trice. The left gel in Figure 7C is namely same as the right one, just as in Figure 4A, only artistically blurred and rotated:
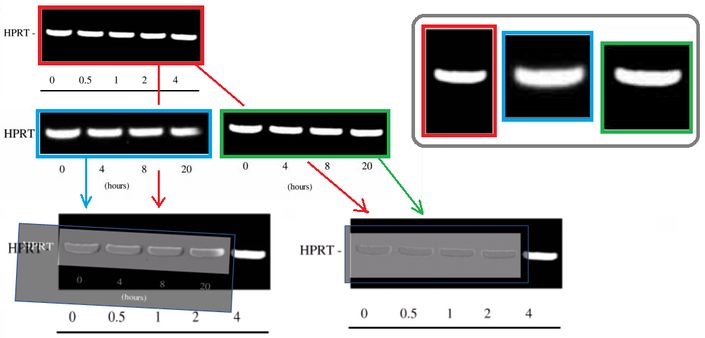
It also was pointed out that the fifth gel band in Figure 4A looks suspiciously too similar to the first band in the same gel (and in its two other incarnations). Hoya camphorifolia explained:
“the PCR results for HPRT in Fig 7C (left-hand panel) are more similar than one would expect to the first four lanes of the results in Fig 4A, slightly defocussed and rotated through 2deg. See, for instance, the enlargements of the first lane(s), at the upper right.
It is certainly true that the respective lanes (and the spacings between them) allow the two panels to overlap and cancel out, when one is black-white reversed and superimposed on the other (at lower left).
But this should not blind us to the observation that the HPRT band from Fig 4A also closely resembles the right-hand band from 7C, without blurring or rotation, as shown in the superimposition at lower right.”
The Ministry of of Higher Education, Research and Innovation reacted swiftly and mercilessly. Same month my article appeared, in October 2018, it issued this statement to the French higher education news service News Tank:
“Charges have been laid against a scientific publication published 17 years ago and co-authored by seven authors, including the current Minister of Higher Education, research and innovation. These unacceptable accusations are totally unfounded. The goal sought by their author is clear: to destabilize the minister. Faced with these accusations, the Minister is studying the possibility to institute legal proceedings.”
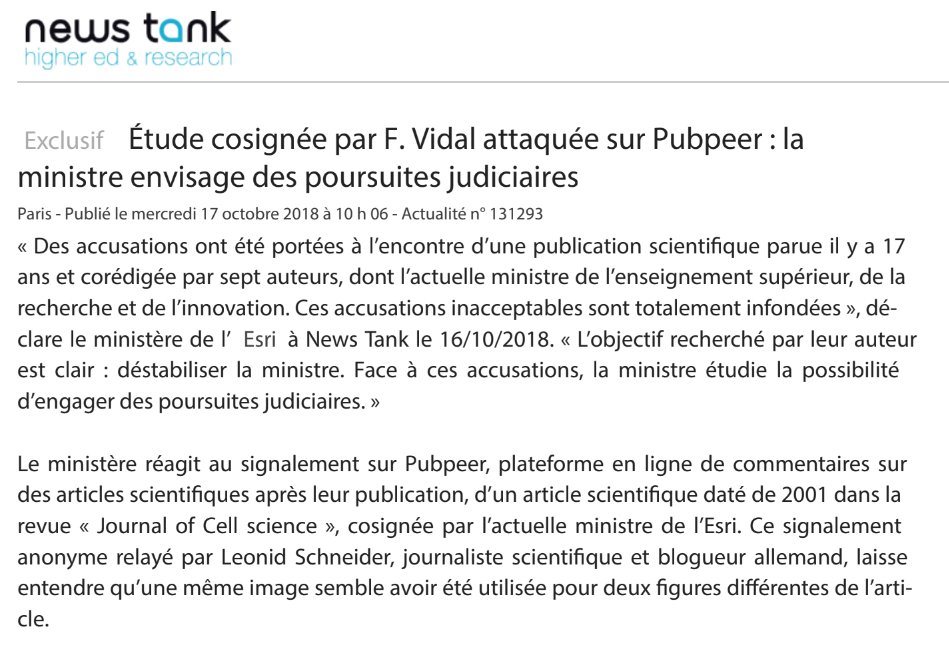
I sent my two-fingered salute back to the Government of France, yet nothing happened since. But now a letter arrived in my mailbox. Not an email, a real letter, in an envelope from France. The unsigned missive presented evidence of more duplicated gel bands in Vidal papers. These being:
Pascal Lopez, Frédérique Vidal, Luc Martin, Luis A. Lopez-Fernandez, Jean-François Rual, Barry S. Rosen, François Cuzin, Minoo Rassoulzadegan
Gene Control in Germinal Differentiation: Rnf6, a Transcription Regulatory Protein in the Mouse Sertoli Cell
Molecular and Cellular Biology (2002) DOI: 10.1128/MCB.22.10.3488-3496.2002
Received November 19, 2001; Returned for modification January 7, 2002; Accepted January 25, 2002; Published online May 15, 2002.
and
Pascal Lopez, Ruken Yaman, Luis A. Lopez-Fernandez, Frédérique Vidal, Daniel Puel, Philippe Clertant, François Cuzin and Minoo Rassoulzadegan
A Novel Germ Line-specific Gene of the Phosducin-like Protein (PhLP) Family
Journal of Biological Chemistry (2002) doi: 10.1074/jbc.M207434200
Received for publication, July 24, 2002, and in revised form, October 28, 2002. Published, November 6, 2002
This is the problem, in overview, the duplicated gel bands are highlighted in colour:
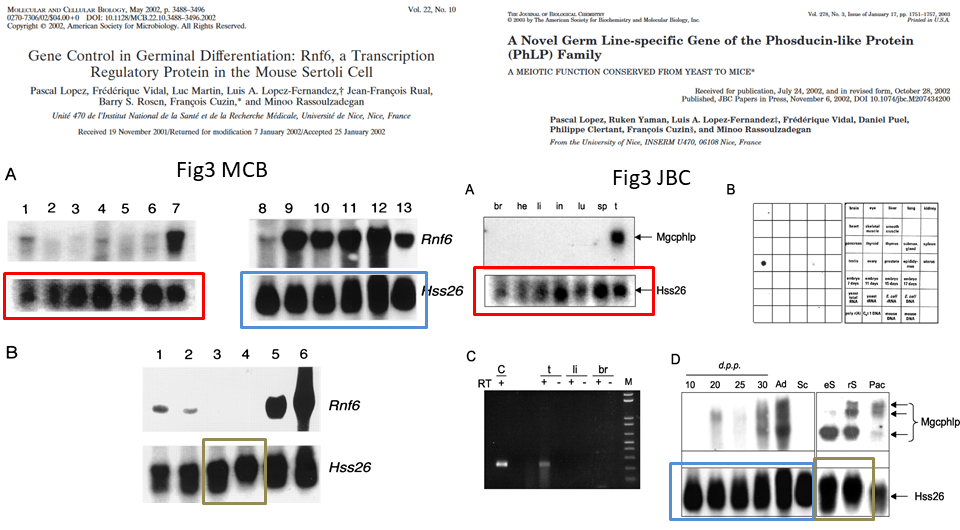
Northern Blot is the method to analyse RNA expression. RNA extracts from cells or tissues are run on a gel and transferred to a blot membrane, which is then incubated with a radioactively labelled specific probe, in this case to detect individual messenger RNAs (mRNA). The images in question show the loading control for the Hss26 mRNA, and these are obviously duplicated between the two papers. However, for the red and blue framed cases, one must say the samples are the same. The MCB paper provides this legend, and it matches the legend in the JBC paper:
“Northern blot analysis of total RNA prepared from brain (lane1), heart (lane 2), liver (lane 3), intestine (lane 4), lung (lane 5), spleen (lane 6), adult testis (lane 7), testis of a 10-day-old mouse (lane 8), testis of a 20-day-old mouse (lane 9), testis of a 25-day-old mouse (lane 10), testis of a 30-day-old mouse (lane 11), testis of a 2-month-old mouse (lane 12), and freshly isolated Sertoli cells from 3-week-old mice (lane 13)”
In both cases, the authors declare respectively: “Hybridization was performed in succession with probes for cDNAs of the Rnf6 and S26 proteins” and “Hybridization was performed in succession with a complete Mgcphlp cDNA probe and with a cDNA probe for the ubiquitous ribosomal S26 protein“. This means that the images for Rnf6 and corresponding Hss26 mRNA signals in MCB come from the same Northern blot membrane, and same is declared for the Mgcphlp and Hss26 signals in JBC.
The authors might be tempted to explain that it was the same Northern blot membrane in both papers, stripped with chemical solutions to remove the previously bound radioactive probe; it would resemble the defence which Lopez-Otin used for his Perennial Northern Blot. Only that is a rather weird scientific approach. Both papers report namely novel studies of Rnf6 and Mgcphlp mRNA expression, in these cases scientists generally prepare new gels and do not reuse old blot membranes, simply because they will not know which signal is correct and which might be a residue from the previously applied probe. They do not know in advance how strong the signal from a new probe would be, while the stripping process flushes out not just the old probe, but also some of the blotted RNA material also. Basically, stripping is something one does not do for new explorative analyses, and this is likely why a blot membrane reuse was not declared in the two papers.
But why the same loading control then? Maybe the authors decided not to probe for loading control again when they repeated the gel for the Mgcplp study, using same samples in same order. That is not really a good practice, because we can’t know how well the samples preserved, entered the gel and transferred to membrane in this repeated experiment.
Another issue highlighted above is the gel lane splicing between lanes 12 and 13 in MCB Figure 3A and corresponding lanes in Figure 3D in JBC paper. It is important because the authors did highlight the splicing one lane further in same figure 3D in JBC, with a clear vertical dividing line. Here it is in close-up:
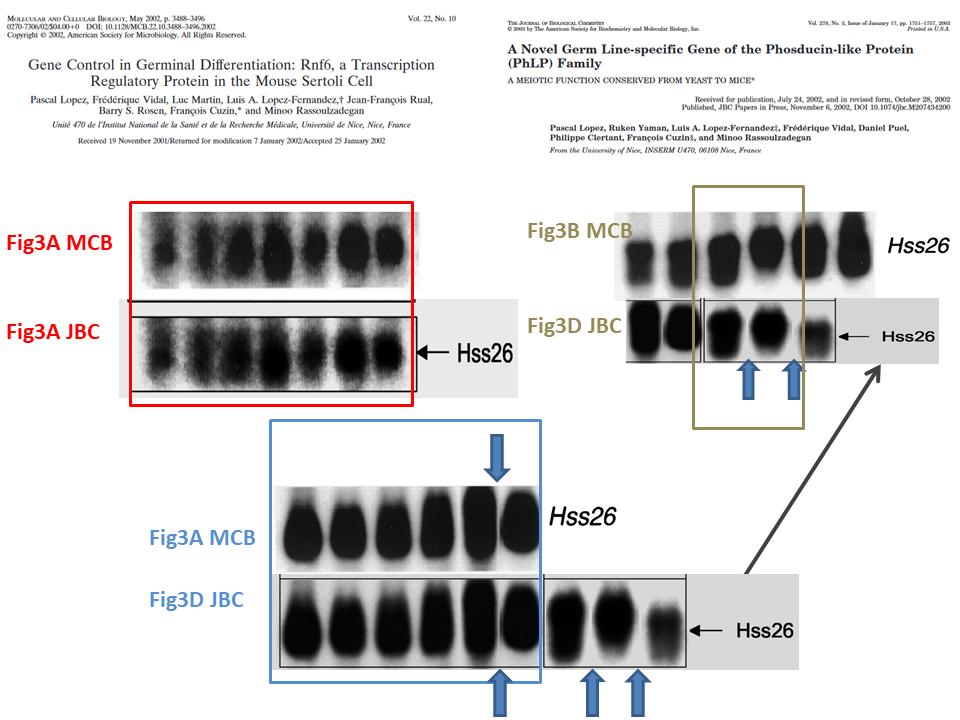
The authors knew splicing of gels must be clearly indicated, and did so in that one instance. But in other instances, they chose to hide gel splicing from peer reviewers and readers, which I indicate above with the blue arrows. And this is where we arrive to a case of intentional data manipulation, highlighted in green.
The two penultimate bands in Hss26 gel in Figure 3D JBC are spliced in, and one sees where they originally come from: the Hss26 gel in MCB paper. But the samples are not same, according to the figure legend. The original MCB legend goes: “Northern blot analysis of total RNA prepared from germinal fractions purified (≥90% pure) by elutriation centrifugation. Fractions: […] 3 and 4, round spermatids“. Compare to the legend of JBC paper for these cloned bands: “eS, rS […], 80–90% pure elutriated preparations of, respectively, elongated spermatids, round spermatids“. Hence, different preparation methodology and different samples. The reuse cannot be explained by mistake, what is presented in Figure 3D JBC as continuous gel is actually a collage of 3 different gel bands which originally showed something else than described.

The shoddy attitude to indicating gel splicing only where one feels safe to do so, is visible again in Figure 7B of the MCB paper:
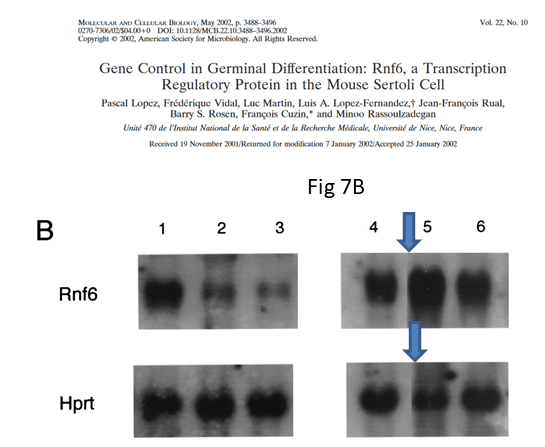
The authors declared “The same membrane was hybridized in succession with probes for the Rnf6 (top) and Hprt (bottom) mRNAs” but now we cannot be sure about that. When scientists start passing off old data as new and even stealthily mixing it into a collage to generate new results, the trust is gone. It might be not a minor issue that one of the authors is a government minister for research, whose ministry’s only reaction so far was a threat of legal action against those who reported the evidence.
Now we know why you protect Peyroche and Jessus so much, Professor Vidal.

Update 25.05.2019. On 15 May an Expression of Concern for the paper Vidal et al 2001 was published:
“Journal of Cell Science was alerted to duplicated HPRT blots in Fig. 4A and Fig. 7C of this paper. The authors state that the conclusions of the paper are not affected by the duplicated control blots, but were unable to locate the original data from almost 20 years ago. Without the original full blots, the journal is unable to determine whether the results and conclusions reported in the paper are compromised.
The journal is publishing this Expression of Concern to make readers aware of these issues. The authors offer cell lines used in the paper for replication by any interested investigators and apologise to readers for any inconvenience caused.”
Another article in News Tank followed on 24.05.2019.
We learn that it was Vidal responsible for the original lab book which now cannot be found. There seems to be a CD-ROM around, but it is not readable with modern technology, according to last author Minoo Rassoulzadegan. But now comes the kicker, in News Tank, the target being again yours truly:
“Frédérique Vidal “always reserves the right” to institute legal proceedings. […] “Frédérique Vidal, as a minister, has not deemed it necessary to prosecute at this time. However, she still reserves the right to do so, should personal accusations be brought against her”
Crazy clowns. Anyway, what is this, from Université de Nice Sophia Antipolis, featuring err… or dear… does this count as a “personal accusation”?
Isabelle Gillot, Cédric Matthews, Daniel Puel, Frédérique Vidal, and Pascal Lopez
Ret Finger Protein: An E3 Ubiquitin Ligase Juxtaposed to the XY Body in Meiosis
Int J Cell Biol. (2009) doi: 10.1155/2009/524858
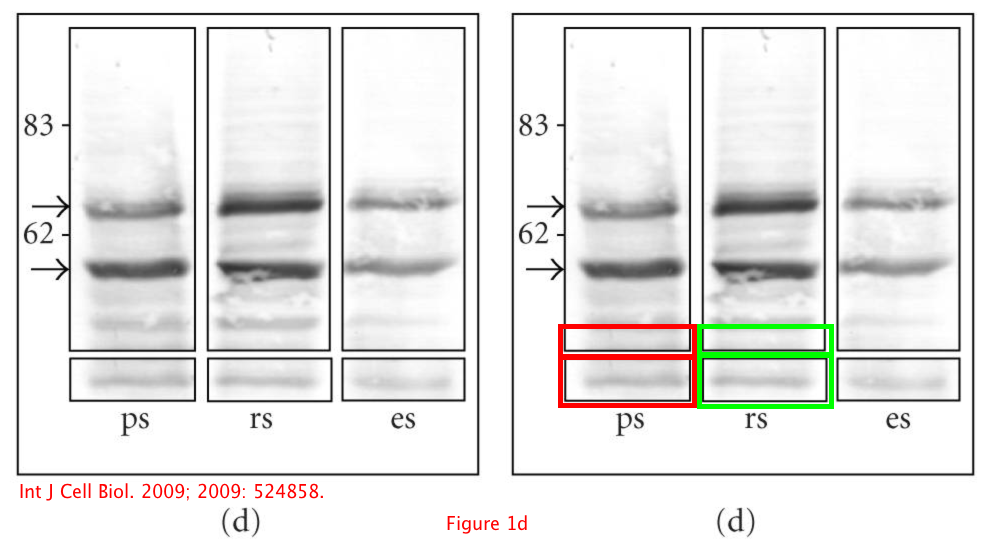

Update 5.06.2019.
In today’s News Tank article (Actualité n° 148672), the senior author Minoo Rassoulzadegan explained everything. Here some translated quotes from the news piece, and please also note her reply on yet another Photoshop-train-wreck figure below.
“Leonid Schneider asserts that the authors “chose to hide the splicing” between bands of gel with results of their research in some figures of these articles. According to Minoo Rassoulzadegan, this selection of information is “necessary” for understanding the results. “Car radios are easy to handle, but not biological results,” she says.“

I was called many things, but never a car radio mechanic. Regarding the reused loading control, Rassoulzadegan explains:
“According to Minoo Rassoulzadegan, “indeed it is the same filter (nitrocellulose membrane) that has been used several times to hybridize with different probes”. “The two candidate genes came out of so-called “gene-trap” experiments. We have identified more than 400 clones and then mass characterized them, carried out gene expression analyses in different tissues, and for this the same filters have been used several times and, I repeat, with different probes.
It is quite normal that the loading controls are identical for both experiments. It is routine to use the same nitrocellulose membrane loaded with different tissues for different analyses, the same filter can be used more than ten times. Research is expensive in terms of time and resources and researchers must avoid waste.“
She doesn’t go into why loading controls were reused between different samples. But then again, Professor Rassoulzadegan is on a roll. Guess how she explained this:
Lopez P, Yaman R, Lopez-Fernandez LA, Vidal F, Puel D, Clertant P, Cuzin F, Rassoulzadegan M.
A novel germ line-specific gene of the phosducin-like protein (PhLP) family. A meiotic function conserved from yeast to mice.
J Biol Chem. 2003 DOI:10.1074/jbc.M207434200
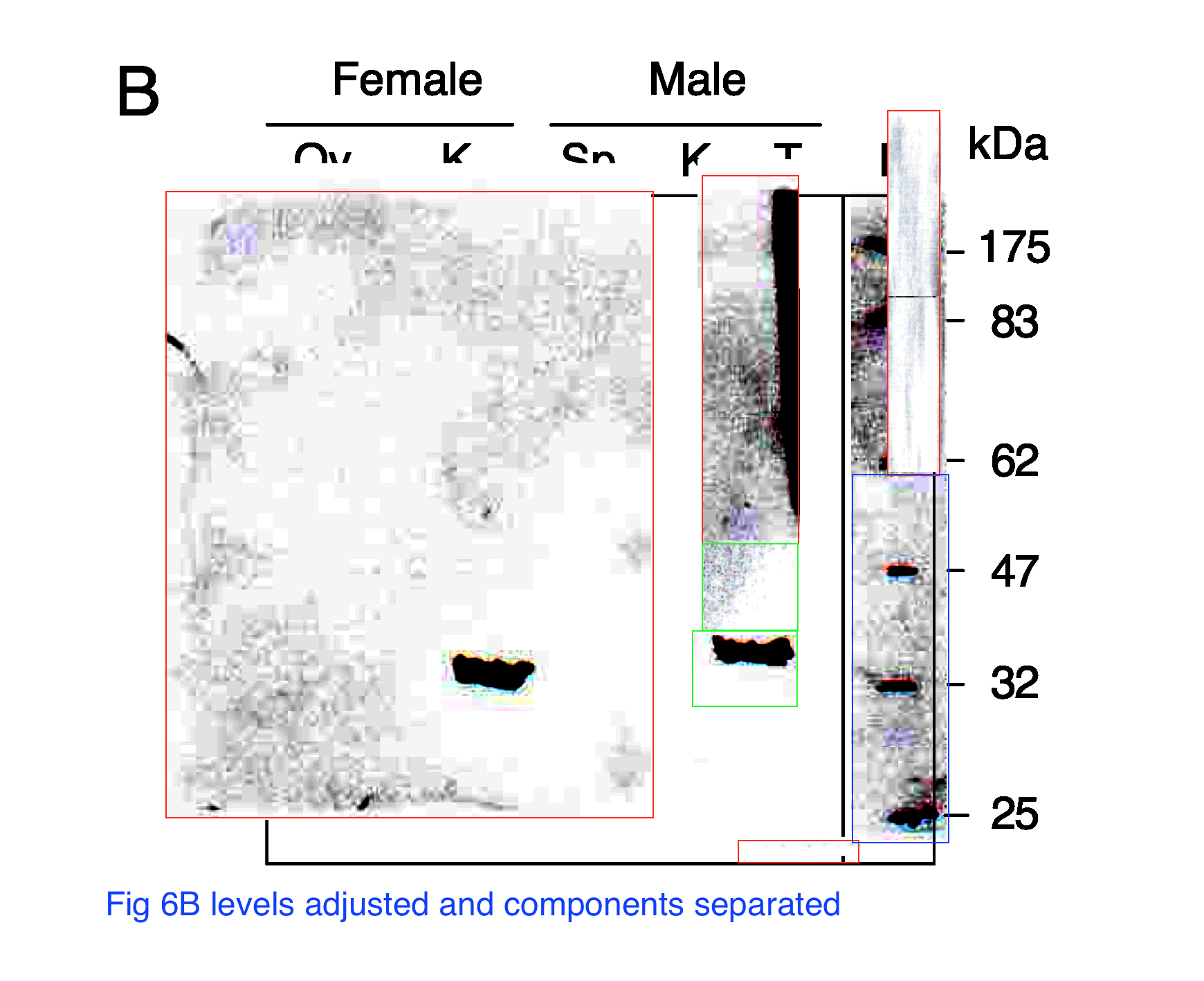
“According to Minoo Rassoulzadegan, “it is clear that in the lane T (testicles), there is much more signal than for the samples in the other lanes and it was necessary to adjust the signal, otherwise we would have seen nothing for example in the Sp lane (sperm) or somatic lanes like K (kidney) “.
Jessus wept.

Donate!
If you are interested to support my work, you can leave here a small tip of $5. Or several of small tips, just increase the amount as you like (2x=€10; 5x=€25). Your generous patronage of my journalism will be most appreciated!
€5.00


Infect Immun. 2003 Feb;71(2):766-73.
Saccharomyces boulardii interferes with enterohemorrhagic Escherichia coli-induced signaling pathways in T84 cells.
Dahan S1, Dalmasso G, Imbert V, Peyron JF, Rampal P, Czerucka D.
Author information
Laboratoire de Gastroentérologie et Nutrition, Faculté de Médecine, Université de Nice-Sophia Antipolis, 06107 Nice Cedex 2, France.
https://pubpeer.com/publications/A360C60C3423A9369A6D6F606CF35C
PLoS One. 2010 Jan 27;5(1):e8925. doi: 10.1371/journal.pone.0008925.
Interaction of Saccharomyces boulardii with Salmonella enterica serovar Typhimurium protects mice and modifies T84 cell response to the infection.
Martins FS1, Dalmasso G, Arantes RM, Doye A, Lemichez E, Lagadec P, Imbert V, Peyron JF, Rampal P, Nicoli JR, Czerucka D.
Author information
Team 4: Inflammation, Cancer, Cancer Stem Cells, Unité INSERM U895, Centre Méditerranéen de Médecine Moléculaire, Nice, France.
https://pubpeer.com/publications/5A014717F973615CC56818FE6ABC48
Br J Cancer. 2008 Jan 29;98(2):335-44. doi: 10.1038/sj.bjc.6604082. Epub 2008 Jan 8.
Pharmacological targeting of NF-kappaB potentiates the effect of the topoisomerase inhibitor CPT-11 on colon cancer cells.
Lagadec P1, Griessinger E, Nawrot MP, Fenouille N, Colosetti P, Imbert V, Mari M, Hofman P, Czerucka D, Rousseau D, Berard E, Dreano M, Peyron JF.
Author information
INSERM U526, Nice F-06000, France.
https://pubpeer.com/publications/9E644D3D952DE9050E835CD018CD98
J Biol Chem. 2012 Jul 13;287(29):24573-84. doi: 10.1074/jbc.M111.333054. Epub 2012 May 31.
Nuclear factor-κB regulates βAPP and β- and γ-secretases differently at physiological and supraphysiological Aβ concentrations.
Chami L1, Buggia-Prévot V, Duplan E, Del Prete D, Chami M, Peyron JF, Checler F.
Author information
Équipe Labellisée “Fondation pour la Recherche Médicale” and “Excellence Laboratory Distalz”, Institut de Pharmacologie Moléculaire et Cellulaire, UMR7275 CNRS/UNSA, 660 Route des Lucioles, 06560, Valbonne, France.
https://pubpeer.com/publications/2073A5EE98B9C529AF152F51214D5D
LikeLike
J Biol Chem. 2001 Nov 30;276(48):45307-19. Epub 2001 Sep 4.
Glial cell line-derived neurotrophic factor-stimulated phosphatidylinositol 3-kinase and Akt activities exert opposing effects on the ERK pathway: importance for the rescue of neuroectodermic cells.
Mograbi B1, Bocciardi R, Bourget I, Busca R, Rochet N, Farahi-Far D, Juhel T, Rossi B.
Author information
INSERM U 364, IFR50, Faculté de Médecine Pasteur, 06107 Nice Cedex 02, France.
https://pubpeer.com/publications/20B134B8DEE1A010CD7AAD1EA18FF2
Infect Immun. 2003 Mar;71(3):1161-9.
Rho GTPase is activated by cytotoxic necrotizing factor 1 in peripheral blood T lymphocytes: potential cytotoxicity for intestinal epithelial cells.
Brest P1, Mograbi B, Hofman V, Loubat A, Rossi B, Auberger P, Hofman P.
Author information
INSERM 364, Faculté de Médecine, 06107 Nice Cédex 02, France.
https://pubpeer.com/publications/61CD52D28573392FAB33DBB6D9EE57
https://www.nature.com/articles/1201413
The multiple endocrine neoplasia type 2B point mutation switches the specificity of the Ret tyrosine kinase towards cellular substrates that are susceptible to interact with Crk and Nck
Renata Bocciardi, Baharia Mograbi, Barbara Pasini, Maria Grazia Borrello, Marco A Pierotti, Isabelle Bourget, Siegmund Fischer, Giovanni Romeo & Bernard Rossi
Oncogene volume 15, pages 2257–2265 (06 November 1997)
Author information
Author notes
Renata Bocciardi & Baharia Mograbi
R Bocciardi and B Mograbi contributed equally to this work
Affiliations
Laboratorio di Genetica Molecolare, Istituto Giannina Gaslini, Genova, Italy
Renata Bocciardi, Baharia Mograbi & Giovanni Romeo
Direzione Scientifica, Istituto Nazionale Tumori, Milano, Italy
Barbara Pasini
Divisione di Oncologia Sperimentale A, Istituto Nazionale Tumori, Milano, Italy
Maria Grazia Borrello & Marco A Pierotti
INSERM U 364, Faculté de Médecine Pasteur, Nice, France
Isabelle Bourget & Bernard Rossi
Institut Cochin de Génétique Moléculaire, INSERM U363, Hôpital Cochin, Paris, France
Siegmund Fischer
International Agency for Research on Cancer, Lyon, France
Giovanni Romeo
Corresponding author
Correspondence to Bernard Rossi.
https://pubpeer.com/publications/FCDBE2B9009A5A9A16BC49F89F7273
LikeLike
J Bone Miner Res. 2003 Oct;18(10):1863-71.
Focal adhesion kinase pp125FAK interacts with the large conductance calcium-activated hSlo potassium channel in human osteoblasts: potential role in mechanotransduction.
Rezzonico R1, Cayatte C, Bourget-Ponzio I, Romey G, Belhacene N, Loubat A, Rocchi S, Van Obberghen E, Girault JA, Rossi B, Schmid-Antomarchi H.
Author information
Faculté de Médecine, INSERM U364, Nice, France.
https://pubpeer.com/publications/4784BC653A3EF163A168A86D9E5DE9
Blood. 2001 Apr 1;97(7):2031-7.
Signal transduction pathways involved in soluble fractalkine-induced monocytic cell adhesion.
Cambien B1, Pomeranz M, Schmid-Antomarchi H, Millet MA, Breittmayer V, Rossi B, Schmid-Alliana A.
Author information
INSERM U364, Facultè de Mèdecine, Nice Cedex 02, France.
https://pubpeer.com/publications/8062A0F364DA11DC293A00C07750C9
Int J Cancer. 2009 Dec 1;125(11):2586-94. doi: 10.1002/ijc.24665.
1 comment on PubPeer (by: Richeria Grandis)
Antagonism of chemokine receptor CXCR3 inhibits osteosarcoma metastasis to lungs.
Pradelli E1, Karimdjee-Soilihi B, Michiels JF, Ricci JE, Millet MA, Vandenbos F, Sullivan TJ, Collins TL, Johnson MG, Medina JC, Kleinerman ES, Schmid-Alliana A, Schmid-Antomarchi H.
Author information
Université de Nice Sophia-Antipolis, Nice, France.
https://pubpeer.com/publications/FC5CAE54259736B9DE00EBDEF253EA
J Immunol. 1999 Nov 1;163(9):5079-85.
Src-regulated extracellular signal-related kinase and Syk-regulated c-Jun N-terminal kinase pathways act in conjunction to induce IL-1 synthesis in response to microtubule disruption in HL60 cells.
Cambien B1, Millet MA, Schmid-Antomarchi H, Brossette N, Rossi B, Schmid-Alliana A.
Author information
Institut National de la Santé et de la Recherche Scientifique Unite 364, Nice, France.
https://pubpeer.com/publications/BD664F156E38F7E530ECC51C454511
Blood. 2001 Jan 15;97(2):359-66.
Signal transduction involved in MCP-1-mediated monocytic transendothelial migration.
Cambien B1, Pomeranz M, Millet MA, Rossi B, Schmid-Alliana A.
Author information
INSERM U364, Faculté de Médecine, Nice, France.
https://pubpeer.com/publications/9CA15D90C6CC021393FB7265E9E71A
LikeLike
Mol Cell Neurosci. 2011 Jul;47(3):223-32. doi: 10.1016/j.mcn.2011.04.008. Epub 2011 May 4.
2 comments on PubPeer (by: Peer 1, Breinlia Mundayi)
ERK1-independent α-secretase cut of β-amyloid precursor protein via M1 muscarinic receptors and PKCα/ε.
Cisse M1, Braun U, Leitges M, Fisher A, Pages G, Checler F, Vincent B.
Author information
Université de Nice-Sophia-Antipolis, Institut de Neuro-Médecine Moléculaire, Unité Mixte de Recherche, 6097 Centre National de la Recherche Scientifique, Equipe labellisée Fondation pour la Recherche Médicale, 660 route des lucioles, Sophia-Antipolis, 06560 Valbonne, France.
https://pubpeer.com/publications/A2BEF190C65401D89A364DE5CDC354
J Biol Chem. 2002 Dec 27;277(52):50980-4. Epub 2002 Oct 22.
Alpha-synuclein lowers p53-dependent apoptotic response of neuronal cells. Abolishment by 6-hydroxydopamine and implication for Parkinson’s disease.
Alves Da Costa C1, Paitel E, Vincent B, Checler F.
Author information
Institut de Pharmacologie Moléculaire et Cellulaire du CNRS, UMR6097, 660 route des Lucioles, 06560 Valbonne, France.
Neuron
Volume 46, Issue 4, 19 May 2005, Pages 541-554
https://www.sciencedirect.com/science/article/pii/S0896627305003211?via%3Dihub
Presenilin-Dependent Transcriptional Control of the Aβ-Degrading Enzyme Neprilysin by Intracellular Domains of βAPP and APLP
RaphaëllePardossi-Piquard1AgnèsPetit1ToshitakaKawarai2ClaireSunyach1CristineAlves da Costa1BrunoVincent1SabineRing3LucianoD’Adamio45JieShen6UlrikeMüller3Peter St. GeorgeHyslop2FrédéricChecler1
1 Institut de Pharmacologie Moléculaire et Cellulaire, Centre National de la Recherche Scientifique, UMR6097 CNRS/UNSA, Valbonne 06560, France
2 Centre for Research in Neurodegenerative Diseases, Department of Medicine, University of Toronto and University Health Network, Toronto Western Hospital Research Institute, 6 Queen’s Park Crescent, Toronto, Ontario M5S 3H2, Canada
3 Institute for Pharmacy and Molecular Biotechnology, University of Heidelberg, 69120 Heidelberg, Germany
4 Albert Einstein College of Medicine, New York, New York
5 Dipartimento di Biochimica e Biotecnologie Mediche, Universita’ Degli Studi di Napoli Federico II, Napoli, Italy
6 Center for Neurologic Diseases, Harvard Medical School, Boston, Massachusetts.
https://pubpeer.com/publications/BE67D5F472B11B75A0F933ECE906DD
J Biol Chem. 2007 Apr 6;282(14):10516-25. Epub 2007 Feb 2.
p53-Dependent Aph-1 and Pen-2 anti-apoptotic phenotype requires the integrity of the gamma-secretase complex but is independent of its activity.
Dunys J1, Kawarai T, Sevalle J, Dolcini V, George-Hyslop PS, Da Costa CA, Checler F.
Author information
Institut de Pharmacologie Moléculaire et Cellulaire, UMR 6097 CNRS/UNSA, Equipe labellisée, Fondation pour la Recherche Médicale, 660 Route des Lucioles, 06560 Valbonne, France.
https://pubpeer.com/publications/3EE4379793F882C57F34966C37079F
Biochem J. 2006 Mar 1;394(Pt 2):501-9.
Catabolism of endogenous and overexpressed APH1a and PEN2: evidence for artifactual involvement of the proteasome in the degradation of overexpressed proteins.
Dunys J1, Kawarai T, Wilk S, St George-Hyslop P, Alves da Costa C, Checler F.
Author information
Institut de Pharmacologie Moléculaire et Cellulaire, Centre National de la Recherche Scientifique, Valbonne, France.
https://pubpeer.com/publications/A5B0107B89615AB8DEDE8B10501463
Neurodegener Dis. 2007;4(2-3):156-63.
Study on the putative contribution of caspases and the proteasome to the degradation of Aph-1a and Pen-2.
Dunys J1, Kawarai T, Giaime E, Wilk S, Herrant M, Auberger P, St George-Hyslop P, Alves da Costa C, Checler F.
Author information
Institut de Pharmacologie Moléculaire et Cellulaire, UMR6097 CNRS/UNSA, Equipe labellisée Fondation pour la Recherche Médicale, Valbonne, France.
https://pubpeer.com/publications/2103522F81154B997B4236F50B981A
J Neurosci. 2009 May 20;29(20):6752-60. doi: 10.1523/JNEUROSCI.0789-09.2009.
p53-Dependent transcriptional control of cellular prion by presenilins.
Vincent B1, Sunyach C, Orzechowski HD, St George-Hyslop P, Checler F.
Author information
Institut de Pharmacologie Moléculaire et Cellulaire, Unité Mixte de Recherche 6097 Centre National de la Recherche Scientifique/Université de Nice-Sophia-Antipolis, 06560 Valbonne, France.
https://pubpeer.com/publications/DCB79632E2579603F640F85D6C6A63#2
https://pubpeer.com/publications/DCB79632E2579603F640F85D6C6A63#3
LikeLike
J Neurochem. 2002 Dec;83(5):1208-14.
Overexpression of PrPc triggers caspase 3 activation: potentiation by proteasome inhibitors and blockade by anti-PrP antibodies.
Paitel E1, Alves da Costa C, Vilette D, Grassi J, Checler F.
Author information
IPMC of CNRS, UMR6097, Valbonne, France INRA, Jouy en Josas, France CEA/Saclay, Gif sur Yvette, France.
https://pubpeer.com/publications/5572903803C87A2C0474AFBF8C2E6E
J Biol Chem. 2011 Aug 19;286(33):29192-206. doi: 10.1074/jbc.M110.208249. Epub 2011 May 17.
The extracellular regulated kinase-1 (ERK1) controls regulated alpha-secretase-mediated processing, promoter transactivation, and mRNA levels of the cellular prion protein.
Cissé M1, Duplan E, Guillot-Sestier MV, Rumigny J, Bauer C, Pagès G, Orzechowski HD, Slack BE, Checler F, Vincent B.
Author information
Institut de Pharmacologie Moléculaire et Cellulaire, Unité Mixte de Recherche, 6097 Centre National de la Recherche Scientifique/Université de Nice-Sophia-Antipolis, Equipe labellisée Fondation pour la Recherche Médicale, 660 route des lucioles, Sophia-Antipolis, 06560 Valbonne, France.
https://pubpeer.com/publications/3186D1114958BCC45251E9DD5847B3
J Biol Chem. 2001 Oct 12;276(41):37743-6. Epub 2001 Jul 26.
The disintegrins ADAM10 and TACE contribute to the constitutive and phorbol ester-regulated normal cleavage of the cellular prion protein.
Vincent B1, Paitel E, Saftig P, Frobert Y, Hartmann D, De Strooper B, Grassi J, Lopez-Perez E, Checler F.
Author information
Institut de Pharmacologie Moléculaire et Cellulaire du CNRS, UMR6097, 06560 Valbonne, France.
https://pubpeer.com/publications/6CCD05FB812362966C2F919527D143
Biochem Biophys Res Commun. 1998 Nov 9;252(1):134-8.
Alzheimer’s disease-linked mutation of presenilin 2 (N141I-PS2) drastically lowers APPalpha secretion: control by the proteasome.
Marambaud P1, Alves da Costa C, Ancolio K, Checler F.
Author information
Institut de Pharmacologie Moléculaire et Cellulaire, du CNRS, UPR411, 660 Route des Lucioles, Sophia Antipolis, Valbonne, 06560, France.
https://pubpeer.com/publications/7B5C3A6187DE0C3A2EFA7905842A24
J Biol Chem. 2005 Dec 9;280(49):40624-31. Epub 2005 Oct 18.
The disintegrin ADAM9 indirectly contributes to the physiological processing of cellular prion by modulating ADAM10 activity.
Cissé MA1, Sunyach C, Lefranc-Jullien S, Postina R, Vincent B, Checler F.
Author information
Institut de Pharmacologie Moléculaire et Cellulaire, du CNRS, UMR6097, Sophia-Antipolis, 06560 Valbonne, France.
https://pubpeer.com/publications/4D482ED31DD121DCF4A0B50D3051F5
J Biol Chem. 2003 Sep 26;278(39):37330-5. Epub 2003 Jul 16.
Beta-synuclein displays an antiapoptotic p53-dependent phenotype and protects neurons from 6-hydroxydopamine-induced caspase 3 activation: cross-talk with alpha-synuclein and implication for Parkinson’s disease.
da Costa CA1, Masliah E, Checler F.
Author information
Institut de Pharmacologie Moléculaire et Cellulaire of Centre National de la Recherche Scientifique, UMR6097, 06560 Valbonne, France.
https://pubpeer.com/publications/E2869E4BEE8D1A43FC1B2BBDE4C203
J Neurochem. 2001 Mar;76(5):1532-9.
Constitutive alpha-secretase cleavage of the beta-amyloid precursor protein in the furin-deficient LoVo cell line: involvement of the pro-hormone convertase 7 and the disintegrin metalloprotease ADAM10.
Lopez-Perez E1, Zhang Y, Frank SJ, Creemers J, Seidah N, Checler F.
Author information
IPMC du CNRS, UMR6097, Valbonne, France.
https://pubpeer.com/publications/3FE16011A63F723E5FD36EEC51186D
LikeLike
J Biol Chem. 2006 Apr 7;281(14):9824-31. Epub 2006 Feb 7.
6-Hydroxydopamine but not 1-methyl-4-phenylpyridinium abolishes alpha-synuclein anti-apoptotic phenotype by inhibiting its proteasomal degradation and by promoting its aggregation.
Alves da Costa C1, Dunys J, Brau F, Wilk S, Cappai R, Checler F.
Author information
1 IPMC du CNRS, UMR6097, Equipe Labellisée FRM, 660 Route des Lucioles, 06560 Valbonne, France.
2013 correction figures 1 and 9.
http://www.jbc.org/content/288/29/21208.short
http://unice.fr/fil/service-communication/actualites/le-dr-frederic-checler-ipmc-recoit-le-prix-2014-de-la-fondation-claude-pompidou-1
J Cell Sci. 2009 Nov 1;122(Pt 21):4003-8. doi: 10.1242/jcs.051169.
3 comments on PubPeer (by: Unregistered Submission, Peer 2)
p53-dependent control of transactivation of the Pen2 promoter by presenilins.
Dunys J1, Sevalle J, Giaime E, Pardossi-Piquard R, Vitek MP, Renbaum P, Levy-Lahad E, Zhang YW, Xu H, Checler F, da Costa CA.
Author information
1
Institut de Pharmacologie Moléculaire et Cellulaire of Centre National de la Recherche Scientifique and Institut de NeuroMédecine Moléculaire, Equipe labellisée Fondation pour la Recherche Médicale, Valbonne, France.
https://pubpeer.com/publications/5EDA66A8634AEC66FBD7651C49CE58#2
https://pubpeer.com/publications/5EDA66A8634AEC66FBD7651C49CE58#3
Biochemical and Biophysical Research Communications
Volume 371, Issue 1, 20 June 2008, Pages 69-74
https://www.sciencedirect.com/science/article/pii/S0006291X08006438?via%3Dihub
TMP21 regulates Aβ production but does not affect caspase-3, p53, and neprilysin
Author links open overlay panelVirginiaDolciniaJulieDunysaJeanSevalleaFushengChenbcMarie-VictoireGuillot-SestieraPeterSt. George-HyslopbcPaul E.FraserbcFrédéricChecleraa
Institut de Pharmacologie Moléculaire et Cellulaire, UMR 6097, Centre National de la Recherche Scientifique-Université Nice-Sophia-Antipolis, Equipe labellisée Fondation pour la Recherche Médicale, 660 Route des Lucioles, 06560 Valbonne, Franceb
Center for Research in Neurodegenerative Diseases, Department of Medicine, University of Toronto, Toronto Western Hospital Research Institute, University Health Network, Canadac
Department of Medical Biophysics, 6 Queen’s Park Crescent, Toronto, Ont., Canada M5S 3H2
https://pubpeer.com/publications/38A49EC5CB24AAE47F6D1A9C20F484
J Neurosci. 2006 Jun 7;26(23):6377-85.
Presenilin-dependent gamma-secretase-mediated control of p53-associated cell death in Alzheimer’s disease.
Alves da Costa C1, Sunyach C, Pardossi-Piquard R, Sévalle J, Vincent B, Boyer N, Kawarai T, Girardot N, St George-Hyslop P, Checler F.
Author information
Institute of Molecular and Cellular Pharmacology, Coeducational Unit of Research 6097, National Center of Scientific Research/Nice-Sophia-Antipolis University, 06560 Valbonne, France.
https://pubpeer.com/publications/8BA44027EF9B6459BE5A37DD890ABB
LikeLike
It seems Nice is trying to compete with Strasbourg!
LikeLike
Senecio Triodon brings yet another masterpiece, and homage to Picasso.
https://pubpeer.com/publications/04842424AF1A40DFB7CD9ABF9C834D#1
LikeLike
This gel was discussed in NewsTank on 5.06.2019, Actualité n° 148672
“Selon Minoo Rassoulzadegan, « il est clair que dans la piste T (testicules), il y a beaucoup plus de signal que pour les mêmes charges dans les autres pistes et il fallait ajuster le signal, sinon on n’aurait rien vu par exemple dans la piste Sp (sperme) ou des pistes somatiques comme K (rein) “
<
blockquote>
Translated
“According to Minoo Rassoulzadegan, “it is clear that in the track T (testicles), there is much more signal than for the same charges in the other tracks and it was necessary to adjust the signal, otherwise we would have seen nothing for example in the Sp track (sperm) or somatic tracks like K (kidney) “.
Is this senior professor trolling or really clueless of basic research integrity? What explanation is worse?

LikeLike
Old story: when scientists are holding hands with politicians….
LikeLike
Cancer Res. 2006 Jul 1;66(13):6861-70.
Disruption of autophagy at the maturation step by the carcinogen lindane is associated with the sustained mitogen-activated protein kinase/extracellular signal-regulated kinase activity.
Corcelle E1, Nebout M, Bekri S, Gauthier N, Hofman P, Poujeol P, Fénichel P, Mograbi B.
Author information
1
Institut National de la Santé et de la Recherche Médicale U670, IFR 50, Faculté de Médecine, Avenue de Valombrose, 06107 Nice Cedex 02, France.
Figure 5C. Much more similar than you would expect.
LikeLike
2019 expression concern J Cell Sci. 2001 Jan;114(Pt 2):435-43.
J Cell Sci. 2001 Jan;114(Pt 2):435-43.
Gene trap analysis of germ cell signaling to Sertoli cells: NGF-TrkA mediated induction of Fra1 and Fos by post-meiotic germ cells.
Vidal F1, Lopez P, López-Fernández LA, Ranc F, Scimeca JC, Cuzin F, Rassoulzadegan M.
Author information
1
Unité 470 de l’Institut National de la Santé et de la Recherche Médicale and Unité Mixte CNRS-Université 6549, Université de Nice, France.
2019 expression of concern.
http://jcs.biologists.org/content/132/10/jcs233692
Journal of Cell Science was alerted to duplicated HPRT blots in Fig. 4A and Fig. 7C of this paper. The authors state that the conclusions of the paper are not affected by the duplicated control blots, but were unable to locate the original data from almost 20 years ago. Without the original full blots, the journal is unable to determine whether the results and conclusions reported in the paper are compromised.
The journal is publishing this Expression of Concern to make readers aware of these issues. The authors offer cell lines used in the paper for replication by any interested investigators and apologise to readers for any inconvenience caused.
http://www.enseignementsup-recherche.gouv.fr/cid116696/biographie-de-frederique-vidal.html
LikeLike
2019 retraction for 2003 Frederique Vidal paper.
http://www.jbc.org/content/278/3/1751.long
January 17, 2003 The Journal of Biological Chemistry 278, 1751-1757.
A Novel Germ Line-specific Gene of the Phosducin-like Protein (PhLP) Family
A MEIOTIC FUNCTION CONSERVED FROM YEAST TO MICE*
Pascal Lopez, Ruken Yaman, Luis A. Lopez-Fernandez‡, Frédérique Vidal, Daniel Puel, Philippe Clertant, François Cuzin§ and Minoo Rassoulzadegan
-Author Affiliations
From the University of Nice, INSERM U470, 06108 Nice, France.
2019 retraction.
http://www.jbc.org/content/294/37/13832
This article has been withdrawn by Pascal Lopez, Luis A. Lopez-Fernandez, François Cuzin, and Minoo Rassoulzadegan. Ruken Yaman, Frédérique Vidal, Daniel Puel, and Philippe Clertant could not be reached. The Hss26 Northern blots from Fig. 3 (A and D) were previously published in Lopez et al. (2002) Mol. Cell. Biol. 22, 3488–3496, with some data representing different experimental conditions. The withdrawing authors state that the same nitrocellulose filter was reused for two different probes and so the control is the same for both publications. The image in Fig. 6B was inappropriately manipulated. The withdrawing authors state that after 20 years, they cannot provide the original data.
LikeLike
They could not even provide the original data back in the day, so zero chance after 20yrs.
LikeLike
Pingback: For Better Science
Pingback: Charbel Massaad et la lutte contre la fraude – For Better Science
Pingback: CEA declares Anne Peyroche's data fakery scientifically correct – For Better Science
I would add some points since I was student at the university where she started her career.
In the beginning, she was head of the Biology department while I was student. We had a very good contact. I knew she wasn’t a good scientist, everyone at the university knew about fake data, and Minoo was also reputated for being a bit special. But I thought she was a good manager though, she helped me in various things I organized for students (I ended up organizing some parties with the EMBO meeting 2012, aiming to get local students in touch with yop scientists).
But at that University, we had a president, Albert Marouani, who was famous for a lot of “affairs”. I had the national “cour des comptes” get involved in university finances, but of course it was the accountant’s mistake and he got fired… I can tell some stories about how some lab heads got paid a normal salary + extra hours for giving classes…. TO THEIR PHD STUDENTS. Yes. having a PHD student could, under certain circumstances (being friend with the right person) allow you to get paid 35 weekly hours for having a phd student in your lab.
So knowing a lot of affairs, especially concerning student representatives, I told Vidal about one important affair concerning FACE06 student corporation. In fact, like i documented in french here : https://www.facebook.com/may.hem.94/posts/10218304977549654?hc_location=ufi the former head of the student association created a little company “myappyzz” and received 10 000 € to program an official university app, which never existed actually. This student corporation, FACE06, is nationally affiliated to FAGE.
So when Vidal ebcame president, we had high hopes. She cleaned and sorted out some of the things (like the extra hours) and I have to admit I was at first pretty confident and happy. But she never used the documentation i published on facebook, proving that student corporation did some unclear things.
Worse. She started befriending with exactly that corporation. She befriended so much that when she turned minister of research, she chose to take the head of FACE06 as a consultant. https://www.nicematin.com/politique/un-nicois-de-24-ans-nomme-conseiller-de-la-ministre-de-lenseignement-superieur-144581
So she knew this corporation was not clean at all and took their leader with her.
But even worse, after macronleaks revealed, we discovered that the head of FAGE secretly negotiated with Macron. So obviously, once elected, she came to their congress :
https://www.fage.org/news/actualites-fage-federations/2018-10-03,fage-intervention-ministre-frederique-vidal-congres-national-2018-nice.htm
Here are the wikileaks documents :
https://wikileaks.org/macron-emails/emailid/54630
What is FACE06 useful for ?
Well a lot of students complained about how science is managed, and how students are selected and can’t get to study because of social injustice. One student immolated in front of the service responsible for student help last year :
https://www.20minutes.fr/societe/2690055-20200108-immolation-feu-etudiant-lyon-deux-mois-apres-toujours-coma
But every time protests rise, FAGE students, like an army, organize counter demonstrations supporting the government decisions, and saying “we want to study stop protesting we wanna work” or “no to the blocus”.
https://www.20minutes.fr/societe/2269331-20180511-universites-72-etudiants-opposes-blocages-selon-fage
A little story ; during my PhD at the University of Nice, Minoo attacked a technician with a STAPLER, she threatened and was like… totally mad. Since then I had a tiny mini-stapler, we called it “mini-minoo”, and it was an emblematic tool of our lab. Nice little anecdote 🙂
LikeLike
Pingback: Interview with JBC research integrity manager Kaoru Sakabe – For Better Science
Pingback: Creating realistic fake data – Brushing Up Science
Pingback: The secret COVID-19 cure of Pasteur Lille – For Better Science