This is a story of scientific success, of German genius, of cancer almost conquered, thanks to the vision of a great man from Hannover. And yet it is also a story of a sad loss for German and international academia. Ruben Plentz, celebrated by German media as “Fighter against the Cancer“, chose to focus his career on clinical medicine, despite his tremendous academic achievements in cancer research. Was it was because malicious critics harassed him with frivolous investigations and made him correct his papers for duplications, envious at all those peer-reviewed cancer cures the great doctor invented?
Professor Plentz honed his scientific skills under the best of the best: the famous telomere researcher Karl Lenhard Rudolph, recently honoured by Leibniz Society and by the DFG, and the oncologist Nabeel Bardeesy in Harvard, USA, whose creative contribution to science is commemorated on PubPeer. The beginning and the centre of Professor Plentz’ academic career has been the famous Hannover Medical School MHH, where tracheas and other organs are grown from stem cells, thanks to works of such international stars like Heike and Thorsten Walles and the winner of MHH best dissertation award, Philipp Jungebluth. MHH also prides itself of hosting Jungebluth’s and Walles’ mentor Paolo Macchiarini as adjunct professor, to this very day.
But this is the story of Professor Plentz, who is now head of the internal medicine clinic at the communal hospital Klinikum Bremen Nord. His former boss at the University of Tübingen, Nissar Malek, described Plentz as “a progressive man of science”, and indeed Plentz, even if far away, retained his adjunct academic affiliation in Tübingen, a professorship he never omits, and neither shall I. What follows is to celebrate Professor Plentz’ creative approach to cancer cure discovery, by presenting some examples a reader of my site posted on PubPeer. If only more German doctors would take his scientific example, cancer would be a thing of the past!

Inspired by his famous MHH mentor K Lenhard Rudolph, Dr Plentz established the role of telomeres in cancer. He predicted and developed what seems to have been a telomere-lenght biomarker assay for liver cancer and liver cirrhosis, which earned Plentz in 2004 a cash award, and later the Venia Legendi habilitation at MHH. It is not clear what else that life-saving biomarker technology invention was used for since. Predicting scientific discoveries is one of key strengths of Professor Plentz, only the best of the best of scientists have such talent. Detecting cancer is nice, but thanks to professor Plentz we now also know how to cure it. Important message in this context:
“The University of Tuebingen investigated these issues and did not find evidence of misconduct”.
Scratchy assay
In order to study cancer cell metastasis, Professor Plentz often applied the so-called “scratch assay”. Metastasis happens when cancers mutae into an invasive phenotype, described as “epithelial mesenchymal transition (EMT)”, where tumour cells leave the original localised growth to squeeze themselves through tissues and blood vessel walls in order to spread throughout the body, via blood and lymph. This transformation is a complex genetic and physiological event, but luckily intellectually superior cancer researchers like Professor Plentz can simulate it with a simple “scratch assay”. There, you take a Petri dish of cells, put them under a climatised microscope, and make a scratch through the cell carpet. From time to time, you look through microscope at that same field of cells again and check if they managed to move into the empty scratch area, and how far did they get. Cultured cells are highly motile, and if you find a way to inhibit their movement, e.g. by some pharmacological inhibitor: you announce a cure for cancer metastasis. Professor Plentz found several such cures, all of them inhibited cells from walking to where they should not be walking. The results were so perfectly reproducible, that even some images of those scratch assays looked exactly the same.
In the paper Palagani et al PLOS One 2012, the Plentz lab in Tübingen discovered the Notch pathway as anti-cancer target and proposed to cure pancreatic cancer with the gamma secretase inhibitor (GSI). It worked like a charm. Plentz’s mentor from MHH, and head of gastroenterology clinic there, Professor Michael Manns is one of the authors and must be immensely proud that the solution to curing pancreatic ductal adenocarcinoma (PDAC) is that simple. Also Plentz’ former superior from Tübingen, Nissar Malik, is coauthor, as he is on the corrected papers discussed below.
This is how it looks on PubPeer, where that reader of my site shared this illustration:
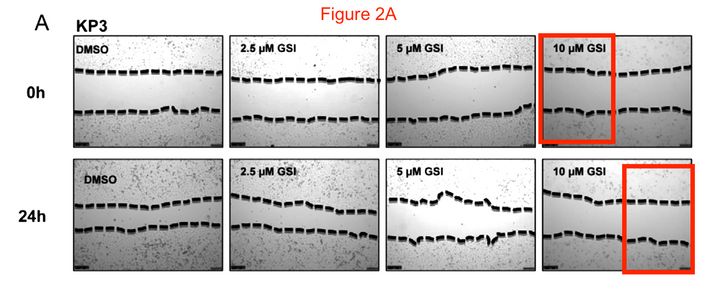
The red-boxed areas look surprisingly same, but they are supposed to be 24 hours apart. Which does prove the inhibitor works best at 10 µM, at that concentration it freezes the cells in time! Strange thing though that the borders were drawn differently.
Two years later: another breakthrough discovery: Professor Plentz found out that combining the Notch inhibitor GSI with an inhibitor of JAK-STAT signalling pathway achieves an even better effect on pancreatic cancer. The paper Palagani et al Carcinogenesis 2014 also used scratch assays to prove that:

Now this a bit confusing. Granted, GSI can freeze cells in time at 10µM as proven before, but why suddenly are cells treated with only solvent DMSO so similar to cells treated with JAK-STAT inhibitor AG-490, 24 hours later? And why do the dish for AG-490 and the dish with combined therapy look so similar to each other? And why again are the borders drawn differently? Never mind such minor issues, the images are probably mere illustrations anyway, for the authors bear good news: “potential therapeutic approach for PDAC” is found.
They do cautiously add: “However, the study design limits the direct transfer into the clinic and the impact of dual treatment in patients with PDAC remains still to be determined“. Maybe it would help duplicating patient’s X-rays, to achieve cancer regression? Combined with the two pharmacological inhibitors? A back scratcher to rub them in, what with the scratch assay evidence? Countless lives can be saved!
The world was not yet ready or still unwilling, so the Plentz lab developed another cure for pancreatic cancer: The proteasome inhibitor Argyrin A, exactly its analogue Argyrin F (AF). It cured mice of cancer, says Professor Plentz, and of course it also prevented cancer cells in scratch assay from going into the no-man’s land. Metastatic cancer, scratched.

Again, somehow the DMSO dish looks exactly like the one treated with AF, only the borders were once again drawn differently. Maybe Plentz’s former teacher from USA, Nabeel Bardeesy, could explain, since he is well versed with image duplications (eg, here)? After all, Bardeesy in turn trained with the great PubPeer star Ronald DePinho, former president of MD Anderson I wrote about before.
Bothersome corrections
Those strange image duplications are a tricky thing. Even impeccable top researchers like Professor Plentz occasionally have to correct papers because of some malicious nitpicking Luddites. Like El-Khatib et al Hepatology 2013, where the Plentz team discovered that bile duct cancer can be cured with an inhibitor of hedgehog molecular signalling pathway. That paper had a duplicated image of cells, in the 2014 correction the publisher Wiley took the blame upon itself as it “regrets the error“. Rightly so, because for that same discovery Professor Plentz was awarded with the Adolf-Kußmaul-Prize of the regional Gastroenterology society in cash.
More recently, in November 2018, PLOS One issued a correction for the paper El-Khatib et al 2013. It namely re-used a picture from the El-Khatib et al Hepatology 2013, because it turned out, bile duct cancer cholangiocacinoma can be cured not just with AF, but even better with the Notch inhibitor GSI on top. Somehow, this Notch inhibition led to even more image duplication, as PLOS One explained:
“This article [1] was republished on October 22, 2018 to address concerns raised post-publication regarding the Figs 3, 4, and 5.
- The image presented for the DMSO/SZ1 experiment in Fig 3A was duplicated from a previous publication (Fig 5A in [2]).
- In Fig 4, duplicate images were shown for SZ1 treated with DMSO and 5μM GSI IX, and the wrong image was shown for the 20μM panel.
- In Fig 5, duplicate images were presented in the “Snail” and “E cadherin” panels of Fig 5A and Fig 5B, and the molecular weight labels for these panels in Fig 5B were incorrect.
The University of Tuebingen investigated these issues and did not find evidence of misconduct.”

Of course there was no misconduct. German doctors are trying to save lives of countless cancer patients, with groundbreaking research on inhibitors, why would anyone in their right mind have a problem with some duplicated Petri dishes or this western blot here?
The Snail and E-cadherin blots looks so similar for different experiments with two different cell lines exactly because the treatment with GSI is so efficient! The doctors at University of Tübingen understand such complex issues and saw that of course there never was any misconduct.
You pedestrian pipette-swingers namely have no clue how great science is done. Professor Plentz’ prediction that bile duct cancer can be cured by Notch inhibitor GSI made it as Zender et al 2013 into the elite journal Cancer Cell, impact factor 23. This should shut up the last of the sceptics, even if the paper had some “duplication of actin blots”. Luckily, the correction made clear that “none of these errors affect the results or conclusions of this work“. The reader of my site visualised it on PubPeer:

Those are some very strange duplications, difficult to have happened by mistake or oversight. Yet if this proves anything, it proves that Professor Plentz is a true visionary genius whose scientific predictions always hit the mark, before the first sample is even loaded on a gel. And now he left science to head a hospital department in Bremen. Hopefully University of Tübingen, or maybe even MHH, will find a way to convince him to return. They should hurry up, because what is their loss, is others’ gain: Bremen politicians plan to open a medical school at the University of Bremen, in collaboration with Bremen hospitals. Readers of my site know what place of excellence University of Bremen is; their local star, diabetes researcher Katrin Maedler , would sure be interested to join creative forces with such fellow expert for curing diseases as Professor Plentz. Let us all pray that the Bremen medical school will become a reality soon. Science needs to have Professor Plentz genius back.
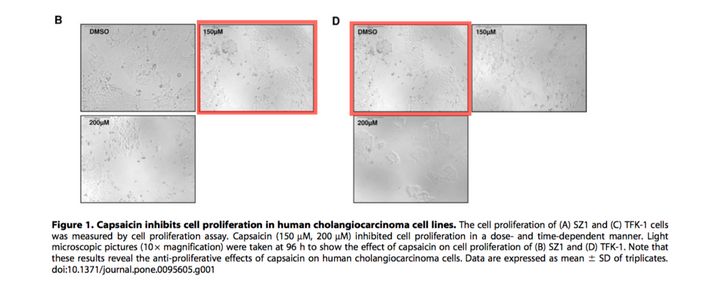
The man simply does not get the recognition he deserves. With such strong body of science in support of GSI Notch inhibitor as cancer therapy it would be highly irresponsible not to listen to Professor Plentz’ scientific advice in the clinic. Yet those philistines, who never discovered anything useful, instead complained of duplications and burdened the Professor and his Tübingen colleagues with pointless investigations into non-existing misconduct. Anyway, did you know Professor Plentz found out that chilli can cure cholangiocacinoma? Yes, chilli peppers, precisely their active ingredient capsaicin. The authors around Plentz concluded:
“The use of capsaicin as a food supplement to inhibit Hedgehog signaling might therefore be of additional therapeutic benefit in patients with CC”.
This was discovered in Wutka et al PLOS One 2014, but the highly innovative study also had to be corrected, two years later, for a duplicated image of cells. Somehow the chilli effect on one cell type made it look just like another cell type, before the chilli treatment.
Those silly loading controls!
In 2016, Professor Plentz published the paper Barat et al Mol Carcinogenesis 2016. The man who keeps discovering new cures for cholangiocarcinoma, had yet another idea: to try the inhibitor of the hepatocyte growth factor receptor-kinase, MET. As it is custom with doctors of supreme intelligence, Plentz’ hypothesis worked out neatly when put in the practice, another cancer cure was identified:
“LY2801653 is an effective inhibitor and suppress the proliferation of CCC cells as well as the growth of xenograft tumors. Therefore, inhibition of c-MET could be a possible alternative approach for the treatment of human CCC”
The technology which worked the magic here, was western blot, like in the Cancer Cell paper before. This is the comment that reader of my site shared on PubPeer:

Most peculiar. Why would same loading control bands apply to untreated cells and those treated with MET inhibitor, as red boxes suggest? Why would loading controls be so interchangeable between utterly unrelated experiments, as highlighted in blue and green? Luckily, and is “just” loading controls, a mere formality some pesky peer reviewers insist on. And we should always recall the Mantra of Tübingen:
“The University of Tuebingen investigated these issues and did not find evidence of misconduct”
Update 25.01.2019. Most peculiar thing: When PLOS One corrected the Figure 5 in Prof Plentz’ paper El Khatib et al 2013, a mathematical peculiarity was overlooked.
According to the labels, the gels shown should have 7 lanes. But some have 6, some have 8 lanes.

Unfortunately, the issues remained in the new version of the figure which was republished in November 2018. But then again, since other panels do contain 7 lanes, the average as well as the mean number of lanes per panel remains 7, with minor error bars, so it’s probably OK?

Donate!
If you are interested to support my work, you can leave here a small tip of $5. Or several of small tips, just increase the amount as you like (2x=€10; 5x=€25). Your generous patronage of my journalism will be most appreciated!
€5.00


2019 expression of concern.
https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0226310
LikeLike
Pingback: mTOR: conclusions not affected? – For Better Science