This new post by Smut Clyde and Tiger BB8 will introduce you to Dr Li Jia, a young Chinese cancer researcher from Dalian whose academic career can only be described with the Olympic motto “Faster, Higher, Stronger”. And just as in Olympia, that was only possible due to unashamed doping. There is no lab technology Jia cannot fake: western blot, flow cytometry, RT-PCR, colony assays, immunohistochemistry: she is the champion athlete of that fraud pentathlon, and beyond. Numerous journals, published by Wiley, Elsevier, Springer Nature and PLoS had the dubious honour of hosting her achievements, often repeatedly.
Nobody wondered until now how Jia, who received her PhD in 2006 and became full professor only two years later, managed to achieve such stupendous research output. And why should anyone wonder? After all, the number of papers and their cumulative impact factor count, not whether the data is actually real and if any experiments were ever performed at all. Li Jia’s research record proves that you can create several independent research datasets using one single picture, if you only apply some elbow grease in Photoshop.
Li Jia, the Research Fraud Olympian
By Smut Clyde and Tiger BB8
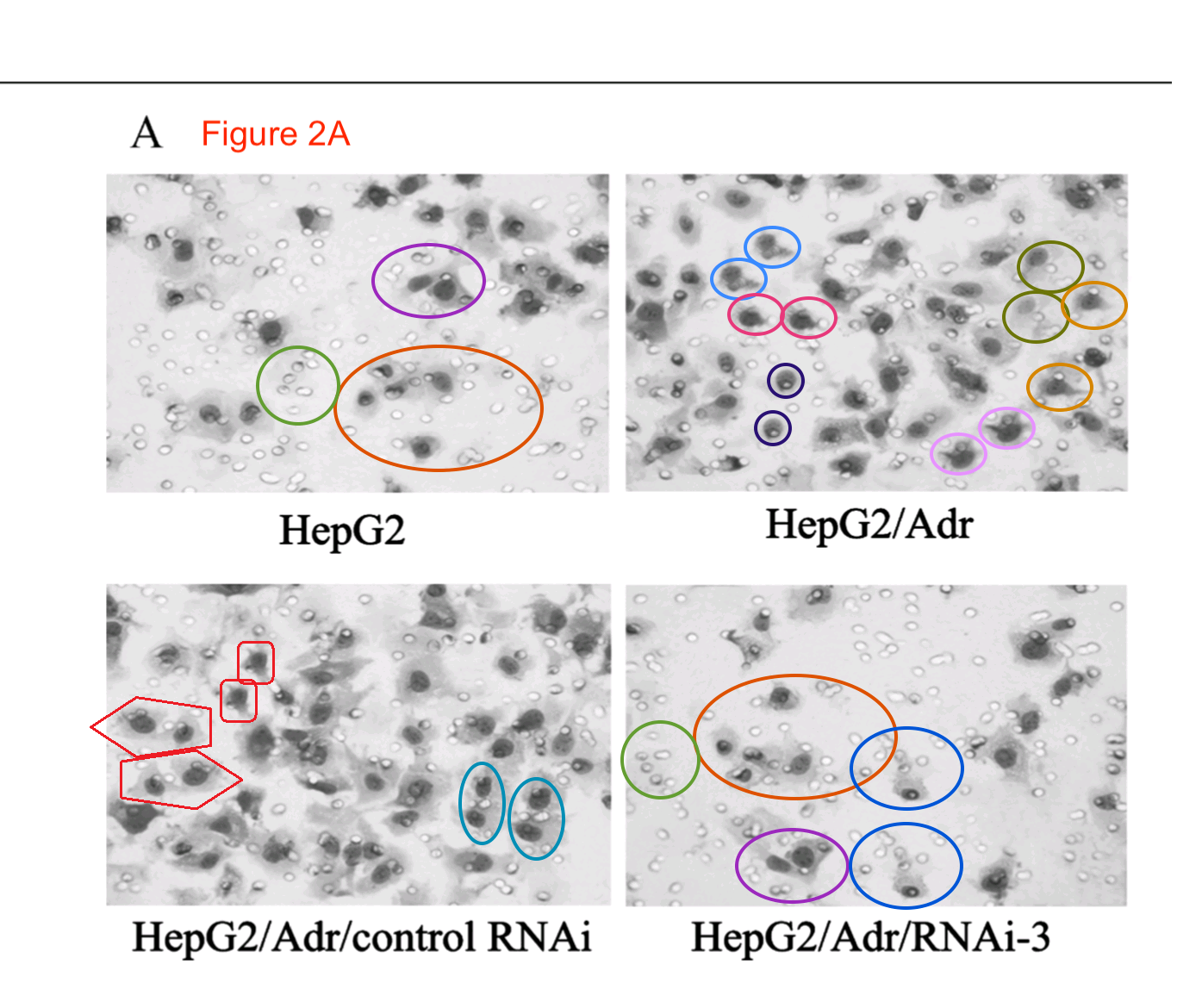
Feel free to admire these panels. The angular silhouettes are not cranes in flight or fish schooling among bubbles, but vat-cultured carcinoma cells from a lineage prone to metastases; they have been lured through pores in a microfilter and then stained, to quantify how different conditions and deterrents affected their invasive tendencies. Sadly in light of the positive results of this ‘migration assay’, it did not exist outside Photoshop, and in comments at PubPeer, ‘Bacillus Mannanilyticus‘ has helpfully circled some of the clone-tool-multiplied cells in Zhao et al Mol Cell Proteomics 2014:


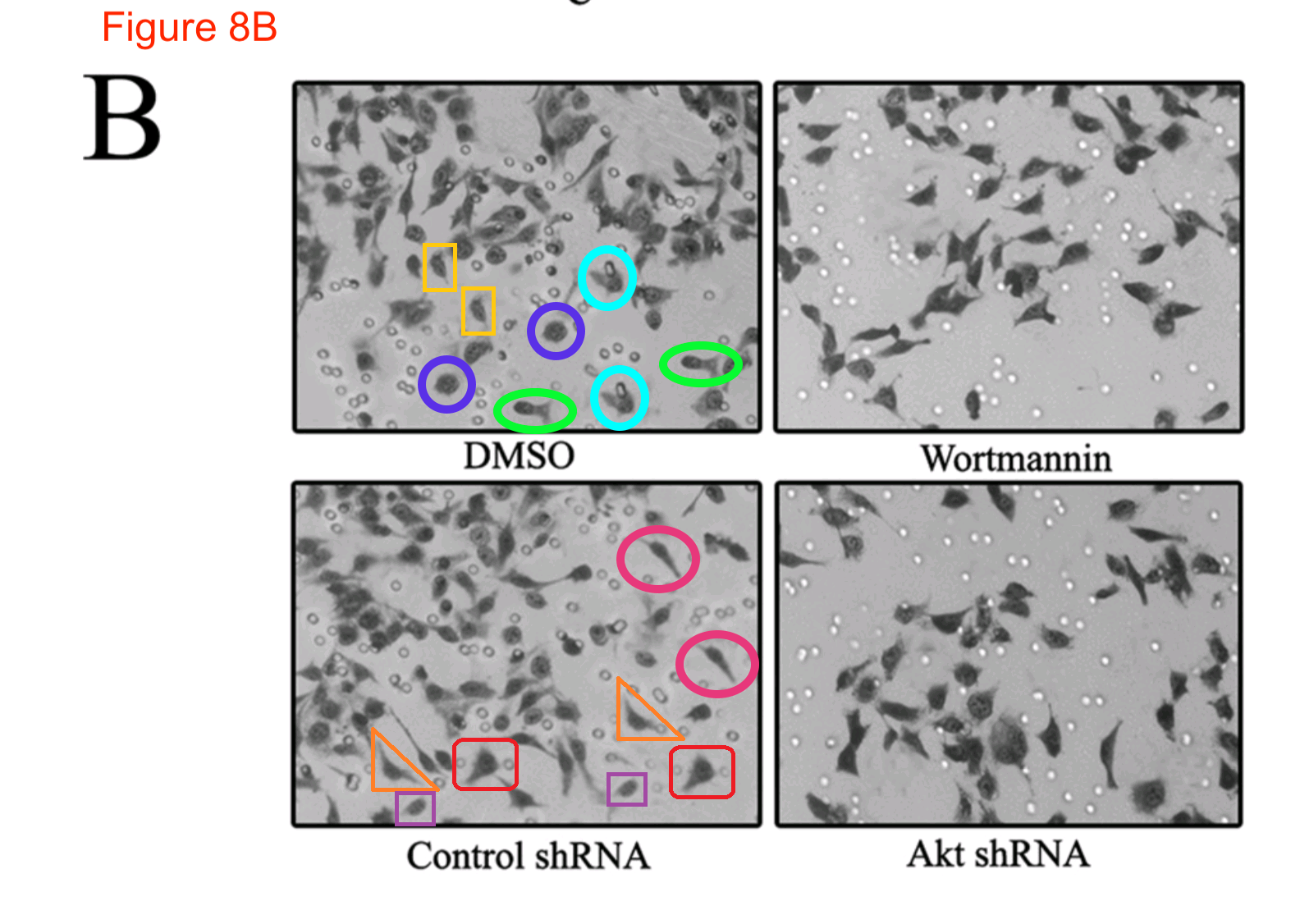
The authors took the criticism to heart and recently retracted the paper —
“This article has been withdrawn at the request of the authors. It contained several inappropriately processed and incorrect figures, Figs. 3C, 3D, 4B, 4C, 4D, 5B, 5D, 6B, 7B, 7C, and 8B. On behalf of all authors, the corresponding author has taken full responsibility and apologizes to the readers of Molecular & Cellular Proteomics for submitting and publishing the erroneous article and any inconvenience caused”.
— though without crediting PubPeer or B. Mannanilyticus… perhaps they learned of the cloning accidents through other channels.
An earlier retraction for Ma et al Biochim Biophys Acta 2014 from the same research group hinged on overlapping microphotographs. Let us say that the number of stained tissue slices (and thus the number of distinct treatments) represented in Figures 5 and 7 was smaller than the number of panels in those subdivided diagrams. Like roofing tiles, or Ordnance Survey maps, the panels share many edges in common.

“This article has been retracted at the request of the authors. It contained several inappropriately processed and incorrect Figures. On behalf of all authors, the corresponding author has taken full responsibility and apologizes to the readers of BBA Molecular Basis of Disease for submitting and publishing the erroneous article and any inconvenience caused.”
This is our introduction to the oeuvre of Dr Li Jia. Now care is required when counting how many of her papers have been discussed at PubPeer (and the ResearchGate archives are little better), for there are some quite separate names in Chinese which become ‘Li Jia’ when romanised by the Pinyin system. Here we are concerned with the Dalian Medical University cancer researcher, who received her Ph.D in 2006 – the basis for her initial papers (seven of them with her Ph.D advisor as main author and herself as first author). That Li Jia.
Advised by Professor Jianing Zhang, a Kyoto University-trained Doctor of pharmaceutical sciences, Li Jia received her PhD on Pathology and Pathophysiology from Dalian Medical University in 2006, with her PhD thesis titled “Effects of CD147 glycosylation on the lymphatic metastasis in murine hepatocarcinoma cells lines”. It won a title of distinguished PhD thesis of the Liaoning Province in 2007. After her PhD, Li Jia remained in her university and became full professor in August 2008, her numerous research grants established her academic success and earned her an own lab and a big team.

What draws us to this oeuvre is the breadth of fake-data styles it encompasses. The versatility is impressive. Future conferences on Data Integrity could use it for an overview of the range of graphical techniques available for inventing results in cell biology. In fact the sheer abundance of material to include has forced me to be systematic, and to arrange everything along a time-line. I have limited its scope to the decade 2006-2015, with 27 papers receiving post-publication review at PubPeer (the two retractions, both from 2014, are [22] and [23]).
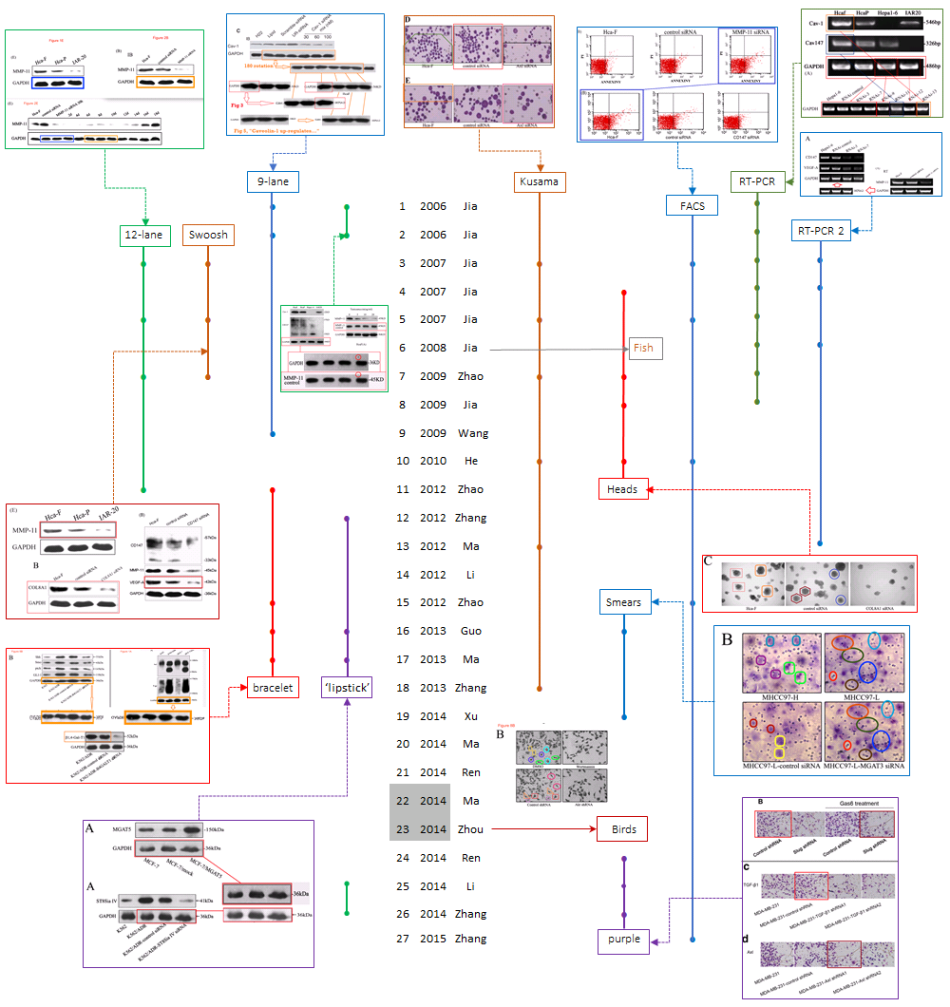
 The time-line is not comprehensive; it features images that tie everything together by occurring more than once. There was no space for exuberant but one-off fabrications, such as Figure 3B from [3] in which hepatocarcinoma cells multiply to form colonies in nutrient agar (peer-reviewers were not concerned that identical colonies arose independently, such as the Mickey Mouse silhouette marked with a blue trapezoid in the first and third panels).
The time-line is not comprehensive; it features images that tie everything together by occurring more than once. There was no space for exuberant but one-off fabrications, such as Figure 3B from [3] in which hepatocarcinoma cells multiply to form colonies in nutrient agar (peer-reviewers were not concerned that identical colonies arose independently, such as the Mickey Mouse silhouette marked with a blue trapezoid in the first and third panels).
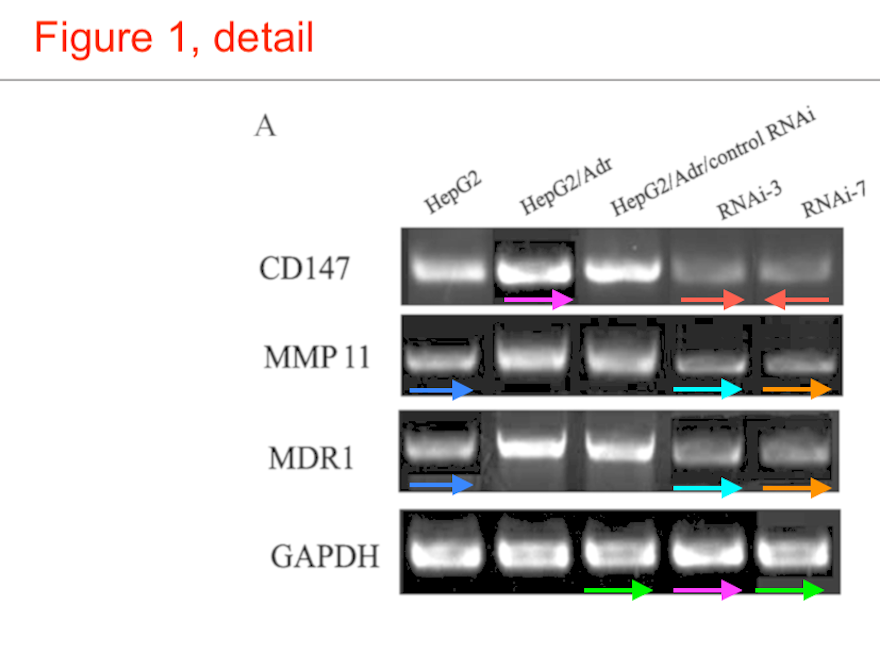
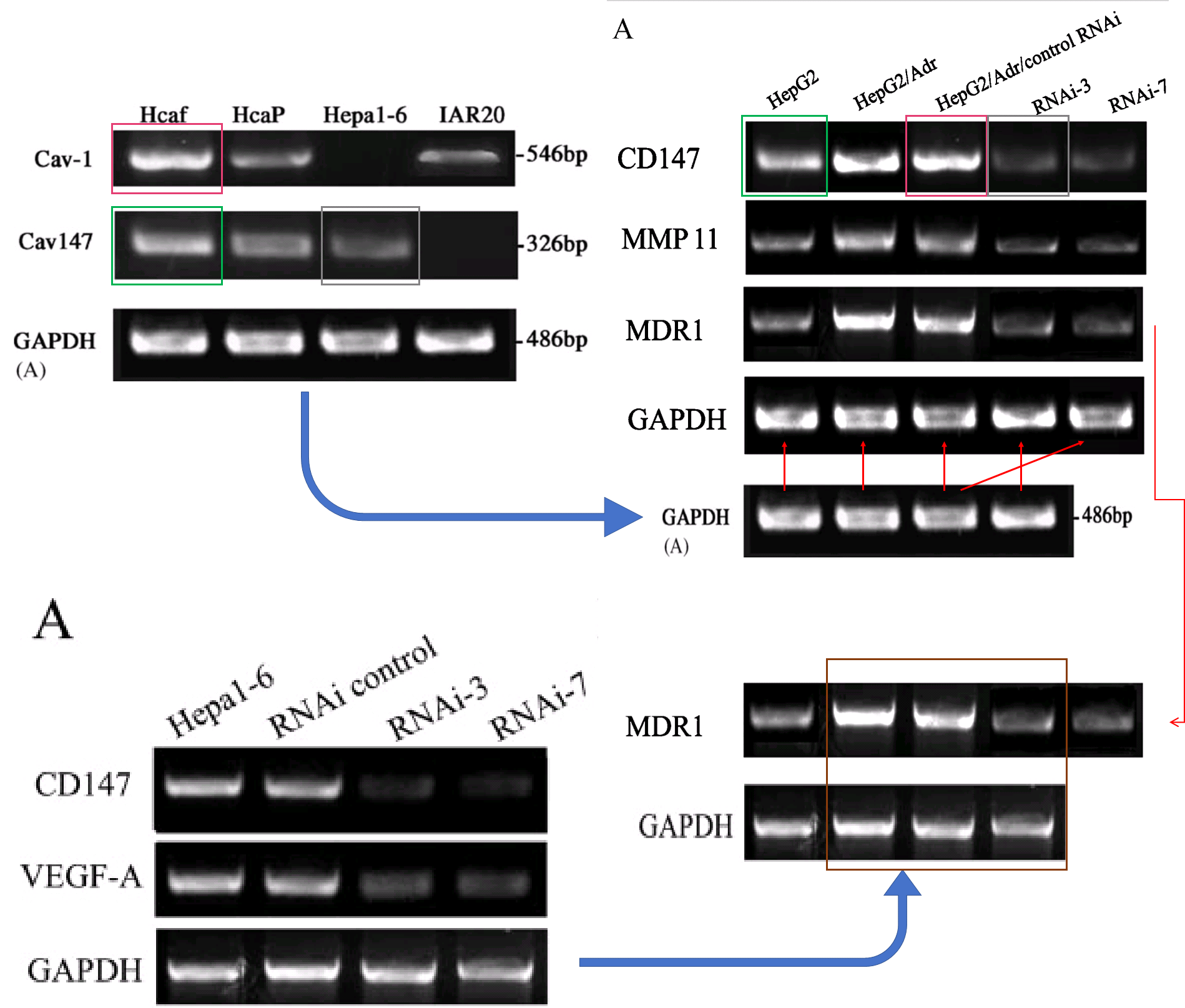
Over to the left of the chronology I have arranged the various manifestations of Western blots. The emphasis is on the metamorphoses of a small number of loading controls, the labels on the lanes changing according to the exigencies of different experiments. Now one school of thought regards loading controls as needless, time-wasting formalities, and relies on the process of harvesting cell contents to capture the same number of cells and yield the same concentration of proteins for each experimental situation (i.e. for each lane in the blot) – making it nugatory to control for that concentration. Jia was evidently of that persuasion.
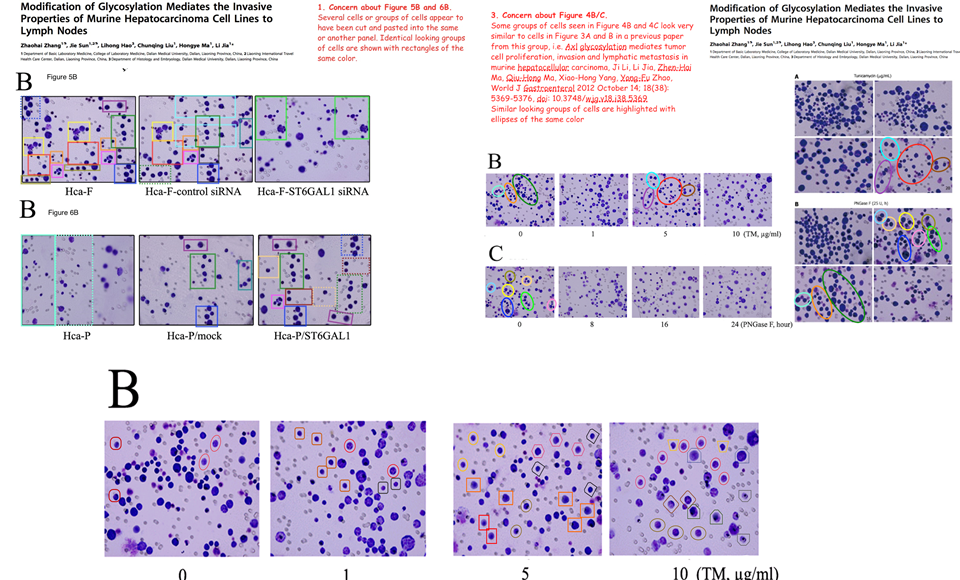
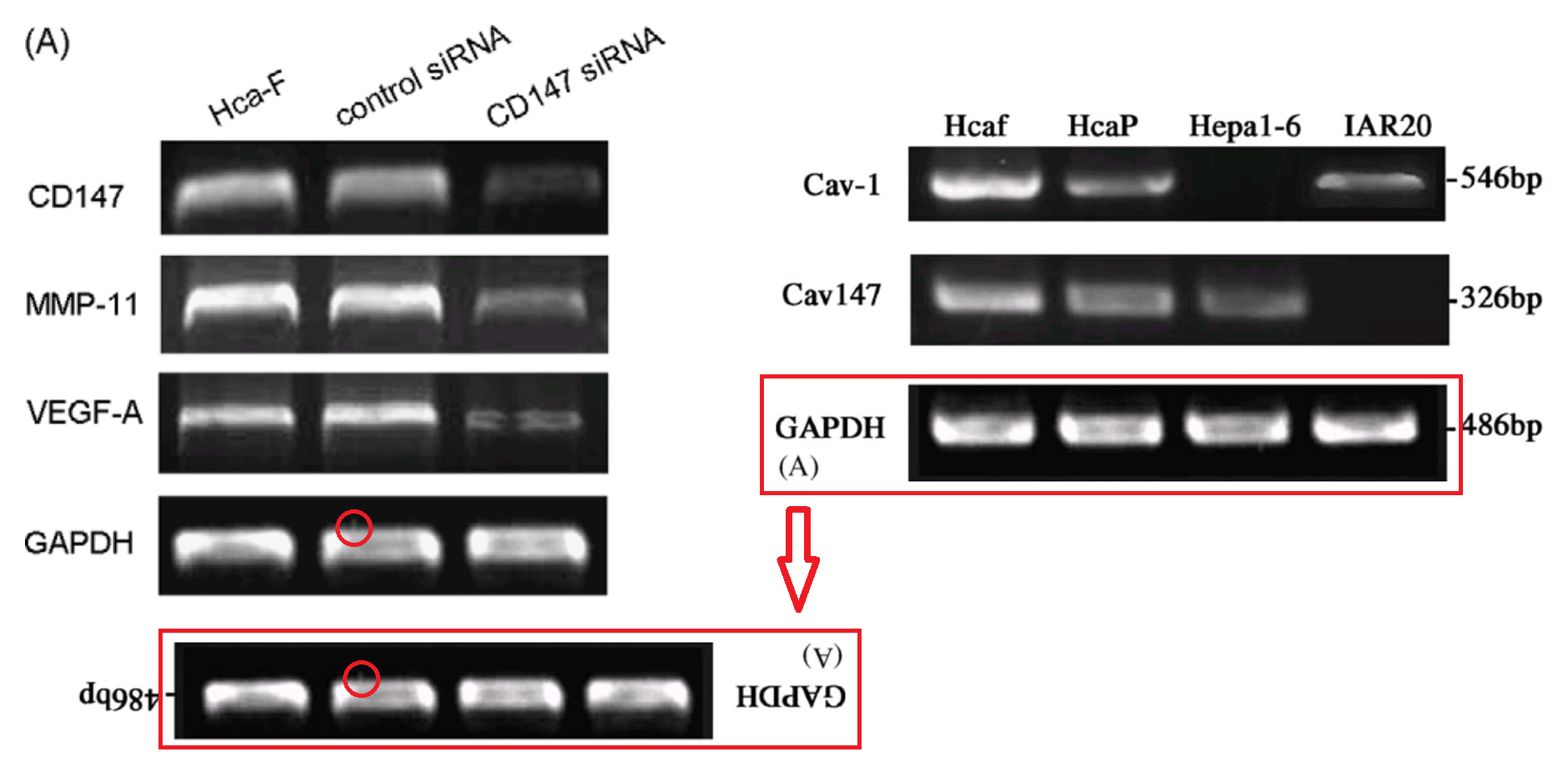
However, that defence does not apply to the “swoosh” series of metamorphoses (from papers #3, #5 and #7), in which the recurring blot serves as empirical evidence for the three substantive proteins MMP-11, COL8A1 and VEGF-A.
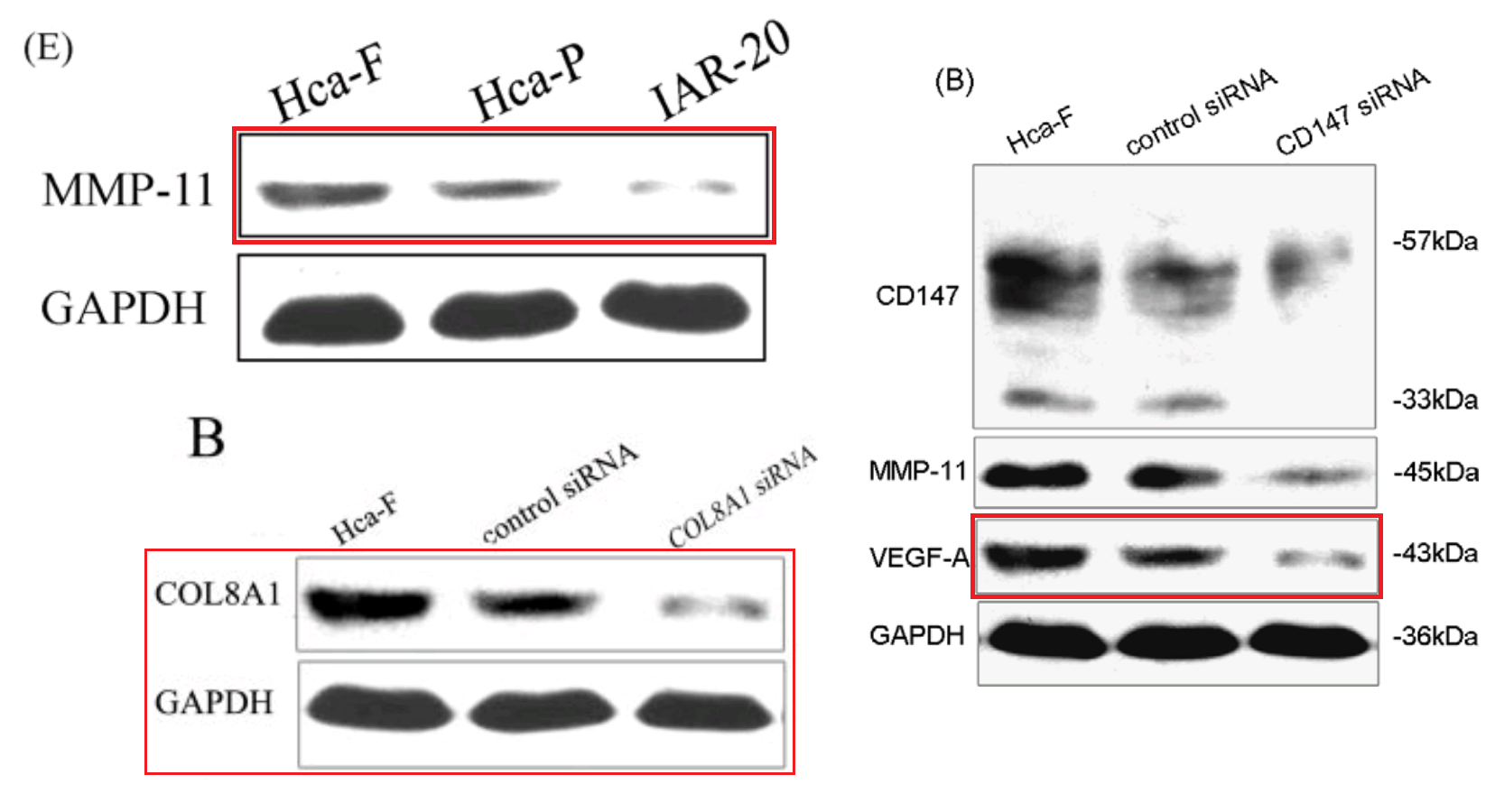
Apparently the same slice of gel can be a GAPDH control from different cell-lines in [1], or Matrix Metalloprotease-11 expression in the same cell-line exposed to different concentrations of tunicamycin, in [2]. None of this is promising for the Ph.D thesis source. Nor is it a good advertisement for the supervisor and main author Jianing Zhang, who had left Dalian Medical University in 2013 and took the position of the Dean of the School of Life Science and Medicine in a neighbouring, but much more prestigious Dalian University of Technology.
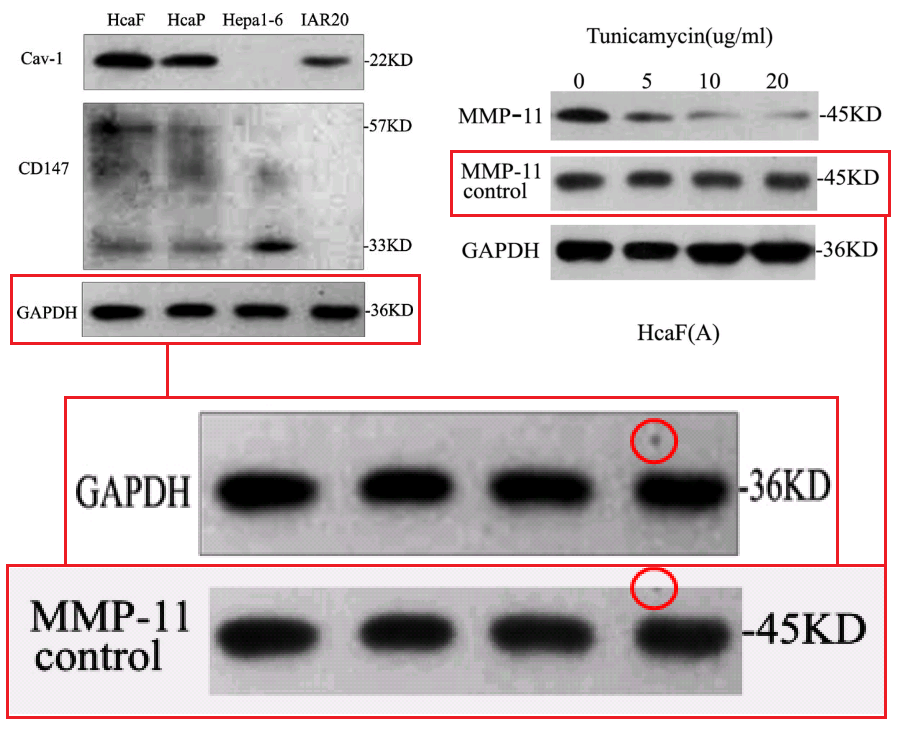
That second manifestation as MMP-11 opens up further vistas of exploration when we look at its nominal loading control. It overlaps with other controls in the same paper, and they with others still… until we can reconstruct a single Ur-blot with nine lanes, from which various segments were extracted to provide the GAPDH controls for five separate blots, comparing multiple gamuts of conditions and spanning papers [1], [2] and [9].

No reconstruction is needed for a 12-lane GAPDH blot, which was shown in full in Figure 2E of [3], ensuring stability over a time-course.
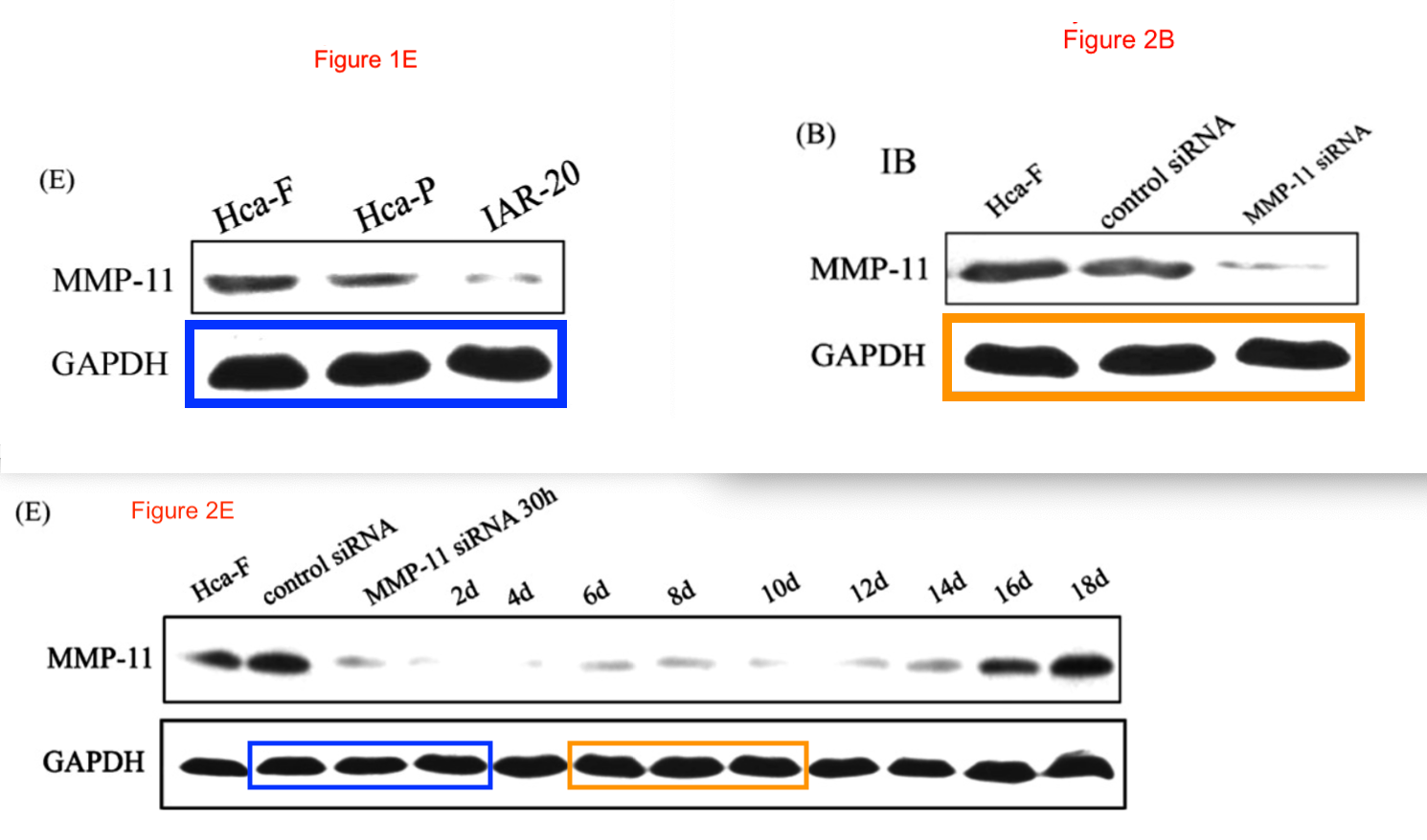
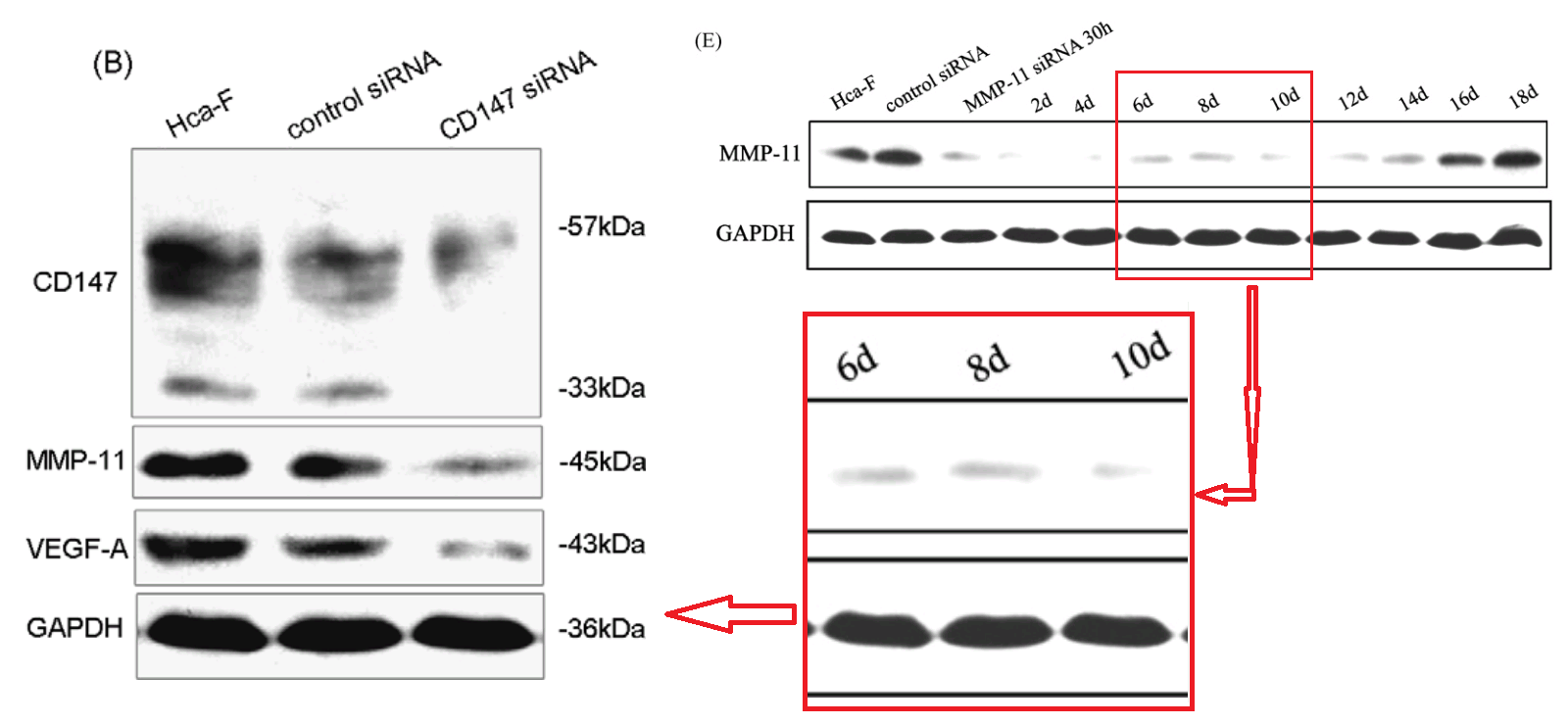

Short excerpts from it can be found controlling for other trials in [3], [5] and [7] (where we met them already), and [11].

In this time period of 2006 to 2010, many of Jia’s image integrity problems can be traced back to her PhD thesis. However, new material did enter the repertoire after 2010. One eight-lane band will always be “lipstick traces” to me. Various fragments featured as Figure 5A of [12], as 6A of [17], and four times in Figure 5 of [16].

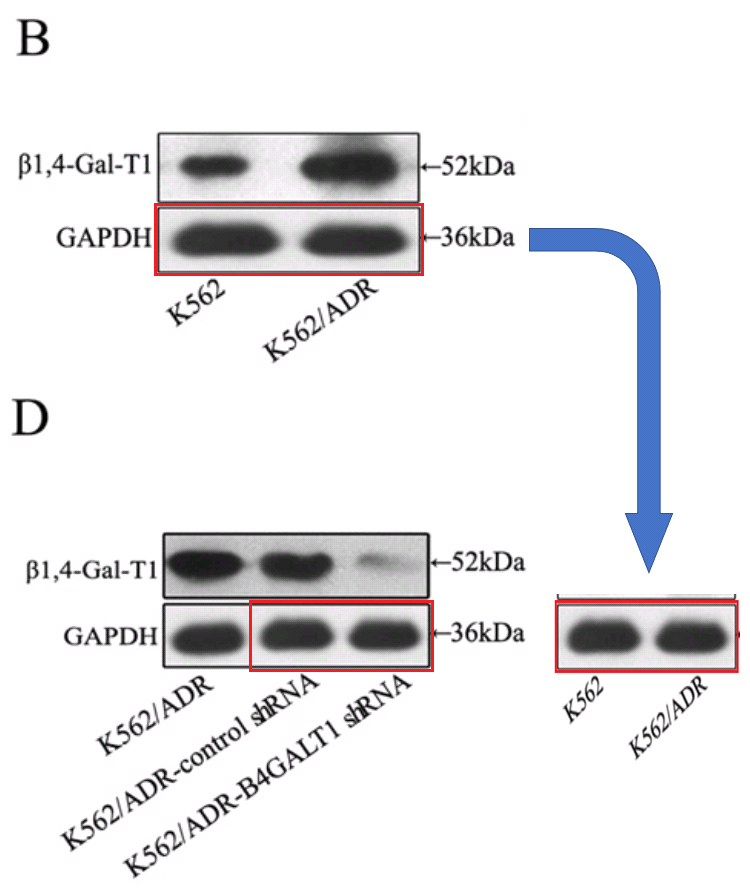
The time-line includes a 2011 gap between [10] and [11], when Jia spent a year in Japan on a research fellowship (May 2010-May 2011) on the basis of her promising youthful productivity. Evidently this provided fresh blots, among them a second six-lane segment featured in overlapping segments in [11], [15], [16] and [17]… I like to think of it as the “charm bracelet”.
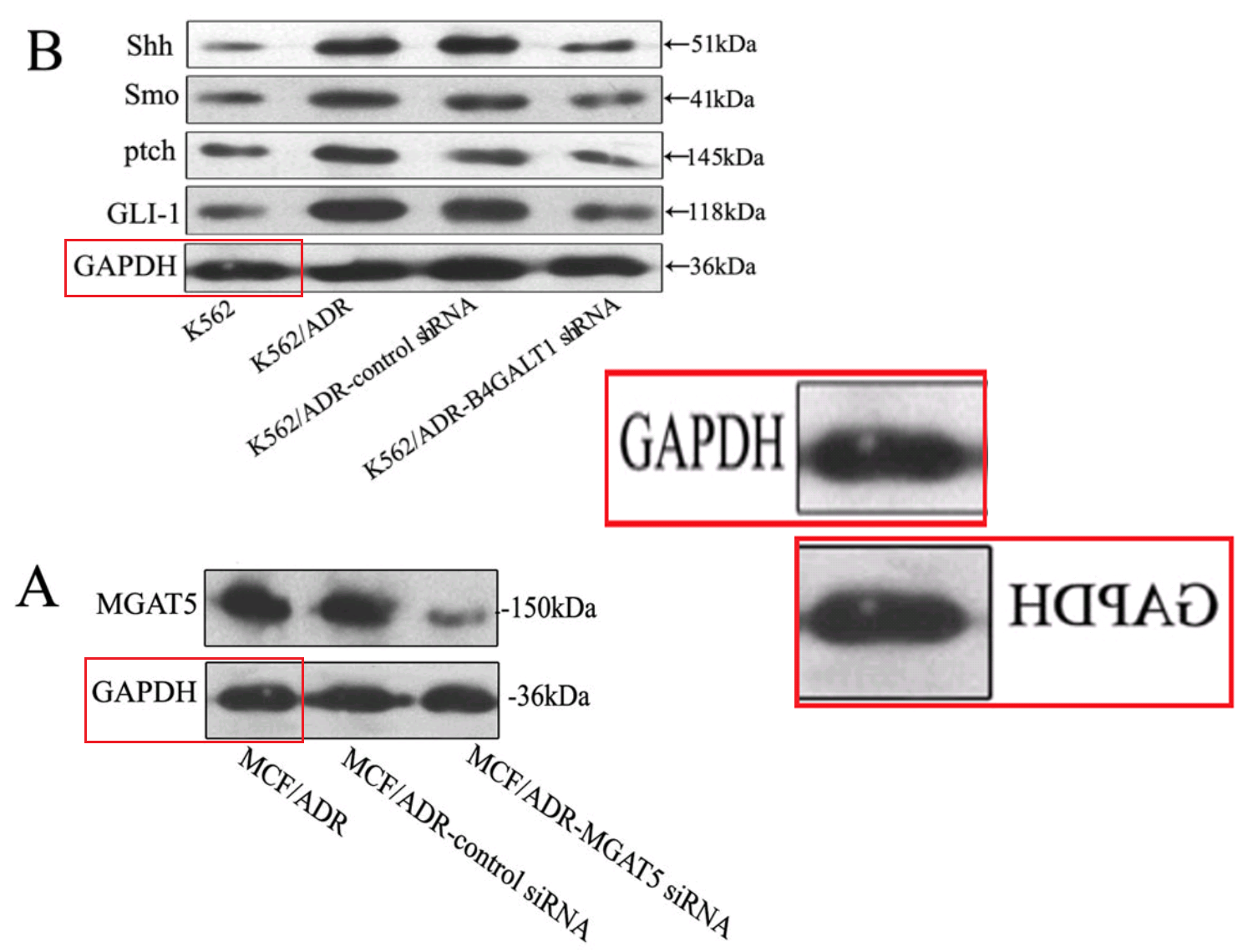
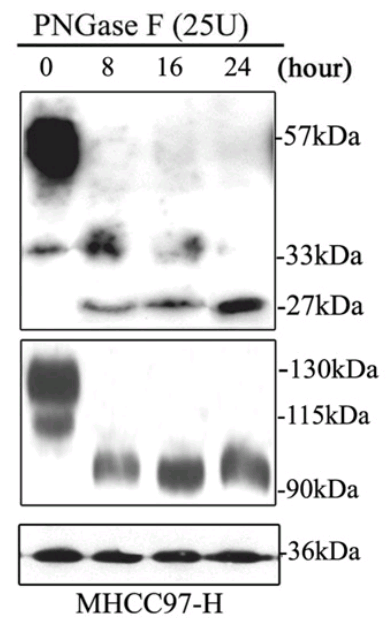
In total, counting Figure 1D of [11] as well as 6B, the charms appeared five times. As its debut it took the form of the substantive protein β1,4-Gal-T1 to justify a crucial claim of the paper: that the expression of that gene was blocked by a targeted shRNA. In that manifestation it acquired a GAPDH loading control of its own, which was less versatile, occurring only twice there and again in [12].

From Western Blots we shift to a genre grouped on the extreme right of the time-line. Here the molecules of interest – to be separated by molecular weight and labelled – are DNA segments, transcribed from RNA in the cellular extracts using Reverse Transcriptase, then amplified with PCR. Repurposed RT-PCR gels are not a novelty, and readers may recall them coming under scrutiny in previous blogging, but Jia took the amplification of RNA message to a new level. Prepare yourself for flotillas of kayaks and Canadian canoes parading across the screen, like a Mohawk war party in a Fenimore Cooper novel.
Two strands of repurposing can be discerned. The less active strand is distinguished by the crumpled prow (or stern) of one of the kayaks, and comprises only [3], [4], [6] and [13].
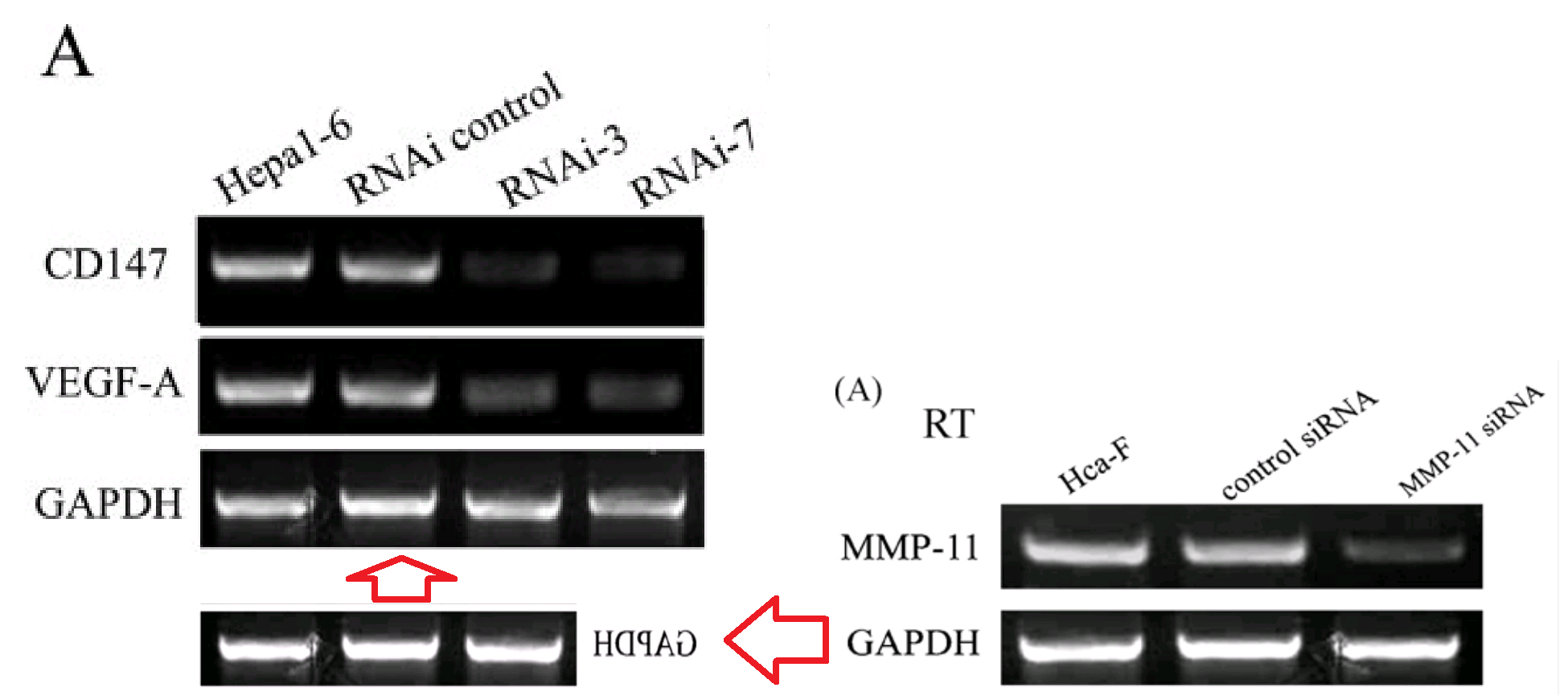


The “core gel” of the second strand is recognisable by the wee peg below the keel of one kayak and by the dot above another one (I surmise that to be the vanguard of the war party, and the leader has raised a small balloon, in the manner of a tour-group leader, to stop the others wandering off and becoming separated). It was cropped to two bands and three lanes for its debut in [1], and to a three-by-four form for [2]

This blot had already been manipulated when it first appeared outside Jia’s thesis: the upturned kayak at the top right corner is a fabrication, spliced in. Anyway, these distinguishing marks allow us to trace the blot through its variants and permutations in [3], [4], [5], [6], [7] and [8]. Sometimes the kayaks are merely flipped vertically [5] or horizontally [7]:
Conversely, in [4] the manipulations reach delirious heights of ransom-note rearrangement to create eight lanes.


The two strands meet in Figure 1A of [6], which is a veritable stamp collection. Even a Helpful Diagram does not really make clear how much the core blots were sliced up and reconfigured.
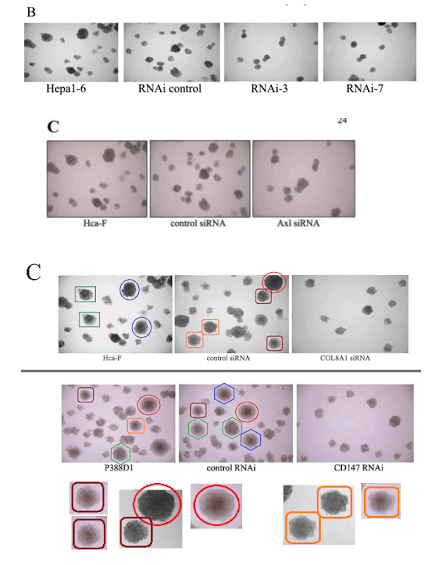
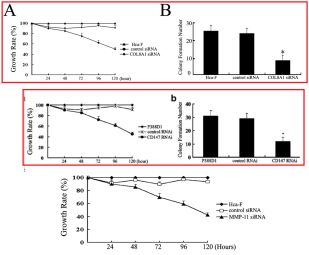
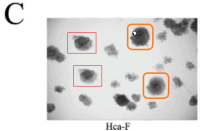
Let’s go back to the Colony Growth assay, and to a group of images recycled across [4], [7], [8] and [10]. Everything can be improved with googly eyes, but also with duplicated colonies. Among other discoveries, it emerged that COLA81 manipulations affect the proliferation of Hca-F cells in exactly the same way as CD147 manipulations affect P388D1 cells… even producing identical colonies (in multiple copies). To my mind this phenomenon deserves a paper in itself.
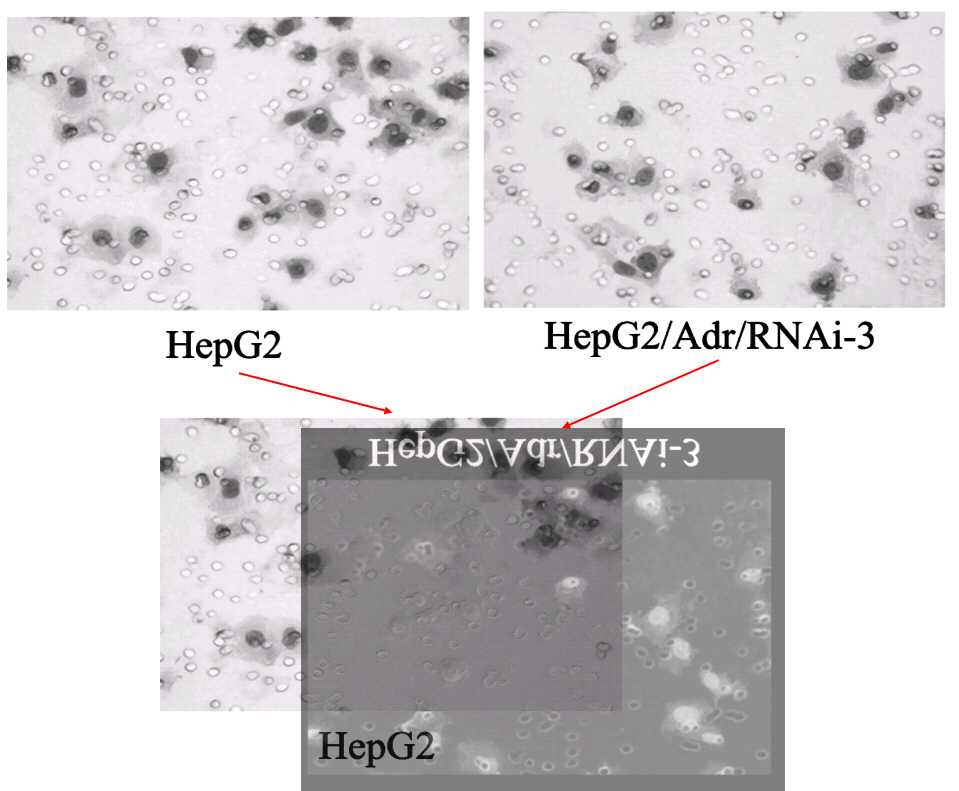
We have already encountered the Invasion / Migration Assay. There is some visual variety in its applications, depending on the specific lineage of cancer cells that had squeezed through pores in the filter. The software enhancement of the images is constant, though, and B. Mannanilyticus noted a number of cloned cells in Figure 3B of [16] (see also 4B).
Some of the repetitions stem from the fact that the upper-right and lower-right panels of 3B are essentially the same, as can be seen when one of them is colour-inverted and superimposed on the other, so that identical areas cancel out. The areas of failed cancellation are where cells were added to one panel, or the other, but not both.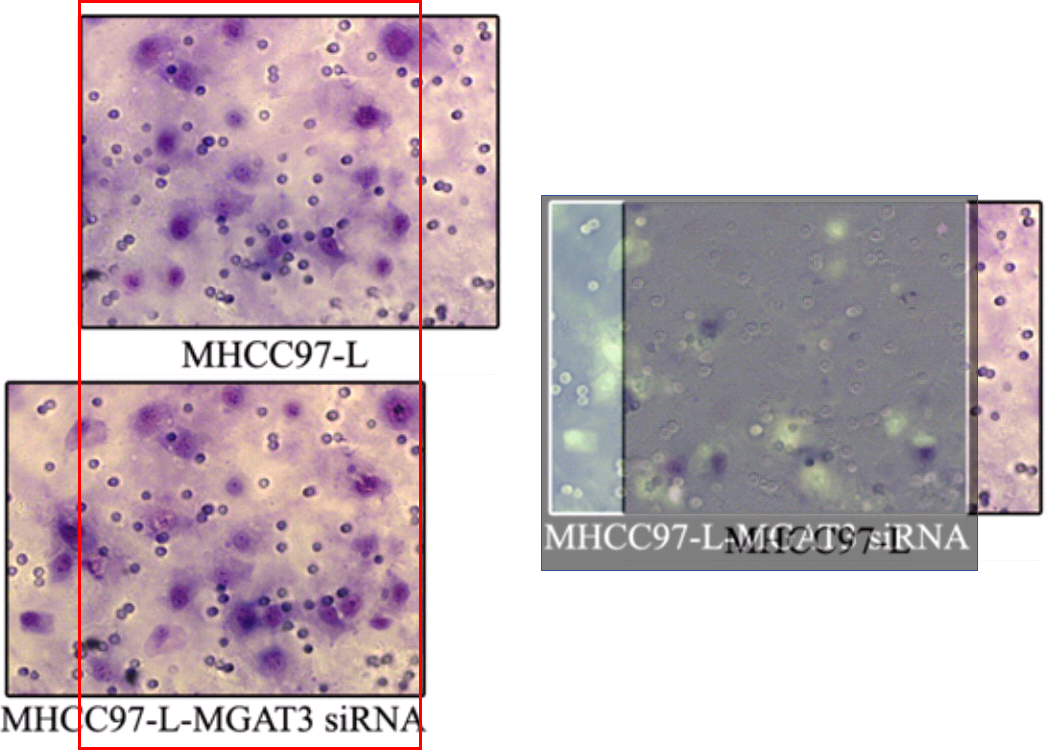
We should also pay attention to the upper-left panel of 3B, for it reappeared in [19] as the top right of Figure 5(b), minus the Photoshop improvements (again, an inversion and superimposition clarify the areas of difference). The repeating sections are confined to other panels of 5(b), and to look at them closely is to be reminded of the crude faces in Appel’s “Vragende Kinderen” paintings.

But we cannot linger on those images for I am impatient to move on to a group of invasion-assay results which may well be hommages to Yayoi Kusama and demanded their own thread in the time-line. Overlapping sections of panels reappear between a half-dozen papers, stretching across seven years, purportedly representing treatment with different siRNAs to discourage the cells’ mobility. This series began in 2007 with Figure 5A of [3], where the most obvious overlaps are with 2E from [10].
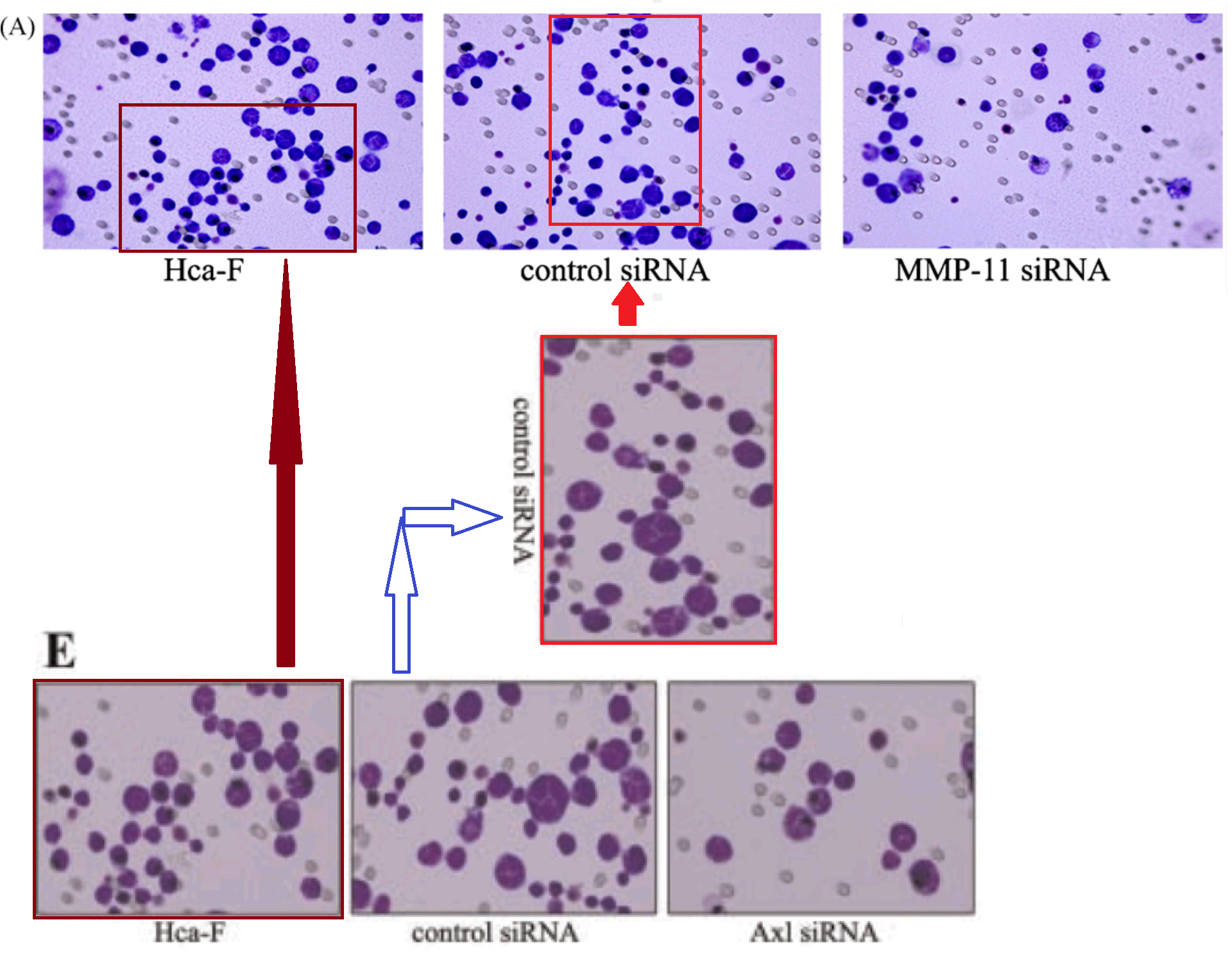
Note that a prominent cell was inserted for the [10] version (or perhaps erased for the [3] version), presumably for aesthetic reasons. The authors continued to improve the images with more dots and manipulations, as if in the thrall of a gaes or whatever is the opposite of trypophobia, and the series build to a crescendo in [13] (2012)…
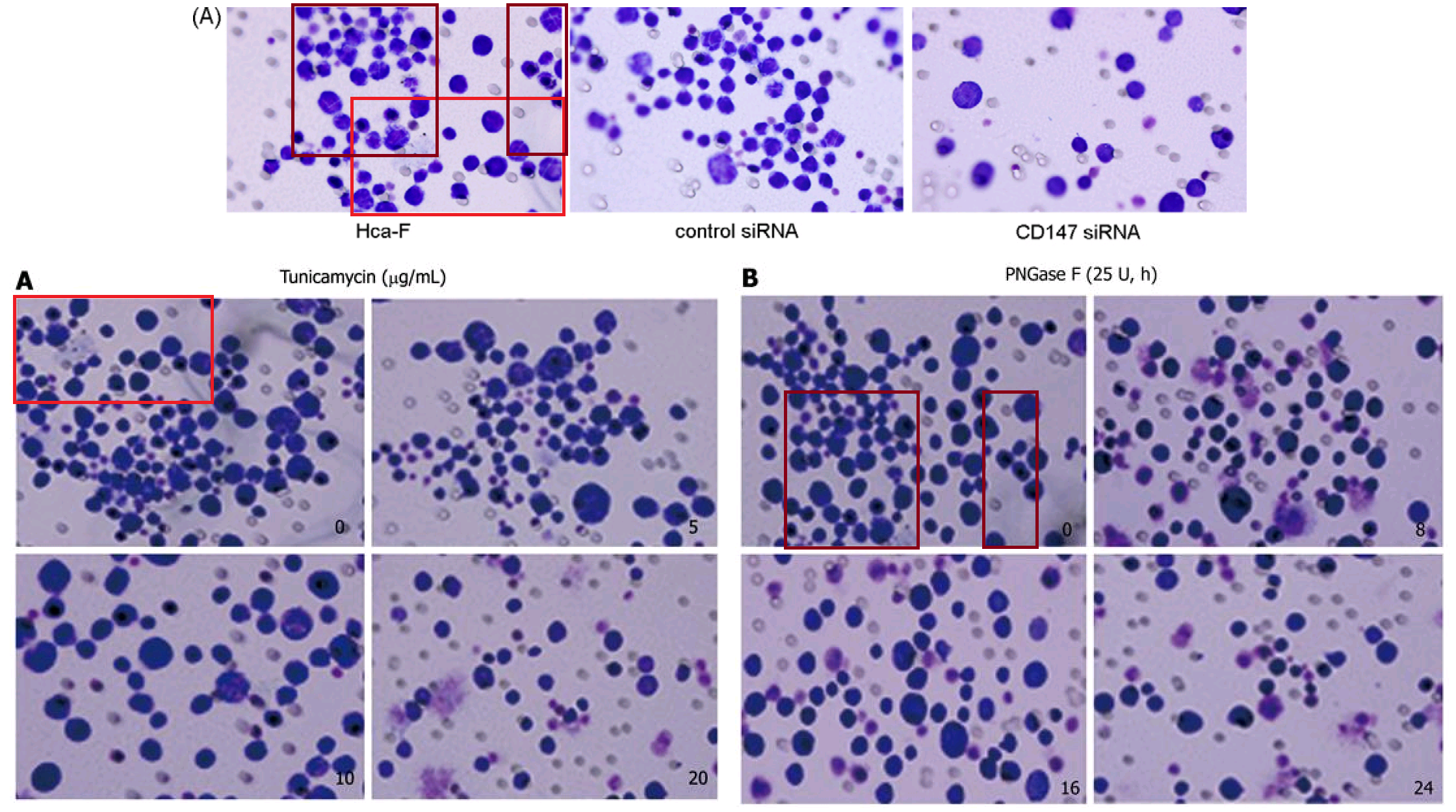
… then reached a dizzying apotheosis of artificiality in [18] (2013).
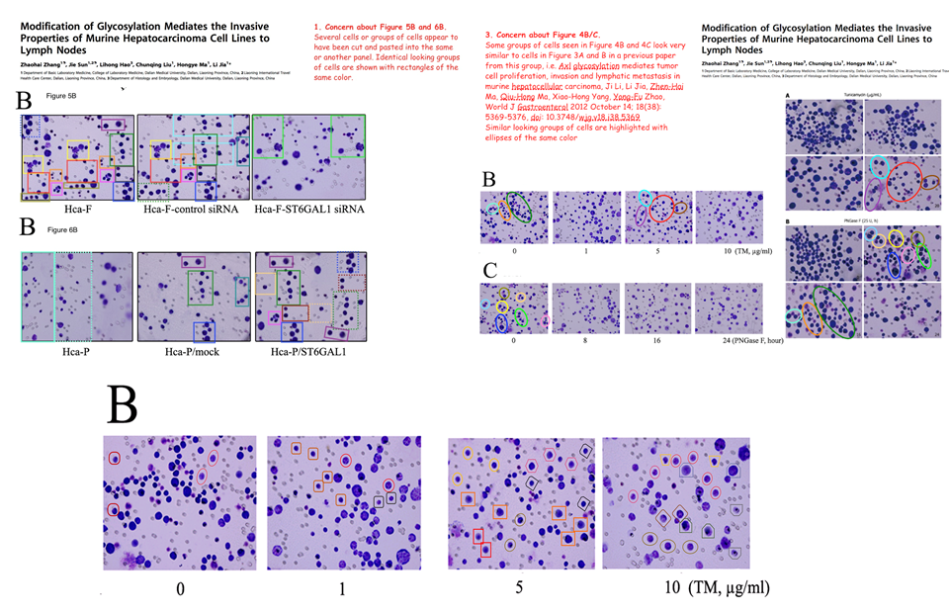

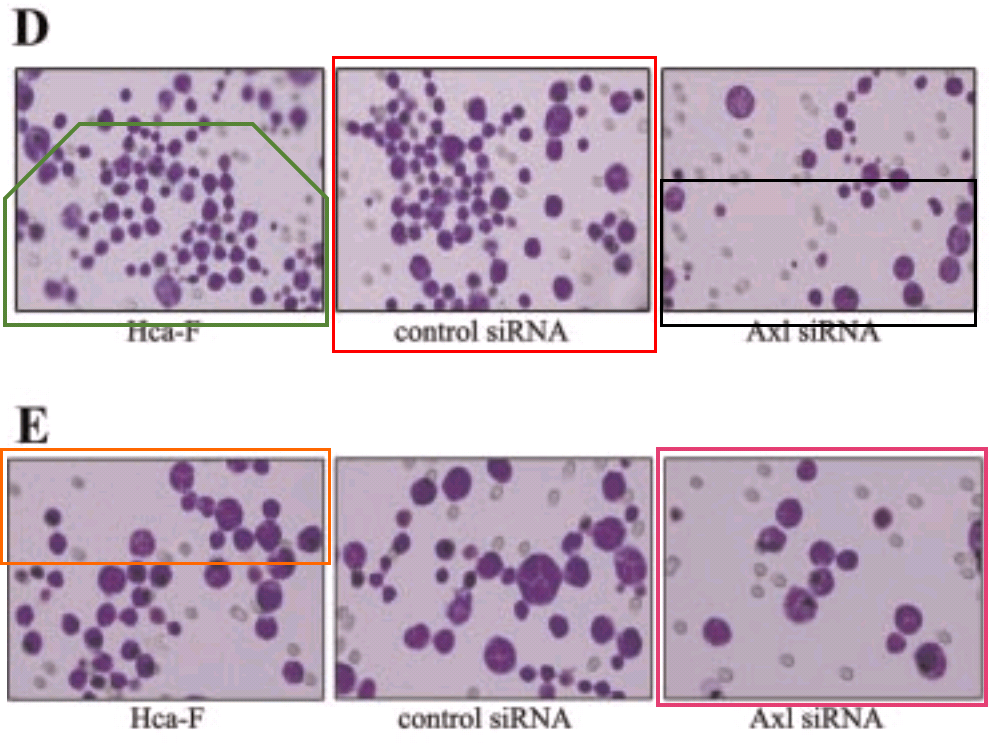 The failure of the PLoS One reviewers to recognise the creativity poured into these figures must have been galling. Anyway, the links of re-use / alteration between papers are spelled out in the PubPeer threads, and rather than take up more space, I have prepared a little diagram to help interested readers follow the routes from one to another.
The failure of the PLoS One reviewers to recognise the creativity poured into these figures must have been galling. Anyway, the links of re-use / alteration between papers are spelled out in the PubPeer threads, and rather than take up more space, I have prepared a little diagram to help interested readers follow the routes from one to another.
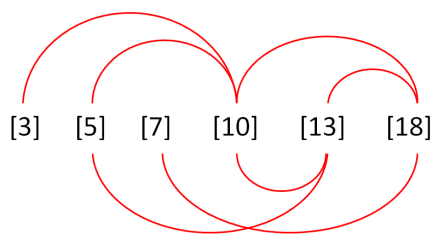
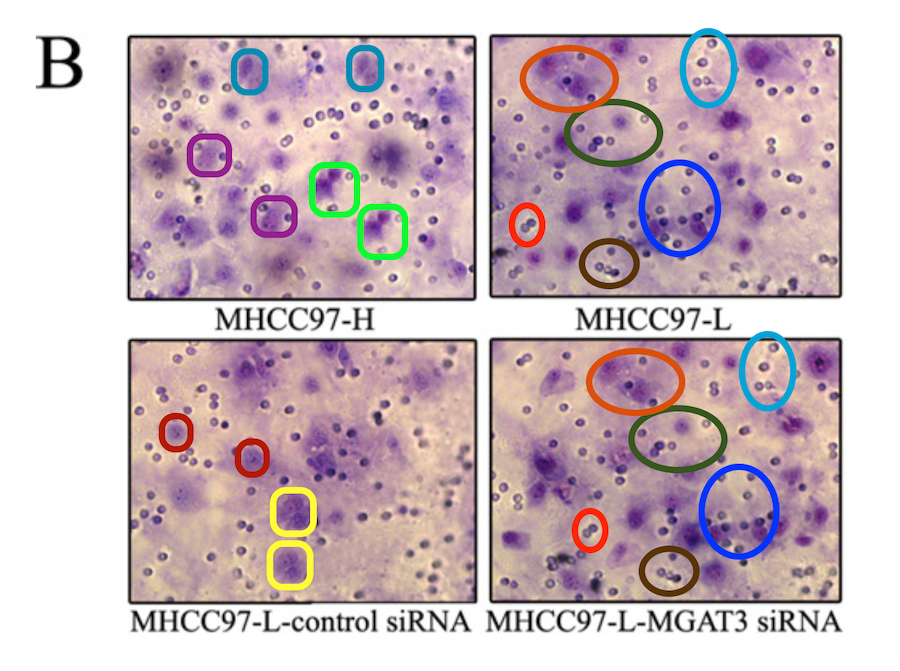
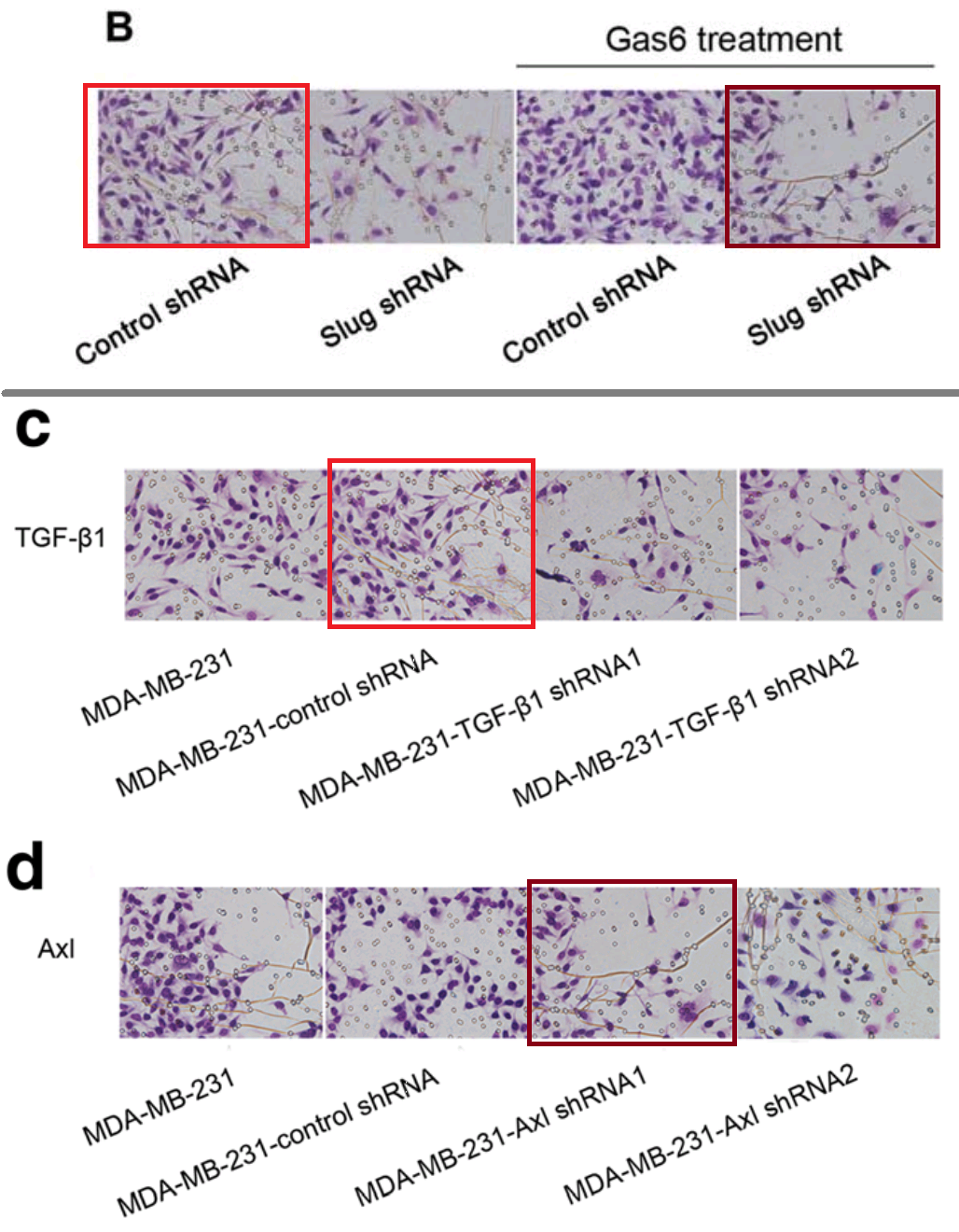 We are nearly finished with Invasion / Migration Assay images. The ones featured in [24], [25] and [26] are close to the angular cranes-in-flight profiles with which we began, though because they are purple I’ll compare them to tropical fish in aquarium tanks. I lied when I wrote that “the software enhancement of the images is constant”, for there is nothing amiss with these examples… except their re-use within and between papers, differently labelled.
We are nearly finished with Invasion / Migration Assay images. The ones featured in [24], [25] and [26] are close to the angular cranes-in-flight profiles with which we began, though because they are purple I’ll compare them to tropical fish in aquarium tanks. I lied when I wrote that “the software enhancement of the images is constant”, for there is nothing amiss with these examples… except their re-use within and between papers, differently labelled.
One visual motif remains to tie these papers together. The FACS-plot strand stretches practically the length of this sample, from [3] to [27], though only five papers take part.
Because the three panels of 3B from [5] portray the protein-expression distributions of murine hepatocarcinoma cells, while panels 1, 2 and 4 of Figure 3A from [7] involve human leukemia cells – measured under different conditions – we expect them not to cancel out when panel is superimposed upon panel. But prepare for disappointment: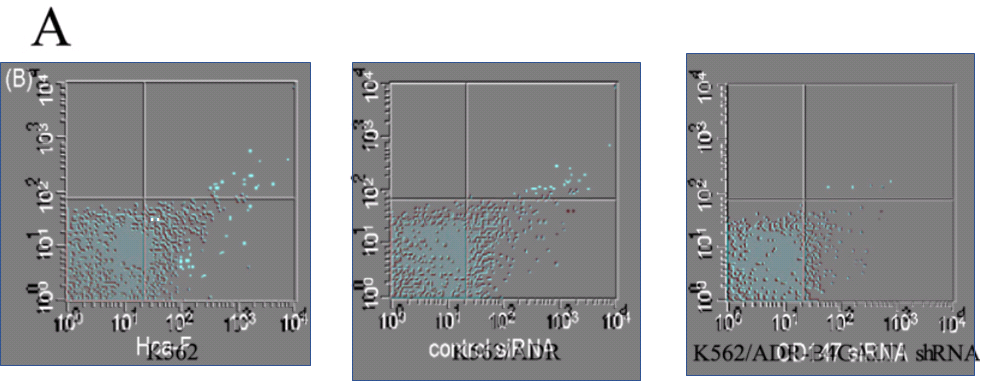
The repetitions are not complete; some points have been added or scraped away (each point indicating a single cell), as if an experimental data-file had been plotted with a different choice of filtering parameters, or simply revisited in Photoshop. As for the third panel of 3A, it is an unnatural chimera, formed by grafting two half-panels together.
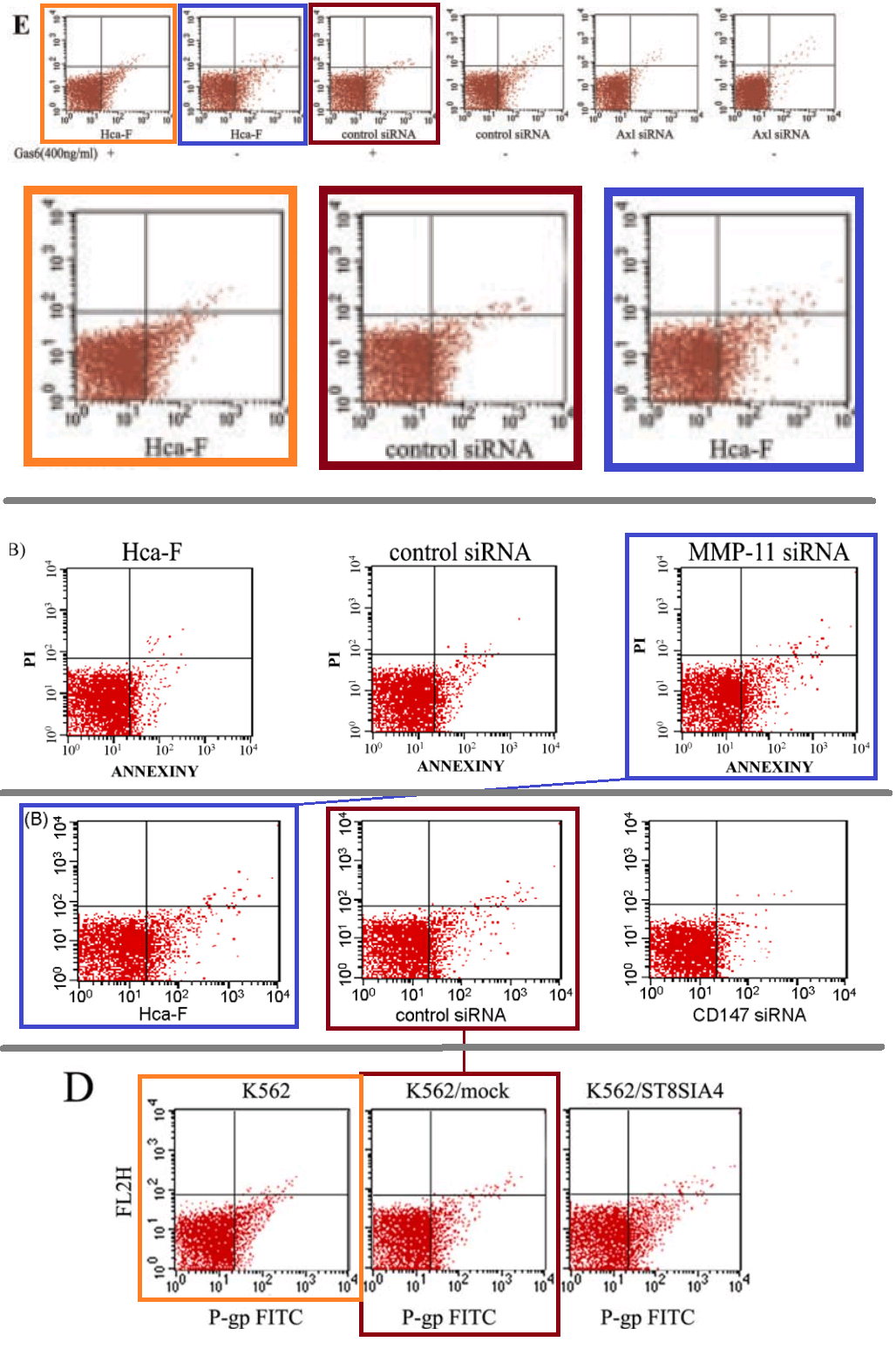 A single color-coded diagram captures the almost-repetitions across Figures 4E from [10], 4B from [3], 3B from [5] and 6D from [27].
A single color-coded diagram captures the almost-repetitions across Figures 4E from [10], 4B from [3], 3B from [5] and 6D from [27].
After 2015, Li Jia’s research shifted direction towards the area of micro-RNA and its role in intra- and inter-cellular regulation. [28] is “Comprehensive N-glycan profiles of hepatocellular carcinoma reveal association of fucosylation with tumor progression and regulation of FUT8 by microRNAs“, from 2016. [29] is “Long non-coding RNA HOTAIR promotes osteoarthritis progression via miR-17-5p/FUT2/β-catenin axis”.
A feature of this newly-popular miRNA genre is its strong visual conventions. Papers follow a template, and as well as Western Blots and FACS plots, readers expect to see immunofluorescent staining and scatterplots of miRNA levels against cancer biomarkers.
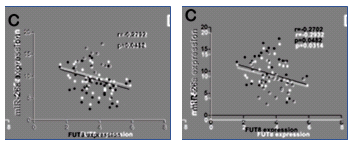
As part of that shift, the Jia Lab left behind egregious displays of forged results. Another 10 threads at PubPeer examine her output since 2015, but with some exceptions they concern trivial image duplications that can be corrected with an embarrassed Erratum notice and a Does-not-affect-the-Conclusions. Indeed, four of the affected papers have been recently amended (coinciding with the Retraction with which we began). Note, too, that these 10 papers are only a fraction of the group’s productivity, with 11 papers published in 2017 and nine in 2018.
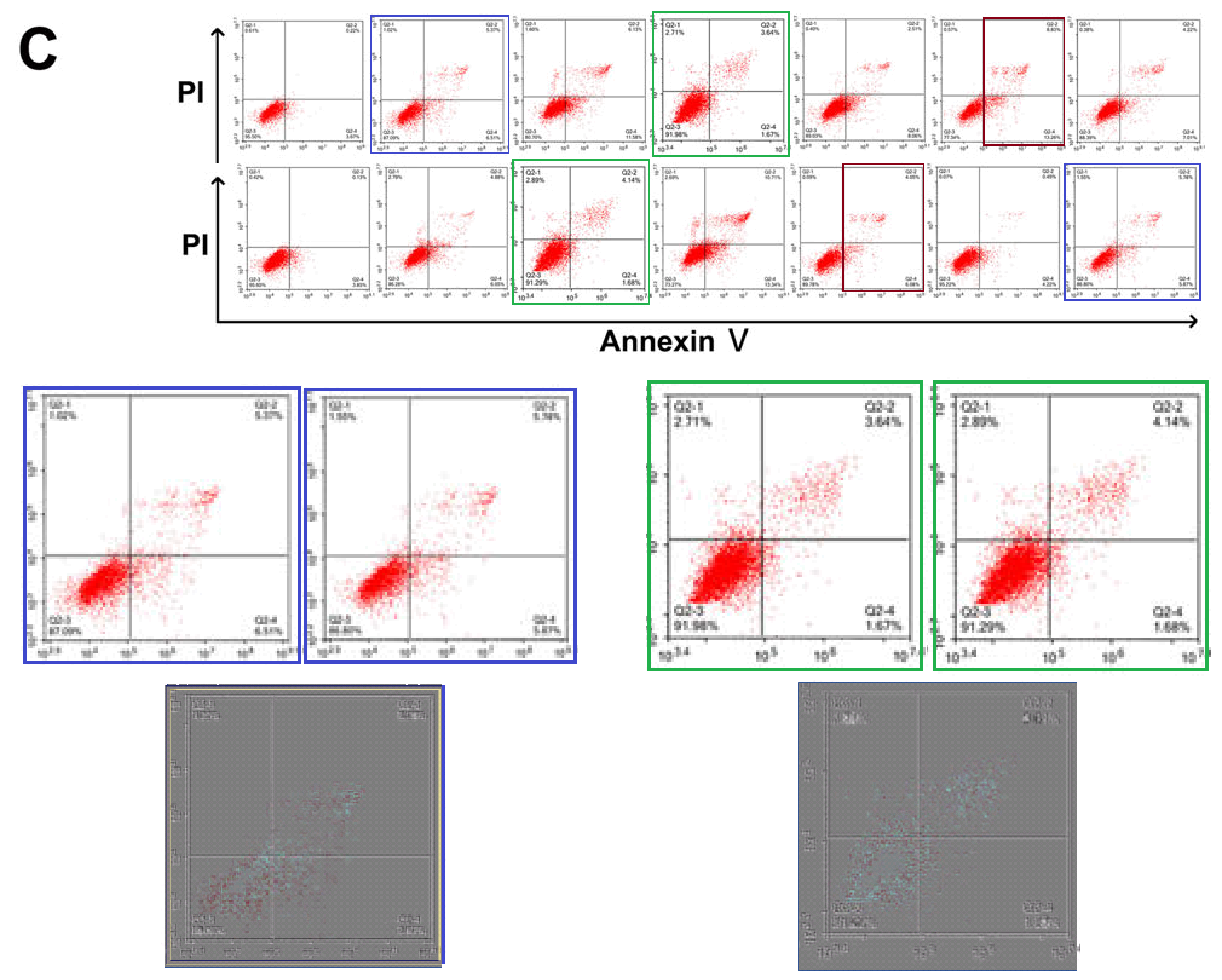
Here are some out-takes. The task of finding the source(s) is left as an exercise for the reader.
 [1] Li Jia, Huimin Zhou, Shujing Wang, Jun Cao, Wei Wei, Jianing Zhang.
[1] Li Jia, Huimin Zhou, Shujing Wang, Jun Cao, Wei Wei, Jianing Zhang.
Deglycosylation of CD147 down-regulates Matrix Metalloproteinase-11 expression and the adhesive capability of murine hepatocarcinoma cell HcaF in vitro
IUBMB Life (2006), DOI:10.1080/15216540600719580 (PubPeer here)
[2] Li Jia, Shujing Wang, Huimin Zhou, Jun Cao, Yichuan Hu, Jianing Zhang.
Caveolin-1 up-regulates CD147 glycosylation and the invasive capability of murine hepatocarcinoma cell lines
Intl J Biochem & Cell Biology (2006), doi: 10.1016/j.biocel.2006.03.019. (PubPeer here)
[3] Li Jia, Shujing Wang, Jun Cao, Huimin Zhou, Wei Wei, Jianing Zhang.
siRNA targeted against matrix metalloproteinase 11 inhibits the metastatic capability of murine hepatocarcinoma cell Hca-F to lymph nodes
Intl J Biochem & Cell Biology (2007). doi: 10.1016/j.biocel.2007.05.023 (PubPeer here)
[4] Li Jia, Huaxin Wang, Shuxian Qu, Xiaoyan Miao, Jianing Zhang.
CD147 regulates vascular endothelial growth factor-A expression, tumorigenicity, and chemosensitivity to curcumin in hepatocellular carcinoma
IUBMB Life (2007), DOI:10.1002/iub.11 (PubPeer here)
[5] Li Jia, Jun Cao, Wei Wei, Shujing Wang, Yunfei Zuo, Jianing Zhang.
CD147 depletion down-regulates matrix metalloproteinase-11, vascular endothelial growth factor-A expression and the lymphatic metastasis potential of murine hepatocarcinoma Hca-F cells
Intl J Biochem & Cell Biology (2007), doi: 10.1016/j.biocel.2007.06.007 (PubPeer here)
[6] Li Jia, Henggui Xu, Yongfu Zhao, Lili Jiang, Jingda Yu, Jianing Zhang.
Expression of CD147 mediates tumor cells invasion and multidrug resistance in hepatocellular carcinoma
Cancer Investigation (2008).DOI:10.1080/07357900802072723 (PubPeer here)
[7] Yongfu Zhao, Li Jia, Xiaoyan Mao, Henggui Xu, Bo Wang, Yongji Liu.
siRNA-targeted COL8A1 inhibits proliferation, reduces invasion and enhances sensitivity to D-limonence treatment in hepatocarcinoma cells
IUBMB Life (2009).DOI:10.1002/iub.151 (PubPeer here)
[8] Li Jia, Wei Wei, Jun Cao, Henggui Xu, Xiaoyan Miao, Jianing Zhang.
Silencing CD147 inhibits tumor progression and increases chemosensitivity in murine lymphoid neoplasm P388D1 cells
Annals of hematology (2009). DOI:10.1007/s00277-008-0678-2 (PubPeer here)
[9] Shujing Wang, Li Jia, Huimin Zhou, Wei Shi, Jianing Zhang.
Knockdown of Caveolin-1 by siRNA Inhibits the Transformation of Mouse Hepatoma H22 Cells In Vitro and In Vivo
Oligonucleotides (2009).DOI:10.1089/oli.2008.0166 (PubPeer here)
[10] Ling He, Jianing Zhang, Lili Jiang, Changgong Jin, Yongfu Zhao, Guang Yang, Li Jia.
Differential expression of Axl in hepatocellular carcinoma and correlation with tumor lymphatic metastasis
Molecular Carcinogenesis (2010).DOI:10.1002/mc.20664 (PubPeer here)
[11] Yongfu Zhao, Xiance Sun, Lili Jiang, Fengjuan Yang, Zhaohai Zhang, Li Jia.
Differential expression of Axl and correlation with invasion and multidrug resistance in cancer cells
Cancer Investigation (2012), doi: 10.3109/07357907.2012.657816 (PubPeer here)
[12] Zhaohai Zhang, Yongfu Zhao, Lili Jiang, Xiaoyan Miao, Huimin Zhou, Li Jia.
Glycomic alterations are associated with multidrug resistance in human leukemia
Intl J Biochem & Cell Biology (2012) doi: 10.1016/j.biocel.2012.04.026 (PubPeer here)
[13] Zhen-Hai Ma, Jin-Hui Ma, Li Jia, Yong-Fu Zhao
Effect of enhanced expression of COL8A1 on lymphatic metastasis of hepatocellular carcinoma in mice
Exper & Therap Medicine (2012).DOI:10.3892/etm.2012.652 (PubPeer here)
[14] Ji Li, Li Jia, Zhen-Hai Ma, Qiu-Hong Ma, Xiao-Hong Yang, Yong-Fu Zhao
Axl glycosylation mediates tumor cell proliferation, invasion and lymphatic metastasis in murine hepatocellular carcinoma
World J Gastroenterology (2012).DOI:10.3748/wjg.v18.i38.5369 (PubPeer here)
[15] Huimin Zhou, Zhaohai Zhang, Chunqing Liu, Changgong Jin, Jianing Zhang, Xiaoyan Miao, Li Jia.
B4GALT1 gene knockdown inhibits the hedgehog pathway and reverses multidrug resistance in the human leukemia K562/adriamycin-resistant cell line
IUBMB Life (2012).DOI:10.1002/iub.1080 (PubPeer here)
[16] Rui Guo, Lei Cheng, Yongfu Zhao, Jianing Zhang, Chunqing Liu, Huimin Zhou, Li Jia.
Glycogenes mediate the invasive properties and chemosensitivity of human hepatocarcinoma cells
Intl J Biochem & Cell Biology (2013). doi: 10.1016/j.biocel.2012.10.006 (PubPeer here)
[17] Hongye Ma, Xiaoyan Miao, Qiuhong Ma, Wenzhi Zheng, Huimin Zhou, Li Jia.
Functional roles of glycogene and N-glycan in multidrug resistance of human breast cancer cells
IUBMB Life (2013).DOI:10.1002/iub.1133 (PubPeer here)
[18] Zhaohai Zhang, Jie Sun, Lihong Hao, Chunqing Liu, Hongye Ma, Li Jia.
Modification of glycosylation mediates the invasive properties of murine hepatocarcinoma cell lines to lymph nodes
PLoS ONE (2013). doi: 10.1371/journal.pone.0065218 (PubPeer here)
[19] Jingchao Xu, Li Jia, Hongye Ma, Yanping Li, Zhenhai Ma, Yongfu Zhao.
Axl gene knockdown inhibits the metastasis properties of hepatocellular carcinoma via PI3K/Akt-PAK1 signal pathway
Tumor Biology (2014). doi: 10.1007/s13277-013-1521-5 (PubPeer here)
[20] Hongye Ma, Lei Cheng, Keji Hao, Yanping Li, Xiaobo Song, Huimin Zhou, Li Jia.
Reversal effect of ST6GAL 1 on multidrug resistance in human leukemia by regulating the PI3K/Akt pathway and the expression of P-gp and MRP1
PLoS ONE (2014). doi: 10.1371/journal.pone.0085113 (PubPeer here)
[21] Dongliang Ren, Li Jia, Yanyan Li, Yanxin Gong, Chen Liu, Xu Zhang, Ning Wang, Yongfu Zhao.
ST6GalNAcII mediates the invasive properties of breast carcinoma through PI3K/Akt/NF-κB signaling pathway
IUBMB Life (2014). DOI:10.1002/iub.1268 (PubPeer here)
[22] Hongye Ma, Huimin Zhou, Peng Li, Xiaobo Song, Xiaoyan Miao, Yanping Li, Li Jia.
Effect of ST3GAL 4 and FUT 7 on sialyl Lewis X synthesis and multidrug resistance in human acute myeloid leukemia
Biochimica et Biophysica Acta (2014). DOI:10.1016/j.bbadis.2014.06.014 (PubPeer here)
[23] Yongfu Zhao, Yanping Li, Hongye Ma, Weijie Dong, Huimin Zhou, Xiaobo Song, Jianing Zhang, Li Jia.
Modification of sialylation mediates the invasive properties and chemosensitivity of human hepatocellular carcinoma
Mol & Cell Proteomics (2014). doi: 10.1074/mcp.m113.034025 (PubPeer here)
[24] Dongliang Ren, Yanyan Li, Yanxin Gong, Jingchao Xu, Xiaolong Miao, Xiangnan Li, Chen Liu, Li Jia, Yongfu Zhao.
Phyllodes tumor of the breast: role of Axl and ST6GalNAcII in the development of mammary phyllodes tumors
Tumor Biology (2014). doi: 10.1007/s13277-014-2254-9 (PubPeer here)
[25] Yanyan Li, Li Jia, Dongliang Ren, Chen Liu, Yanxin Gong, Ning Wang, Xu Zhang, Yongfu Zhao.
Axl mediates tumor invasion and chemosensitivity through PI3K/Akt signaling pathway and is transcriptionally regulated by slug in breast carcinoma
IUBMB Life (2014).DOI:10.1002/iub.1285 (PubPeer here)
[26] Yanyan Li, Li Jia, Chen Liu, Yanxin Gong, Dongliang Ren, Ning Wang, Xu Zhang, Yongfu Zhao.
Axl as a downstream effector of TGF-β1 via PI3K/Akt-PAK1 signaling pathway promotes tumor invasion and chemoresistance in breast carcinoma
Tumor Biology (2014). doi: 10.1007/s13277-014-2677-3 (PubPeer here)
[27] Xu Zhang, Weijie Dong, Huimin Zhou, Hongshuai Li, Ning Wang, Xiaoyan Miao, Li Jia.
α-2,8-Sialyltransferase Is Involved in the Development of Multidrug Resistance via PI3K/Akt Pathway in Human Chronic Myeloid Leukemia
IUBMB Life (2015). DOI:10.1002/iub.1351 (PubPeer here)
[28] Lei Cheng, Shuhang Gao, Xiaobo Song, Weijie Dong, Huimin Zhou, Lifen Zhao, Li Jia
Comprehensive N-glycan profiles of hepatocellular carcinoma reveal association of fucosylation with tumor progression and regulation of FUT8 by microRNAs
Oncotarget (2016). doi: 10.18632/oncotarget.11284. (PubPeer here)
[29] Jialei Hu, Zi Wang, Yujia Shan, Yue Pan, Jia Ma, Li Jia
Long non-coding RNA HOTAIR promotes osteoarthritis progression via miR-17-5p/FUT2/β-catenin axis
Cell Death & Disease (2018). doi: 10.1038/s41419-018-0746-z (PubPeer here)

Donate!
If you are interested to support my work, you can leave here a small tip of $5. Or several of small tips, just increase the amount as you like (2x=€10; 5x=€25). Your generous patronage of my journalism will be most appreciated!
€5.00



A lot of these crap papers in crap journals have a hefty number of citations. Has anyone looked to see if they mention “as faked by Lia et al.” in the Methods?
LikeLike
Let me introduce a more exaggerated professor of Dalian Medical University!!Li dan,Sun,director of Dalian Friendship Hospital affiliated Dalian Medical University.Let me first introduce her title:Master’s tutor of Dalian Medical University, the first author of National Natural Science Foundation and the Liaoning Province Natural Science Foundation, presided over a number of scientific funds, published many SCI, but she has never learned English.Here is the detail,do not laugh out loud!!!
(1)7-O-geranylquercetin-induced autophagy contributes to apoptosis via ROS generation in human non-small cell lung cancer cells,(2)Effects of specific egg yolk immunoglobulin on pan-drug-resistant Acinetobacter baumannii.About these two articles’ corresponding author :Lidan Sun’s misconduct.
1.As a graduate tutor of Dalian Medical University, director of respiratory department of Dalian Friendship Hospital of China,Lidan Sun has no research ability and has never studied English. As is known to all in China that passing through CET-4 (College English Test- rank 4)English examination is the most basic proof that one has learnt English.In the life of her learning language, from childhood to growing up, she never learnt English,Lidan,Sun’s first language is China,her second language is Russian,she has never passed through any English examination to prove that she has learnt English,even she has never learnt English by herself, but she owns many titles, such as graduate tutors, The first author of the National Natural Science Foundation and Provincial Natural Science Foundation, this behavior does not meet the moral standards of scientific research, her achievements are questionable”.She has a lot of privilege ,because she get used to plagiarizing and bribery for years to achieve so called “scientific research expert”.This is so ridiculous that in the two articles above, she is the corresponding author of these two articles, and she has the contribution of designing research!!That is so ridiculous and absurd!! How can a person who has never learnt English design a research from reading a lot of English paper??That is the biggest joke!!
LikeLiked by 1 person
Ah! Ah! That’s very cool!
LikeLike
Research papers from China are unfortunately untrustworthy for several reasons:
1) The chinese society is highly hierarchical, and in a research environment with a big boss, the PhD students and young researchers do and deliver the results they ask for.
2) The generally high corruption level in China has many unfortunate consequences, including hiring leaders in research institutes.
3) Production and sales of counterfeit research products is a huge problem in China and the few respectable researchers that will try to do proper research will most probably be misleaded by using the domestic fake products.
4) Prestige in the chinese state apparatus and lack of democracy and transparancy.
Then we get the fakery shown here in this excellent presentation.
LikeLike
Hi Morty,
It’s sad to see my home country is being commented this way. But I don’t blame you. I myself don’t read any Chinese papers any more and have been rejecting invitations for reviewing manuscripts from Chinese authors.
Tiger
LikeLike
These factors can explain given cases, but I would not generalize.
LikeLike
I am afraid you are most probably wrong Weiss.
Just to exemplify one of my points and how bad it is:
https://www.nature.com/news/the-secret-war-against-counterfeit-science-1.21960
Tiger: I am deeply sorry, but what I wrote was not meant to hurt anyone. Hopefully your excellent documentation and all discussions about these problems can lead to some improvements for the best of research and society!
LikeLike
Hi Morty
You didn’t hurt anybody, truth hurts.
Tiger
LikeLike
Actually a major issue is the destruction of morale and individualism during the MAO years. Both of these are key to scientific integrity. It will take many many years to recreate these and I am afraid during the current communistic dictatorship it will take even longer. Perhaps centuries. So despite some Chinese science being ‘clean’ this is predominantly from people who have ‘moved back’. Of course one would hope there are also local scientists who will be ‘clean’ but the general problems will remain as a concern at least. It will be a long process.
LikeLike
sadly and pathetically using one example to attack the whole community, and still call yourself a scientist? Go morty mortified.
LikeLike
Hi Morty, still, it is one thing to produce consumer products without proper intellectual property standards (i.e. fake watches/shoes for too low price, what actually you suspect to be a fake anyway), and to say that Chinese researchers in general produce falsified science. Besides, I feel much more dangerous the practice of those researchers, who do data picking, force statistical power, use loading control libraries, hide sample variance, hide uncut blots, run identical samples in FACS analysis and label them distinctly, bully their fellows, in short: produce fake data and create a toxic work environment of a data factory, without provable signs. I can mention here concrete examples where no action has been taken, or pathetic corrections have been released. They are all non-Chinese, and indeed, are influental figures of our time.
LikeLike
I would not say that Chinese researchers ‘in general” faking data, but the truth is that making up data is widely accepted or acknowledged as fairly common practice you’ve got to do there. The worst? No one really cares.
LikeLike
If you get positive results, there is no need to fake data. Its when you face demanding publication standards and kids to feed and you dont have positive results is when data might be faked.
LikeLike
Precisely for also the reason I mentioned above
LikeLike
In many cases it is not a “good data / bad data” issue. There are no data at all. When people go to a paper-mill to have a paper assembled from hand-drawn FACS plots and odd scraps of Western blot and then published in their name, it is not because they have been doing experiments themselves but failing.
Case in point, the production line of micro-RNA papers
LikeLike
“I can mention here concrete examples where no action has been taken, or pathetic corrections have been released. They are all non-Chinese, and indeed, are influental figures of our time.”
Could you mention the “concrete examples examples where no action has been taken, or pathetic corrections have been released.” I believe that those things happens, It would be useful to compare lists.
LikeLike
https://pubpeer.com/publications/2D084E91EDC85D6C23BC63ACF5E6A8
https://pubpeer.com/publications/1C1292425188400AB6A2B0C4B6E2D0
https://pubpeer.com/publications/6CC01D626651DFF7D30AECF0B762AB
https://pubpeer.com/publications/8362F93170BAE224FC3006A9425993
https://pubpeer.com/publications/E89EBC1AE561A534E027E9D19AB9F9
https://pubpeer.com/publications/454C6F77CE3CB3C28C3A6AF75A371D
https://pubpeer.com/publications/DEA53E3CBABFAAB1F4D6959A46DA9E
https://pubpeer.com/publications/2EF48BB746D7ACEB9F7FA310D24D73
etc etc etc
Many of them have been reported/discussed by others as well.
LikeLike
This reply is to NMH: Then these must-fake-data-otherwise-starving-their-own-kids individuals would be much better off finding another career for a living.
To make it clear: Doing science is not for everybody. Being an engineer is not for everyone either. The same logic applies to every profession. People have the right choosing what to do but following the standard practice and obeying the rules in the chosen field. It is that simple.
No one points a gun to your head to force you fake data for publication. Can’t publish? then don’t! The world is full of opportunities and fun outside a science lab.
LikeLike
Well said!
LikeLike
“No one points a gun to your head to force you fake data for publication. Can’t publish? then don’t! The world is full of opportunities and fun outside a science lab.”: It’s not that easy…. sometimes faking data is the only way to survive for some people. Specially, for people from certain countries it is the only way to a better life. It’s ridiculous to say people should obey “rules” but at the time you’ve got bad results nobody will help you or even will give you any useful hint or advice and they’re just saying to correct it before deadline without asking for more resources or help. I personally think if someone fake something, the “publish or perish” academic system is the who that should be blamed.. Obviously, nobody has sadist for faking data without any apparent reason. It’s really ridiculous that researchers have the lowest income with highest stress and pressure in their life and society at the general level doesn’t give a shit to researchers’ situation but only according to “rules” highest ethical standards are expected from researchers. It’s not fair…
LikeLike
” It’s really ridiculous that researchers have the lowest income “. Supply and demand surely? Too many people want to do it.
LikeLike
One could argue that data generation is as important as management of the generators. But in the end the data generators are not paid enough for a middle-class lifestyle in the USA. I know…Im a permanent post-doc in the US, nearing retirement. It stinks. Im in no way suggesting that people should cheat, but I can certainly see why they do to get ahead. Its too much of a big jump from the pay of a data generator to the pay of a manager.
LikeLike
Do you look at faculty hiring in USA lately ? High emphasis on NIH grants, CNS papers and outrageous H INDEX. The only way to succeed in academia is to work for superstars in the hot fields who can give you papers and grants and promote your career at the expense of slavery, bullying and misconduct.
LikeLike
Reply to zmudzka June 15, 2019
What you write is true.
虎仔 (@TigerBB8) June 15, 2019 gives very good advice;
“much better off finding another career for a living”.
Too many of us, me included, have, or had, an idealised picture of how science is practised.
I think we have learned this idealised picture from children’s books, or had an inspirng, but misguided science teacher.
LikeLike
In my country, we consider academia as an “ivory tower”. Scientists, especially those in universities as professors, are the group of people with the highest moral standard. Traditionally, we also have the understanding of “Once your teacher, forever your father”, a very high respect to professors. But in reality, scientists/professors are the same corrupted people as in gov and in industry.
LikeLike
Yeah, I have heard this teacher-father kitsch before from some undergrad fellows who were on the way of deceiving me 🙂 Students should consider their teachers as teachers, and their fathers as fathers – and these two should not be mixed up, at least in my culture 🙂 Anyway, my opinion remains the same I have stated above: I do not like tags and general statements about countries/people/Chinese researchers. Though I accept the arguments above that certain countries due to their social rules and cultures warrant attention when it comes to their scientific publications.
LikeLike
Reply to Weiss June 16, 2019
Thanks for posting Pubpeer pages with problematic data. Looks like 3 or 4 groups of researchers.
It would be more useful if you listed each group and then put the Pubpeer pages for them,
I had only heard of one of the groups, headed by Richard Moriggl.
This suggests there is a lot more out there.
LikeLike
Answer to Zebedee: RM has many shared papers with JPT, and all those have western blot loading controls run on separate gels. Some of them are on Pub Peer.
LikeLike
Pingback: De coccio - Ocasapiens - Blog - Repubblica.it
Let me introduce a more exaggerated professor of Dalian Medical University!!Li dan,Sun,director of Dalian Friendship Hospital affiliated Dalian Medical University.Let me first introduce her title:Master’s tutor of Dalian Medical University, the first author of National Natural Science Foundation and the Liaoning Province Natural Science Foundation, presided over a number of scientific funds, published many SCI, but she has never learned English.Here is the detail,do not laugh out loud!!!
(1)7-O-geranylquercetin-induced autophagy contributes to apoptosis via ROS generation in human non-small cell lung cancer cells,(2)Effects of specific egg yolk immunoglobulin on pan-drug-resistant Acinetobacter baumannii.About these two articles’ corresponding author :Lidan Sun’s misconduct.
1.As a graduate tutor of Dalian Medical University, director of respiratory department of Dalian Friendship Hospital of China,Lidan Sun has no research ability and has never studied English. As is known to all in China that passing through CET-4 (College English Test- rank 4)English examination is the most basic proof that one has learnt English.In the life of her learning language, from childhood to growing up, she never learnt English,Lidan,Sun’s first language is China,her second language is Russian,she has never passed through any English examination to prove that she has learnt English,even she has never learnt English by herself, but she owns many titles, such as graduate tutors, The first author of the National Natural Science Foundation and Provincial Natural Science Foundation, this behavior does not meet the moral standards of scientific research, her achievements are questionable”.She has a lot of privilege ,because she get used to plagiarizing and bribery for years to achieve so called “scientific research expert”.This is so ridiculous that in the two articles above, she is the corresponding author of these two articles, and she has the contribution of designing research!!That is so ridiculous and absurd!! How can a person who has never learnt English design a research from reading a lot of English paper??That is the biggest joke!!
LikeLike
Maybe she uses Google Translator. But yeah, kinda´ absurd story. Anyway, there are labs where the technicians are first authors, and I also have seen papers where undergraduates are corresponding authors (that one is not easy to catch, you should know the persons).
LikeLike
If you have not learnt Chinese,Japanese,how do you use translator to review Chinese or Japanese article? At least someone must have language foundation that can use translator.The google translator can not translate full of all,it must combine with someone’s reading English ability!!
LikeLike
Pingback: Blagosklonny’s lawyer threatens me to love Oncotarget or else – For Better Science
Pingback: The full-service paper mill and its Chinese customers – For Better Science
2020 retraction:
Cell Death Dis. 2017 May 25;8(5):e2830. doi: 10.1038/cddis.2017.223.
Functional screen analysis reveals miR-3142 as central regulator in chemoresistance and proliferation through activation of the PTEN-AKT pathway in CML.
Zhao L1, Shan Y1, Liu B1, Li Y1, Jia L1.
Author information
1
College of Laboratory Medicine, Dalian Medical University, Dalian 116044, Liaoning Province, China.
https://pubpeer.com/publications/CD27B1662B633DDC750D9E7C774255
2020 retraction notice.
https://www.nature.com/articles/s41419-020-2319-1
The Editors-in-Chief have retracted this article because the PTEN-shRNA panel in Fig. 6d appears to be the same as the Anti3142+PTEN-shRNA panel in Fig. 6h. The Editors-in-Chief therefore no longer have confidence in the integrity of the data in Fig. 6. All of the authors disagree with this retraction.
LikeLike
2020 retraction:
Cell Death Dis. 2018 Jun 7;9(6):688. doi: 10.1038/s41419-018-0706-7.
Aberrant mannosylation profile and FTX/miR-342/ALG3-axis contribute to development of drug resistance in acute myeloid leukemia.
Liu B1, Ma X1,2, Liu Q1, Xiao Y1, Pan S1, Jia L3.
Author information
1
College of Laboratory Medicine, Dalian Medical University, Dalian, 116044, Liaoning Province, China.
2
Department of Clinical Laboratory, the First Affiliated Hospital of Dalian Medical University, Dalian, 116011, Liaoning Province, China.
3
College of Laboratory Medicine, Dalian Medical University, Dalian, 116044, Liaoning Province, China. jiali0386@sina.com
https://pubpeer.com/publications/8074E3B2620A4F09B2399FFB40D85F
2020 retraction notice.
https://www.nature.com/articles/s41419-020-2320-8
The Editors-in-Chief have retracted this article because there appears to be replication between a group of data points in the 10.95% quadrant of the U/A (VCR) ALG3 shRNA panel of Fig. 3f and a group of data points in the 4.52% quadrant of the T/A (Paxcitaxel) ALG3 shRNA panel of Fig. 3f. The Editors-in-Chief therefore no longer have confidence in the integrity of the data in Fig. 3. All of the authors disagree with this retraction.
LikeLike
The 2 retractions by Cell Death and Disease draw attention that the journal is diseased itself.
https://www.nature.com/cddis/editors
For starters:
Richard Knight, one the Editors-in-Chief, 6 retractions, but not a dickybird from the journal.
The readers are left ignorant of that fact.
Enhanced IL-17 signalling following myocardial ischaemia/reperfusion injury.
Barry SP, Ounzain S, McCormick J, Scarabelli TM, Chen-Scarabelli C, Saravolatz LII, Faggian G, Mazzucco A, Suzuki H, Thiemermann C, Knight RA, Latchman DS, Stephanou A.
Int J Cardiol. 2013 Mar 10;163(3):326-334. doi: 10.1016/j.ijcard.2011.08.849. Epub 2011 Oct 24.
Retraction in: Int J Cardiol. 2019 Jan 1;274:404. PMID: 22030025
ERK and the F-box protein betaTRCP target STAT1 for degradation.
Soond SM, Townsend PA, Barry SP, Knight RA, Latchman DS, Stephanou A.
J Biol Chem. 2008 Jun 6;283(23):16077-83. doi: 10.1074/jbc.M800384200. Epub 2008 Mar 31.
Retraction in: J Biol Chem. 2018 Mar 30;293(13):4954. PMID: 18378670
Cardiac release of urocortin precedes the occurrence of irreversible myocardial damage in the rat heart exposed to ischemia/reperfusion injury. Knight RA, Chen-Scarabelli C, Yuan Z, McCauley RB, Di Rezze J, Scarabelli GM, Townsend PA, Latchman D, Saravolatz L, Faggian G, Mazzucco A, Chowdrey HS, Stephanou A, Scarabelli TM. FEBS Lett. 2008 Mar 19;582(6):984-90. doi: 10.1016/j.febslet.2008.02.035. Epub 2008 Feb 22.
Retraction in: FEBS Lett. 2018 Feb;592(4):657. PMID: 18295601
STAT-1 facilitates the ATM activated checkpoint pathway following DNA damage.
Townsend PA, Cragg MS, Davidson SM, McCormick J, Barry S, Lawrence KM, Knight RA, Hubank M, Chen PL, Latchman DS, Stephanou A.
J Cell Sci. 2005 Apr 15;118(Pt 8):1629-39. Epub 2005 Mar 22.
Retraction in: J Cell Sci. 2015 Mar 1;128(5):1064. PMID: 15784679
The carboxyl-terminal activation domain of the STAT-1 transcription factor enhances ischemia/reperfusion-induced apoptosis in cardiac myocytes.
Stephanou A, Scarabelli TM, Townsend PA, Bell R, Yellon D, Knight RA, Latchman DS.
FASEB J. 2002 Nov;16(13):1841-3. Epub 2002 Sep 5.
Retraction in: FASEB J. 2018 Apr;32(4):2315.
PMID: 12223448
Antiapoptotic activity of the free caspase recruitment domain of procaspase-9: a novel endogenous rescue pathway in cell death.
Stephanou A, Scarabelli TM, Knight RA, Latchman DS.
J Biol Chem. 2002 Apr 19;277(16):13693-9. Epub 2002 Feb 1.
Retraction in: J Biol Chem. 2015 Jan 16;290(3):1454. PMID: 11825888
LikeLike
All of the authors disagree with this retraction. How refreshing to see that.
LikeLike
So this happened: 15 papers by Jia Li were condemned by the National Natural Science Foundation, after a thorough investigation (possibly reading this post).
Decision on handling misconduct cases investigated and handled in 2021 (third batch).
The first of 22 Decisions is as follows (thanks Google Translate for translation). Criticism of the majority of those papers has not seen any response from the journal or the publisher, but perhaps the NNSF show-trial verdicts will goad them into doing something. The second Decision in the webpage relates to five of the same papers, dwelling on the role of another author.
Regarding the problem of image duplication and rotation in the papers published by Jia Li et al.
And provide false information in the project application, progress/final report
Processing decision
National Science and Technology Supervision Office (2021) No. 78
The Supervision Committee of the National Natural Science Foundation of China launched an investigation into the suspected academic misconduct in the papers published by Jia Li of Dalian Medical University. The papers involved are as follows:
Paper 1: “Li Jia, Huimin Zhou, Shujing Wang, Jun Cao, Wei Wei, Jianing Zhang*. Deglycosylation of CD147 down-regulates Matrix Metalloproteinase-11 expression and the adhesive capability of murine hepatocarcinoma cell HcaF in vitro. IUBMB Life, 2006 , 58(4):209-216.” (marked with fund number 30470400)
Paper 2: “Li Jia, Shujing Wang, Jun Cao, Huimin Zhou, Wei Wei, Jianing Zhang*. siRNA targeted against matrix metalloproteinase 11 inhibits the metastatic capability of murine hepatocarcinoma cell HcaF to lymph nodes. The International Journal of Biochemistry & Cell Biology , 2007, 39(11): 2049-2062.” (marked with fund numbers 30470400, 30670466)
Paper 3: “Li Jia, Huaxin Wang, Shuxian Qu, Xiaoyan Miao, Jianing Zhang*. CD147 regulates vascular endothelial growth factor-A expression, tumorigenicity, and chemosensitivity to curcumin in hepatocellular carcinoma. IUBMB Life, 2008, 60(1): 57-63.” (marked with fund numbers 30470400, 30670466)
Paper 4: “Li Jia, Henggui Xu, Yongfu Zhao, Lili Jiang, Jingda Yu, Jianing Zhang*. Expression of CD147 Mediates Tumor Cells Invasion and Multidrug Resistance in Hepatocellular Carcinoma. Cancer Investigation, 2008, 26(10):977-983 .” (marked with fund number 30670466)
Paper 5: “Li Jia, Wei Wei, Jun Cao, Henggui Xu, Xiaoyan Miao, Jianing Zhang*. Silencing CD147 inhibits tumor progression and increases chemosensitivity in murine lymphoid neoplasm P388D1 cells. Annals of Hematology, 2009, 88(8):753 -760.” (marked with fund number 30670466)
Paper 6: “Zhaohai Zhang # , Yongfu Zhao # , Lili Jiang, Xiaoyan Miao, Huimin Zhou, Li Jia*. Glycomic alterations are associated with multidrug resistance in human leukemia. The International Journal of Biochemistry & Cell Biology, 2012, 44(8 ):1244-1253.” (marked with fund number 8107415)
Paper 7: “Huimin Zhou, Zhaohai Zhang, Chunqing Liu, Changgong Jin, Jianing Zhang, Xiaoyan Miao, Li Jia*. B4GALT1 gene knockdown inhibits the hedgehog pathway and reverses multidrug resistance in the human leukemia K562/adriamycin-resistant cell line. IUBMB Life, 2012, 64(11):889-900.” (marked with fund number 81071415)
Paper 8: “RuiGuo # , Lei Cheng # , Yongfu Zhao, Jianing Zhang, Chunqing Liu, Huimin Zhou, Li Jia*. Glycogenes mediate the invasive properties and chemosensitivity of human hepatocarcinoma cells. The International Journal of Biochemistry & Cell Biology, 2013, 45(2):347-358.” (marked with fund number 81071415)
Paper 9: “Hongye Ma # , Xiaoyan Miao # , Qiuhong Ma, Wenzhi Zheng, Huimin Zhou, Li Jia*. Functional roles of glycogene and N-glycan in multidrug resistance of human breast cancer cells. IUBMB Life, 2013, 65(5 ): 409-422.” (marked with fund number 81071415)
Paper 10: “Zhaohai Zhang # , Jie Sun # , Lihong Hao, Chunqing Liu, Hongye Ma, Li Jia*. Modification of Glycosylation Mediates the Invasive Properties of Murine Hepatocarcinoma Cell Lines to Lymph Nodes. PLoS One, 2013, 8(6) :e65218.” (marked with fund numbers 81071415, 81271910)
Paper 11: “Hongye Ma # , Lei Cheng # , Keji Hao, Yanping Li, Xiaobo Song, Huimin Zhou, Li Jia*. Reversal Effect of ST6GAL1 on Multidrug Resistance in Human Leukemia by Regulating the PI3K/Akt Pathway and the Expression of P -gp and MRP1. PLoS One, 2014, 9(1):e85113.” (marked with fund numbers 81071415, 81271910)
Paper 12: “Hongye Ma # , Huimin Zhou # , Peng Li, Xiaobo Song, Xiaoyan Miao, Yanping Li, Li Jia*. Effect of ST3GAL4 and FUT7 on sialyl Lewis X synthesis and multidrug resistance in human acute myeloid leukemia. Biochimica et Biophysica Acta, 2014, 1842(9):1681-1692.” (marked with fund number 81271910)
Paper 13: “Yongfu Zhao, Yanping Li, Hongye Ma, Weijie Dong, Huimin Zhou, Xiaobo Song, Jianing Zhang, Li Jia*. Modification of Sialylation Mediates the Invasive Properties and Chemosensitivity of Human Hepatocellular Carcinoma. Molecular and Cellular Proteomics, 2014, 13(2):520-536.” (marked with fund numbers 81071415, 81271910)
Paper 14: “Lei Cheng # , Shuhang Gao # , Xiaobo Song, Weijie Dong, Huimin Zhou, Lifen Zhao, Li Jia*. Comprehensive N-glycan profiles of hepatocellular carcinoma reveal association of fucosylation with tumor progression and regulation of FUT8 by microRNAs. Oncotarget, 2016, 7(38):61199-61214.” (marked with fund numbers 81271910, 81472014)
Paper 15: “Jialei Hu # , Zi Wang # , Yujia Shan, Yue Pan, Jia Ma, Li Jia*. Long non-coding RNA HOTAIR promotes osteoarthritis progression via miR-17-5p/FUT2/β-catenin axis. Cell Death and Disease, 2018, 9(7):711.” (labeled fund number 81772277)
After investigation, the above-mentioned papers have problems such as photo duplication and rotation. As the corresponding author of papers 7, 8, 9, 10, 11, 12, 14, 15 and the first author of papers 1, 2, 3, 4, and 5, Jia Li is mainly responsible for the misconduct of papers 6, 13 The behavior bears secondary responsibility. The papers 1, 2, 3, 4, 5, and 6 are also listed in the application form of the National Natural Science Foundation of China (approval number 81271910), and the papers 2, 3, 4, and 5 are listed in the country In the application form of the Natural Science Foundation Project (approval number 81071415), the papers 6, 7, 8, 9, 10, 11, and 13 are listed in the application form of the National Natural Science Foundation Project (approval number 81472014), and the papers 6, 7 , 8, 9, 10, 11, 13, and 14 were included in the application of the National Natural Science Foundation of China (approval number 81772277), and the papers 6, 7, 8, 9, 13 were included in the National Natural Science Foundation of China (approved No. 81071415) In the progress report, Papers 10, 11, and 13 are included in the progress report of the National Natural Science Foundation of China (approval number 81271910), and Paper 14 is included in the progress report of the National Natural Science Foundation of China (approval number 81472014) , Included papers 6, 7, 8, 9, 10, and 11 in the final report of the National Natural Science Foundation of China (approval number 81071415), and included papers 10, 11, and 14 in the National Natural Science Foundation of China (approval number) 81271910) in the final report.
After deliberation at the 10th meeting of the 5th Supervisory Committee of the National Natural Science Foundation of China (Life Medicine Professional Committee), and approved by the 13th Executive Meeting of the National Natural Science Foundation of China in 2021, it was decided to comply with the 35th National Natural Science Foundation Regulation Article 4, “The National Natural Science Foundation of China Supervision Committee’s Measures for Handling Misconduct in the Funding of Science (for Trial Implementation)” Article 16, Paragraph 2 and Article 17, Paragraph 3, revoke Jia Li The National Natural Science Foundation of China project “Exploration of glycoprotein N-sugar chain as a marker for predicting tumor resistance” (approval number 81071415), “Characteristic changes of glycogenes in leukemia multidrug resistance and its diagnostic value” (approved No. 81271910), “Screening, identification and functional study of leukemia drug-resistant sugar complex markers” (Approval No. 81472014), “Screening of carbohydrate markers for liver metastasis of colorectal cancer” (Approval No. 81772277), withdrawn Funds have been allocated for the above four projects, and Jia Li’s national natural science fund project application qualification has been cancelled for 5 years (July 20, 2021 to July 19, 2026), and Jia Li has been criticized in a circular.
National Natural Science Foundation of China
September 19, 2021
LikeLike